

Table of SynGAP1 Isoform α2 (UniProt Q96PV0-1) Missense Variants.
| c.dna | Variant | SGM Consensus | Domain and Structure information: based on WT protein | Annotated databases | Deep learning-based pathogenicity predictions | Folding stability-based pathogenicity predictions | Sequence/structure-based pathogenicity predictions | Phase Separation | Evolutionary/physical properties | Molecular Dynamics-based analysis | DOI | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Domain | IUPred2 | ANCHOR2 | AlphaFold | MobiDB | PhosphoSitePlus | ClinVar | gnomAD | ESM1b | AlphaMissense | FoldX | Rosetta | Foldetta | PremPS | REVEL | PROVEAN | PolyPhen-2 HumDiv | PolyPhen-2 HumVar | FATHMM | SIFT | PSMutPred | PAM | Physical | SASA | Normalized B-factor backbone | Normalized B-factor sidechain | SynGAP Structural Annotation | |||||||||||||||||||||||||||||||||||||||||||||
| Score | Prediction | Score | Prediction | pLDDT | disorder | disorder | LTP | HTP | KL | PTM | Clinical Status | Review | Subm. | ID | Allele count | Allele freq. | LLR score | Prediction | Pathogenicity | Class | Optimized | Average ΔΔG | Prediction | StdDev | ΔΔG | Prediction | ΔΔG | Prediction | ΔΔG | Prediction | Score | Prediction | Score | Prediction | pph2_prob | Prediction | pph2_prob | Prediction | Nervous System Score | Prediction | Prediction | Status | Conservation | Sequences | IP RF | SP RF | Prediction | PAM250 | PAM120 | Hydropathy Δ | MW Δ | Average | Δ | Δ | StdDev | Δ | StdDev | Secondary | Tertiary bonds | Inside out | GAP-Ras interface | At membrane | No effect | MD Alert | Verdict | Description | |||||
| c.1862G>A | R621Q 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant R621Q is listed in ClinVar (ID 578137.0) as benign and is present in gnomAD (variant ID 6‑33440914‑G‑A). Functional prediction tools that agree on a benign effect include only FATHMM, whereas the remaining tools—REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default—consistently predict a pathogenic impact. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. No evidence from FoldX, Rosetta, or Foldetta supports a benign outcome. Overall, the preponderance of predictions indicates a likely pathogenic effect, which contradicts the benign classification reported in ClinVar. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.222385 | Structured | 0.084420 | Uncertain | 0.945 | 0.216 | 0.000 | Likely Benign | 1 | 6-33440914-G-A | 19 | 1.18e-5 | -14.682 | Likely Pathogenic | 0.910 | Likely Pathogenic | Ambiguous | 0.81 | Ambiguous | 0.1 | 1.13 | Ambiguous | 0.97 | Ambiguous | 1.35 | Destabilizing | 0.621 | Likely Pathogenic | -3.98 | Deleterious | 1.000 | Probably Damaging | 0.997 | Probably Damaging | 2.82 | Benign | 0.01 | Affected | 3.37 | 35 | 0.2590 | 0.1963 | 1 | 1 | 1.0 | -28.06 | 243.7 | 54.3 | 0.0 | 0.0 | -0.4 | 0.2 | X | X | Potentially Pathogenic | The guanidinium group of Arg621, located in an α helix (res. Glu617-Asn635), forms a salt bridge with Glu525 in a nearby loop and stacks with Leu635. In the variant simulations, the carboxamide side chain of Gln621, which can act as both a hydrogen bond acceptor and donor, also stacks with Leu635 but can only sporadically hydrogen bond with Glu525.Accordingly, the residue swap could affect the tertiary structure integrity by disrupting the salt bridge formation. Additionally, due to its location at the GAP-Ras interface, the residue swap could impact the complex formation with the GTPase, but this cannot be investigated using solvent-only simulations. | ||||||||||||
| c.1891C>A | Q631K 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant Q631K is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions come from FoldX, Foldetta, and FATHMM, while pathogenic predictions arise from REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and the SGM‑Consensus score. Uncertain or inconclusive results are reported by Rosetta, premPS, and AlphaMissense‑Optimized. High‑accuracy methods give a mixed picture: Foldetta, which integrates FoldX‑MD and Rosetta outputs, predicts a benign effect; the SGM‑Consensus, a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, indicates a likely pathogenic outcome; AlphaMissense‑Optimized remains uncertain. Overall, the majority of evidence points toward a pathogenic impact, and this assessment does not conflict with the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.041405 | Structured | 0.038963 | Uncertain | 0.948 | 0.230 | 0.000 | -15.194 | Likely Pathogenic | 0.953 | Likely Pathogenic | Ambiguous | -0.37 | Likely Benign | 0.1 | 1.13 | Ambiguous | 0.38 | Likely Benign | 0.88 | Ambiguous | 0.596 | Likely Pathogenic | -3.98 | Deleterious | 0.958 | Probably Damaging | 0.931 | Probably Damaging | 2.79 | Benign | 0.01 | Affected | 0.1288 | 0.2157 | 1 | 1 | -0.4 | 0.04 | |||||||||||||||||||||||||||||
| c.1990T>G | L664V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L664V is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. Tools that agree on a pathogenic effect include SGM‑Consensus, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. Rosetta and Foldetta are uncertain, providing no definitive evidence. High‑accuracy assessments show AlphaMissense‑Optimized predicts benign, while the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts pathogenic (3 pathogenic vs 1 benign). Foldetta’s stability prediction is uncertain. Overall, the majority of reliable tools predict a pathogenic impact. Therefore, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.100716 | Structured | 0.089318 | Uncertain | 0.937 | 0.339 | 0.000 | -11.515 | Likely Pathogenic | 0.755 | Likely Pathogenic | Likely Benign | 2.01 | Destabilizing | 0.1 | 1.13 | Ambiguous | 1.57 | Ambiguous | 1.37 | Destabilizing | 0.378 | Likely Benign | -2.99 | Deleterious | 0.993 | Probably Damaging | 0.776 | Possibly Damaging | 2.91 | Benign | 0.01 | Affected | 0.0984 | 0.2628 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.723A>C | K241N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K241N is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that agree on a benign effect include only FATHMM, whereas the remaining tools—REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN)—all predict a pathogenic or likely pathogenic impact. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is inconclusive. Based on the collective predictions, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.196879 | Structured | 0.349250 | Uncertain | 0.797 | 0.347 | 0.000 | -10.198 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 0.92 | Ambiguous | 0.1 | 1.13 | Ambiguous | 1.03 | Ambiguous | 0.89 | Ambiguous | 0.663 | Likely Pathogenic | -4.23 | Deleterious | 0.995 | Probably Damaging | 0.829 | Possibly Damaging | 5.81 | Benign | 0.03 | Affected | 0.3426 | 0.1872 | 1 | 0 | 0.4 | -14.07 | |||||||||||||||||||||||||||||
| c.723A>T | K241N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K241N is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only FATHMM, whereas the majority of tools predict a pathogenic impact: REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Uncertain results are reported for FoldX, Rosetta, Foldetta, and premPS and are not used as evidence. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta as uncertain. Based on the overall consensus of the available predictions, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.196879 | Structured | 0.349250 | Uncertain | 0.797 | 0.347 | 0.000 | -10.198 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 0.92 | Ambiguous | 0.1 | 1.13 | Ambiguous | 1.03 | Ambiguous | 0.89 | Ambiguous | 0.663 | Likely Pathogenic | -4.23 | Deleterious | 0.995 | Probably Damaging | 0.829 | Possibly Damaging | 5.81 | Benign | 0.03 | Affected | 0.3426 | 0.1872 | 1 | 0 | 0.4 | -14.07 | |||||||||||||||||||||||||||||
| c.757A>T | N253Y 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N253Y is not reported in ClinVar and has no entries in gnomAD. Prediction tools that indicate a benign effect include FoldX, FATHMM, and premPS, whereas a larger set—SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default—consistently predict pathogenicity. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta as uncertain. Because the majority of evidence points to a deleterious impact, the variant is most likely pathogenic; this conclusion does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.513880 | Disordered | 0.201744 | Uncertain | 0.771 | 0.298 | 0.250 | -14.749 | Likely Pathogenic | 0.920 | Likely Pathogenic | Ambiguous | 0.27 | Likely Benign | 0.1 | 1.13 | Ambiguous | 0.70 | Ambiguous | 0.29 | Likely Benign | 0.896 | Likely Pathogenic | -7.01 | Deleterious | 0.998 | Probably Damaging | 0.994 | Probably Damaging | 5.55 | Benign | 0.01 | Affected | 0.0642 | 0.7055 | -2 | -2 | 2.2 | 49.07 | |||||||||||||||||||||||||||||
| c.1114G>T | G372W 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G372W has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include premPS, PROVEAN, polyPhen‑2 HumVar, and AlphaMissense‑Optimized. Those that agree on a pathogenic effect are REVEL, FoldX, polyPhen‑2 HumDiv, SIFT, ESM1b, and FATHMM. The remaining tools (Rosetta, Foldetta, AlphaMissense‑Default) give uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points to a pathogenic impact, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.433034 | Structured | 0.430335 | Uncertain | 0.322 | 0.774 | 0.375 | -8.262 | Likely Pathogenic | 0.478 | Ambiguous | Likely Benign | 2.28 | Destabilizing | 0.5 | 1.14 | Ambiguous | 1.71 | Ambiguous | 0.21 | Likely Benign | 0.649 | Likely Pathogenic | -1.25 | Neutral | 0.657 | Possibly Damaging | 0.075 | Benign | -0.74 | Pathogenic | 0.00 | Affected | 0.1057 | 0.4368 | -7 | -2 | -0.5 | 129.16 | ||||||||||||||||||||||||||||||
| c.1465C>T | L489F 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L489F is listed in ClinVar with an uncertain significance (ClinVar ID 522018.0) and is present in the gnomAD database (gnomAD ID 6‑33438497‑C‑T). In silico prediction tools that assess pathogenicity all converge on a deleterious effect: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all report a pathogenic outcome, while no tool predicts a benign effect. High‑accuracy assessments reinforce this consensus: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic; Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is inconclusive. No prediction or folding‑stability result is missing or ambiguous. **Thus, the variant is most likely pathogenic based on the collective predictions, and this does not contradict the ClinVar uncertain status.** Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.191378 | Structured | 0.326126 | Uncertain | 0.949 | 0.234 | 0.125 | Uncertain | 2 | 6-33438497-C-T | 1 | 6.20e-7 | -12.066 | Likely Pathogenic | 0.965 | Likely Pathogenic | Likely Pathogenic | 1.72 | Ambiguous | 0.5 | 1.14 | Ambiguous | 1.43 | Ambiguous | 0.56 | Ambiguous | 0.724 | Likely Pathogenic | -3.76 | Deleterious | 1.000 | Probably Damaging | 0.997 | Probably Damaging | -1.51 | Pathogenic | 0.01 | Affected | 3.37 | 35 | 0.0791 | 0.3729 | 2 | 0 | -1.0 | 34.02 | 246.4 | -17.8 | 0.0 | 0.0 | 0.6 | 0.1 | X | Potentially Benign | The iso-butyl side chain of Leu489, located in the α-helix (res. Leu489-Glu519) within an inter-helix space of four helices (res. Ala461-Phe476, res. Val441-Ser457, and res. Met414-Glu436), packs with hydrophobic residues (e.g., Cys432, Ala448, Lys444, Ala493, Val447, Met468) in the inter-helix space. In the variant simulations, the phenyl ring of the Phe489 side chain can also pack favorably in the hydrophobic region. However, due to the size difference, the aromatic side chain of Phe489 tends to reposition to escape the tight region to accommodate the larger side chain, stacking with Lys444. Although no apparent negative changes are observed during the variant simulation, the size difference between the swapped residues could affect the protein folding process. | |||||||||||||
| c.1760G>A | R587K 2D  3DClick to see structure in 3D Viewer AISynGAP1 R587K is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions from SIFT and AlphaMissense‑Optimized, and pathogenic predictions from REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and FATHMM. The remaining tools (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Default) give uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while the SGM‑Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—indicates likely pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta outputs, remains uncertain. Overall, the majority of evidence points toward a pathogenic effect, which is consistent with the SGM‑Consensus prediction but contradicts the benign calls from SIFT and AlphaMissense‑Optimized. Thus, the variant is most likely pathogenic, and this conclusion aligns with the lack of ClinVar annotation rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.054297 | Structured | 0.077330 | Uncertain | 0.862 | 0.216 | 0.000 | -10.220 | Likely Pathogenic | 0.433 | Ambiguous | Likely Benign | 0.63 | Ambiguous | 0.1 | 1.14 | Ambiguous | 0.89 | Ambiguous | 0.88 | Ambiguous | 0.539 | Likely Pathogenic | -2.55 | Deleterious | 0.967 | Probably Damaging | 0.955 | Probably Damaging | -1.16 | Pathogenic | 0.07 | Tolerated | 0.5313 | 0.3897 | Weaken | 3 | 2 | 0.6 | -28.01 | ||||||||||||||||||||||||||||
| c.2054T>G | L685W 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L685W is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only FATHMM, while the majority of tools (REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus) predict a pathogenic impact. FoldX, Rosetta, and Foldetta provide uncertain results and are not considered evidence for either side. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as likely pathogenic, and Foldetta as uncertain. Taken together, the preponderance of evidence indicates that the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar annotation because no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.175930 | Structured | 0.162061 | Uncertain | 0.913 | 0.280 | 0.000 | -15.885 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 1.53 | Ambiguous | 0.2 | 1.14 | Ambiguous | 1.34 | Ambiguous | 1.14 | Destabilizing | 0.509 | Likely Pathogenic | -5.99 | Deleterious | 1.000 | Probably Damaging | 0.984 | Probably Damaging | 3.23 | Benign | 0.00 | Affected | 0.0700 | 0.2328 | -2 | -2 | -4.7 | 73.05 | |||||||||||||||||||||||||||||
| c.673T>C | S225P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S225P is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split opinion: benign calls come from premPS, polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized, while pathogenic calls arise from REVEL, FoldX, PROVEAN, and ESM1b. Two tools—Rosetta and Foldetta—return uncertain results and are treated as unavailable. High‑accuracy assessments further clarify the picture: AlphaMissense‑Optimized predicts benign, whereas the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive due to a 2‑vs‑2 split, and Foldetta remains unavailable. Overall, the majority of evidence leans toward a benign effect, and this conclusion does not conflict with the lack of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.150080 | Structured | 0.344191 | Uncertain | 0.817 | 0.319 | 0.250 | -8.084 | Likely Pathogenic | 0.156 | Likely Benign | Likely Benign | 2.22 | Destabilizing | 1.8 | 1.14 | Ambiguous | 1.68 | Ambiguous | 0.15 | Likely Benign | 0.503 | Likely Pathogenic | -3.31 | Deleterious | 0.396 | Benign | 0.133 | Benign | 5.91 | Benign | 0.17 | Tolerated | 0.2128 | 0.6946 | 1 | -1 | -0.8 | 10.04 | ||||||||||||||||||||||||||||||
| c.830A>T | K277M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 K277M missense variant is not reported in ClinVar and has no entries in gnomAD. Prediction tools cluster into two groups: benign predictions come from FoldX and premPS, while the majority of tools—SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized—label the change as pathogenic. High‑accuracy methods reinforce this trend: AlphaMissense‑Optimized predicts pathogenic, the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) reports likely pathogenic, and Foldetta, which integrates FoldX‑MD and Rosetta outputs, yields an uncertain result. Overall, the preponderance of evidence points to a pathogenic effect, and this assessment does not conflict with the absence of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.061840 | Structured | 0.321811 | Uncertain | 0.649 | 0.247 | 0.250 | -13.918 | Likely Pathogenic | 0.993 | Likely Pathogenic | Likely Pathogenic | 0.18 | Likely Benign | 0.0 | 1.14 | Ambiguous | 0.66 | Ambiguous | 0.15 | Likely Benign | 0.712 | Likely Pathogenic | -5.52 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.80 | Pathogenic | 0.00 | Affected | 0.0945 | 0.2584 | 0 | -1 | 5.8 | 3.02 | |||||||||||||||||||||||||||||
| c.1103C>T | P368L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P368L is not reported in ClinVar (ClinVar ID None) but is present in gnomAD (ID 6‑33438008‑C‑T). Prediction tools that agree on a benign effect include REVEL, ESM1b, and AlphaMissense‑Optimized. Those that agree on a pathogenic effect are PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM. Predictions that are uncertain or inconclusive are FoldX, Rosetta, premPS, AlphaMissense‑Default, and Foldetta. High‑accuracy assessments give AlphaMissense‑Optimized a benign score, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta outputs, is uncertain. Based on the overall distribution of predictions, the variant is most likely pathogenic. This assessment does not contradict ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.363090 | Structured | 0.439989 | Uncertain | 0.580 | 0.677 | 0.250 | 6-33438008-C-T | 1 | 6.33e-7 | -6.520 | Likely Benign | 0.444 | Ambiguous | Likely Benign | 1.52 | Ambiguous | 0.7 | 1.15 | Ambiguous | 1.34 | Ambiguous | 0.52 | Ambiguous | 0.248 | Likely Benign | -6.61 | Deleterious | 0.991 | Probably Damaging | 0.831 | Possibly Damaging | 1.77 | Pathogenic | 0.00 | Affected | 3.42 | 19 | 0.2336 | 0.7125 | -3 | -3 | 5.4 | 16.04 | |||||||||||||||||||||||||
| c.1192C>G | P398A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P398A is not reported in ClinVar (ClinVar ID None) but is present in gnomAD (ID 6‑33438097‑C‑G). Functional prediction tools that agree on a benign effect include REVEL, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Only PROVEAN predicts a pathogenic outcome. Tools with inconclusive results—FoldX, Rosetta, Foldetta, and premPS—are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as Likely Benign, and Foldetta as Uncertain. Overall, the majority of evidence points to a benign effect for P398A, and this conclusion does not contradict the ClinVar status, which currently has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.436924 | Structured | 0.401041 | Uncertain | 0.891 | 0.525 | 0.250 | 6-33438097-C-G | 2 | 1.24e-6 | -5.321 | Likely Benign | 0.184 | Likely Benign | Likely Benign | 1.80 | Ambiguous | 0.2 | 1.15 | Ambiguous | 1.48 | Ambiguous | 0.92 | Ambiguous | 0.290 | Likely Benign | -5.17 | Deleterious | 0.008 | Benign | 0.005 | Benign | 5.55 | Benign | 0.12 | Tolerated | 3.40 | 16 | 0.3644 | 0.5487 | -1 | 1 | 3.4 | -26.04 | ||||||||||||||||||||||||
| c.1439A>C | E480A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E480A is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only SIFT, whereas a majority of tools (REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default) predict a pathogenic impact. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM Consensus (derived from the four pathogenic‑predicted tools above) as likely pathogenic, and Foldetta as uncertain. Because most evidence points to a deleterious effect, the variant is most likely pathogenic, and this conclusion does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.216401 | Structured | 0.426867 | Uncertain | 0.798 | 0.250 | 0.000 | -13.192 | Likely Pathogenic | 0.931 | Likely Pathogenic | Ambiguous | 0.91 | Ambiguous | 0.1 | 1.15 | Ambiguous | 1.03 | Ambiguous | 0.55 | Ambiguous | 0.694 | Likely Pathogenic | -5.04 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | -1.25 | Pathogenic | 0.09 | Tolerated | 0.3468 | 0.6635 | 0 | -1 | 5.3 | -58.04 | |||||||||||||||||||||||||||||
| c.1657A>G | K553E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 K553E missense variant is not reported in ClinVar (status: None) and has no entries in gnomAD. Prediction tools that agree on a benign effect include only SIFT, whereas the remaining tools (REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized) uniformly predict pathogenicity. FoldX and Rosetta give uncertain results, and Foldetta also reports an uncertain stability change. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points to a deleterious effect. Thus, the variant is most likely pathogenic based on current predictions, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.012270 | Structured | 0.006539 | Uncertain | 0.949 | 0.246 | 0.000 | -17.415 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 1.08 | Ambiguous | 0.3 | 1.15 | Ambiguous | 1.12 | Ambiguous | 1.04 | Destabilizing | 0.828 | Likely Pathogenic | -3.85 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | -1.35 | Pathogenic | 0.12 | Tolerated | 0.2890 | 0.0650 | 0 | 1 | 0.4 | 0.94 | |||||||||||||||||||||||||||||
| c.1723C>G | R575G 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant R575G is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Prediction tools that agree on benign effect include only SIFT, whereas the remaining tools—REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and the SGM Consensus—consistently predict pathogenicity. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Because the majority of evidence points to deleterious impact, the variant is most likely pathogenic; this conclusion does not contradict ClinVar status, which currently has no entry for R575G. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.061840 | Structured | 0.021362 | Uncertain | 0.916 | 0.259 | 0.000 | -13.104 | Likely Pathogenic | 0.914 | Likely Pathogenic | Ambiguous | 2.18 | Destabilizing | 0.0 | 1.15 | Ambiguous | 1.67 | Ambiguous | 1.23 | Destabilizing | 0.772 | Likely Pathogenic | -4.22 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.31 | Pathogenic | 0.13 | Tolerated | 0.2889 | 0.1755 | -3 | -2 | 4.1 | -99.14 | |||||||||||||||||||||||||||||
| c.1769G>T | S590I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S590I is not reported in ClinVar and is absent from gnomAD. Prediction tools that classify it as benign include FoldX, premPS, and FATHMM, whereas the majority of tools predict it to be pathogenic: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Two tools give uncertain results: Rosetta and Foldetta. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is pathogenic. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus as pathogenic, and Foldetta as uncertain. Overall, the preponderance of evidence from multiple independent predictors indicates that S590I is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.022667 | Structured | 0.088943 | Uncertain | 0.918 | 0.199 | 0.000 | -17.956 | Likely Pathogenic | 0.982 | Likely Pathogenic | Likely Pathogenic | -0.04 | Likely Benign | 0.5 | 1.15 | Ambiguous | 0.56 | Ambiguous | 0.31 | Likely Benign | 0.686 | Likely Pathogenic | -5.78 | Deleterious | 0.998 | Probably Damaging | 0.948 | Probably Damaging | 3.05 | Benign | 0.02 | Affected | 0.0975 | 0.5025 | -1 | -2 | 5.3 | 26.08 | |||||||||||||||||||||||||||||
| c.1915T>C | F639L 2D  AIThe SynGAP1 missense variant F639L is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign predictions come from REVEL, polyPhen2_HumVar, and FATHMM, whereas pathogenic predictions are returned by SGM‑Consensus, FoldX, premPS, PROVEAN, polyPhen2_HumDiv, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Rosetta and Foldetta provide uncertain results. High‑accuracy assessments indicate that AlphaMissense‑Optimized predicts pathogenicity, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates likely pathogenic, while Foldetta’s stability analysis is inconclusive. Based on the majority of predictions, the variant is most likely pathogenic, which is consistent with the lack of ClinVar annotation and gnomAD absence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.038042 | Structured | 0.117665 | Uncertain | 0.942 | 0.284 | 0.000 | -11.660 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 2.12 | Destabilizing | 0.7 | 1.15 | Ambiguous | 1.64 | Ambiguous | 1.43 | Destabilizing | 0.499 | Likely Benign | -5.98 | Deleterious | 0.994 | Probably Damaging | 0.380 | Benign | 3.12 | Benign | 0.03 | Affected | 0.2347 | 0.2726 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1917T>A | F639L 2D  AIThe SynGAP1 missense variant F639L is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign predictions come from REVEL, polyPhen2_HumVar, and FATHMM, whereas pathogenic predictions are returned by SGM‑Consensus, FoldX, premPS, PROVEAN, polyPhen2_HumDiv, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Rosetta and Foldetta provide uncertain results. High‑accuracy assessments indicate that AlphaMissense‑Optimized predicts pathogenicity, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates likely pathogenic, while Foldetta’s stability analysis is inconclusive. Based on the majority of predictions, the variant is most likely pathogenic, which is consistent with the lack of ClinVar annotation and gnomAD absence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.038042 | Structured | 0.117665 | Uncertain | 0.942 | 0.284 | 0.000 | -11.660 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 2.12 | Destabilizing | 0.7 | 1.15 | Ambiguous | 1.64 | Ambiguous | 1.43 | Destabilizing | 0.336 | Likely Benign | -5.98 | Deleterious | 0.994 | Probably Damaging | 0.380 | Benign | 3.12 | Benign | 0.03 | Affected | 0.2347 | 0.2726 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1917T>G | F639L 2D  AIThe SynGAP1 missense variant F639L is not reported in ClinVar or gnomAD. Prediction tools that indicate a benign effect include REVEL, polyPhen‑2 HumVar, and FATHMM, whereas a majority of tools predict pathogenicity: SGM‑Consensus, FoldX, premPS, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Uncertain results come from Rosetta and Foldetta. High‑accuracy methods specifically show AlphaMissense‑Optimized as pathogenic, SGM‑Consensus as likely pathogenic, and Foldetta as inconclusive. Overall, the balance of evidence points to a pathogenic effect for F639L, and this assessment does not conflict with any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.038042 | Structured | 0.117665 | Uncertain | 0.942 | 0.284 | 0.000 | -11.660 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 2.12 | Destabilizing | 0.7 | 1.15 | Ambiguous | 1.64 | Ambiguous | 1.43 | Destabilizing | 0.336 | Likely Benign | -5.98 | Deleterious | 0.994 | Probably Damaging | 0.380 | Benign | 3.12 | Benign | 0.03 | Affected | 0.2347 | 0.2726 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1951G>A | E651K 2D  AIThe SynGAP1 E651K missense variant has no ClinVar entry and is not reported in gnomAD. Prediction tools cluster into two groups: benign calls (REVEL, FoldX, premPS, polyPhen‑2 HumVar, SIFT, FATHMM) and pathogenic calls (PROVEAN, polyPhen‑2 HumDiv, ESM1b, AlphaMissense‑Default). Three tools (Rosetta, Foldetta, AlphaMissense‑Optimized) give uncertain or inconclusive results. High‑accuracy assessments further clarify the picture: AlphaMissense‑Optimized is uncertain; the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—labels the variant as likely pathogenic; Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, remains uncertain. Overall, the majority of evidence points toward a pathogenic effect, and this conclusion does not contradict any ClinVar status because none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.088832 | Structured | 0.365409 | Uncertain | 0.955 | 0.340 | 0.000 | -8.714 | Likely Pathogenic | 0.818 | Likely Pathogenic | Ambiguous | 0.11 | Likely Benign | 0.4 | 1.15 | Ambiguous | 0.63 | Ambiguous | 0.08 | Likely Benign | 0.211 | Likely Benign | -2.92 | Deleterious | 0.921 | Possibly Damaging | 0.303 | Benign | 3.39 | Benign | 0.17 | Tolerated | 0.2768 | 0.5803 | 0 | 1 | -0.4 | -0.94 | |||||||||||||||||||||||||||||
| c.2014A>G | T672A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant T672A is listed in ClinVar as Benign (ClinVar ID 2154412.0) and is present in gnomAD (variant ID 6‑33441273‑A‑G). Prediction tools that agree on a benign effect include REVEL, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Only PROVEAN predicts a pathogenic outcome. Uncertain results are reported for FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as Benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as Likely Benign, and Foldetta as Uncertain. Overall, the preponderance of evidence points to a benign effect, and this conclusion is consistent with the ClinVar designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.116183 | Structured | 0.102069 | Uncertain | 0.586 | 0.362 | 0.000 | Benign | 1 | 6-33441273-A-G | 3 | 1.86e-6 | -6.524 | Likely Benign | 0.109 | Likely Benign | Likely Benign | 0.51 | Ambiguous | 0.3 | 1.15 | Ambiguous | 0.83 | Ambiguous | 0.65 | Ambiguous | 0.046 | Likely Benign | -3.20 | Deleterious | 0.006 | Benign | 0.002 | Benign | 3.44 | Benign | 0.12 | Tolerated | 3.40 | 25 | 0.3687 | 0.4380 | 1 | 0 | 2.5 | -30.03 | 188.5 | 42.5 | -0.1 | 0.3 | 0.2 | 0.0 | X | Potentially Pathogenic | The hydroxyl group of Thr672, located in an entangled α-α loop connecting the two α-helices (res. Ser641-Glu666 and res. Leu685-Val699), is involved in a highly coordinated hydrogen-bonding network between residues from two α-helices (res. Ser641-Glu666 and res. Arg563-Glu578) and from the α-α loop itself, such as Lys566, Glu666, and Asn669. In the variant simulations, Ala672 can only form a hydrogen bond with Lys566 via its backbone carbonyl group. Consequently, it cannot maintain the Lys566-Glu666 salt bridge through hydrogen bonding, leading to a significant disruption of the intricate and stable hydrogen-bond network between the loop and the helices. | |||||||||||||
| c.2053T>G | L685V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L685V is not reported in ClinVar (ClinVar ID = None) and is absent from gnomAD (gnomAD ID = None). Prediction tools that agree on a benign effect include REVEL and FATHMM, whereas a majority of tools predict a pathogenic impact: PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also indicates a likely pathogenic effect. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM Consensus as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. No other folding‑stability methods provide definitive evidence. Overall, the preponderance of pathogenic predictions, including the SGM Consensus, suggests that the variant is most likely pathogenic; this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.175930 | Structured | 0.162061 | Uncertain | 0.913 | 0.280 | 0.000 | -11.418 | Likely Pathogenic | 0.935 | Likely Pathogenic | Ambiguous | 1.87 | Ambiguous | 0.0 | 1.15 | Ambiguous | 1.51 | Ambiguous | 0.97 | Ambiguous | 0.214 | Likely Benign | -2.99 | Deleterious | 0.993 | Probably Damaging | 0.694 | Possibly Damaging | 3.33 | Benign | 0.02 | Affected | 0.1414 | 0.3010 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.707C>G | A236G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A236G is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are REVEL, premPS, and PROVEAN. Four tools (FoldX, Rosetta, Foldetta, and ESM1b) return uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—also favors benign, while Foldetta remains uncertain. Overall, the majority of available predictions support a benign impact, and there is no ClinVar annotation to contradict this assessment. Thus, the variant is most likely benign. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.185198 | Structured | 0.329926 | Uncertain | 0.775 | 0.330 | 0.000 | -7.412 | In-Between | 0.162 | Likely Benign | Likely Benign | 0.92 | Ambiguous | 0.1 | 1.15 | Ambiguous | 1.04 | Ambiguous | 1.09 | Destabilizing | 0.615 | Likely Pathogenic | -3.35 | Deleterious | 0.067 | Benign | 0.028 | Benign | 5.74 | Benign | 0.07 | Tolerated | 0.2200 | 0.3899 | 1 | 0 | -2.2 | -14.03 | ||||||||||||||||||||||||||||||
| c.724T>A | W242R 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant W242R is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that assess evolutionary conservation and protein function uniformly indicate a deleterious effect: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the change as pathogenic. No tool predicts a benign outcome; the only inconclusive results come from FoldX, Rosetta, and Foldetta, which are treated as unavailable evidence. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta remains uncertain. Consequently, the variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.328603 | Structured | 0.352582 | Uncertain | 0.847 | 0.341 | 0.000 | -11.948 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 0.85 | Ambiguous | 0.5 | 1.15 | Ambiguous | 1.00 | Ambiguous | 1.34 | Destabilizing | 0.859 | Likely Pathogenic | -12.71 | Deleterious | 0.995 | Probably Damaging | 0.854 | Possibly Damaging | 1.52 | Pathogenic | 0.00 | Affected | 3.40 | 14 | 0.4206 | 0.0771 | -3 | 2 | -3.6 | -30.03 | |||||||||||||||||||||||||||
| c.724T>C | W242R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant W242R is reported in gnomAD (ID 6‑33435575‑T‑C) but has no ClinVar entry. Across the evaluated predictors, every tool that provides a definitive call classifies the substitution as pathogenic: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No predictor reports a benign effect. FoldX, Rosetta, and Foldetta return uncertain results and are treated as unavailable evidence. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates likely pathogenic, while Foldetta remains inconclusive. Consequently, the variant is most likely pathogenic based on the consensus of pathogenic predictions, and this assessment does not contradict any ClinVar status because no ClinVar claim exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.328603 | Structured | 0.352582 | Uncertain | 0.847 | 0.341 | 0.000 | 6-33435575-T-C | 2 | 1.24e-6 | -11.948 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 0.85 | Ambiguous | 0.5 | 1.15 | Ambiguous | 1.00 | Ambiguous | 1.34 | Destabilizing | 0.858 | Likely Pathogenic | -12.71 | Deleterious | 0.995 | Probably Damaging | 0.854 | Possibly Damaging | 1.52 | Pathogenic | 0.00 | Affected | 3.40 | 14 | 0.4206 | 0.0771 | -3 | 2 | -3.6 | -30.03 | ||||||||||||||||||||||||
| c.823C>A | P275T 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant P275T is reported in gnomAD (ID 6‑33437728‑C‑A) but has no ClinVar entry. Functional prediction tools cluster into two groups: benign predictions come from REVEL, AlphaMissense‑Default, and AlphaMissense‑Optimized; pathogenic predictions arise from FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and FATHMM. The high‑accuracy AlphaMissense‑Optimized score is benign, whereas the SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta outputs, is uncertain and therefore not considered evidence. No other tools provide conclusive results. Overall, the majority of predictions, including the SGM‑Consensus, indicate a pathogenic effect, and this assessment does not contradict any ClinVar classification because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.059222 | Structured | 0.353469 | Uncertain | 0.811 | 0.208 | 0.250 | 6-33437728-C-A | 3 | 1.86e-6 | -8.708 | Likely Pathogenic | 0.309 | Likely Benign | Likely Benign | 2.44 | Destabilizing | 0.3 | 1.15 | Ambiguous | 1.80 | Ambiguous | 0.69 | Ambiguous | 0.425 | Likely Benign | -5.38 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.75 | Pathogenic | 0.01 | Affected | 3.38 | 19 | 0.1808 | 0.4000 | -1 | 0 | 0.9 | 3.99 | ||||||||||||||||||||||||
| c.1379A>C | K460T 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant K460T is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two consensus groups: benign predictions come from REVEL, SIFT, and FATHMM, whereas pathogenic predictions are made by PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Pathogenic). Stability‑based methods (FoldX, Rosetta, premPS, Foldetta) yield uncertain or inconclusive results. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenicity, the SGM‑Consensus majority vote (AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates Likely Pathogenic, while Foldetta’s combined FoldX‑MD/Rosetta analysis remains inconclusive. Overall, the majority of evidence points to a pathogenic impact for K460T, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.155435 | Structured | 0.289547 | Uncertain | 0.938 | 0.150 | 0.125 | -11.187 | Likely Pathogenic | 0.967 | Likely Pathogenic | Likely Pathogenic | 1.23 | Ambiguous | 0.1 | 1.16 | Ambiguous | 1.20 | Ambiguous | 0.77 | Ambiguous | 0.208 | Likely Benign | -5.02 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.35 | Benign | 0.07 | Tolerated | 0.1858 | 0.3848 | 0 | -1 | 3.2 | -27.07 | |||||||||||||||||||||||||||||
| c.2081C>G | A694G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A694G is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv and SIFT. Predictions that are inconclusive or unavailable are FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as likely benign, and Foldetta as uncertain. Overall, the majority of evidence points to a benign impact. The variant’s predicted benign status does not contradict any ClinVar annotation, as none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.127496 | Structured | 0.352199 | Uncertain | 0.938 | 0.269 | 0.000 | -3.420 | Likely Benign | 0.144 | Likely Benign | Likely Benign | 0.86 | Ambiguous | 0.0 | 1.16 | Ambiguous | 1.01 | Ambiguous | 0.86 | Ambiguous | 0.093 | Likely Benign | -1.90 | Neutral | 0.866 | Possibly Damaging | 0.171 | Benign | 3.50 | Benign | 0.05 | Affected | 0.2015 | 0.3387 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.614T>A | I205N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 I205N missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, FATHMM, and AlphaMissense‑Optimized; pathogenic predictions come from premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), and ESM1b. Four tools (FoldX, Rosetta, Foldetta, AlphaMissense‑Default) give uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. No evidence from these tools contradicts the lack of ClinVar annotation. Overall, the majority of predictions (five pathogenic vs. four benign) and the pathogenic SGM Consensus suggest the variant is most likely pathogenic, with no ClinVar status to conflict this assessment. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.264545 | Structured | 0.409933 | Uncertain | 0.821 | 0.414 | 0.125 | -10.928 | Likely Pathogenic | 0.552 | Ambiguous | Likely Benign | 0.66 | Ambiguous | 0.1 | 1.16 | Ambiguous | 0.91 | Ambiguous | 1.57 | Destabilizing | 0.138 | Likely Benign | -3.56 | Deleterious | 0.940 | Possibly Damaging | 0.641 | Possibly Damaging | 4.06 | Benign | 0.07 | Tolerated | 0.0802 | 0.0212 | -2 | -3 | -8.0 | 0.94 | ||||||||||||||||||||||||||||||
| c.1046C>T | P349L 2D  AIThe SynGAP1 missense variant P349L is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are REVEL and AlphaMissense‑Optimized, whereas the majority of tools (SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default) predict a pathogenic impact. High‑accuracy methods give conflicting results: AlphaMissense‑Optimized reports a benign outcome, SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates a likely pathogenic effect, and Foldetta (combining FoldX‑MD and Rosetta outputs) is uncertain. Other stability‑based predictors (FoldX, Rosetta, premPS) are also inconclusive. Overall, the preponderance of evidence from the consensus of multiple in‑silico tools points to a pathogenic effect for P349L. This prediction does not contradict ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.167087 | Structured | 0.348607 | Uncertain | 0.947 | 0.396 | 0.000 | -11.734 | Likely Pathogenic | 0.650 | Likely Pathogenic | Likely Benign | 0.70 | Ambiguous | 0.6 | 1.17 | Ambiguous | 0.94 | Ambiguous | 0.57 | Ambiguous | 0.326 | Likely Benign | -8.04 | Deleterious | 1.000 | Probably Damaging | 0.997 | Probably Damaging | 1.51 | Pathogenic | 0.00 | Affected | 0.2222 | 0.6867 | -3 | -3 | 5.4 | 16.04 | |||||||||||||||||||||||||||||
| c.1196C>A | A399D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A399D is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. Those that agree on a pathogenic effect are REVEL, FoldX, SIFT, ESM1b, and AlphaMissense‑Default. The remaining tools—Rosetta, Foldetta, premPS, and AlphaMissense‑Optimized—return uncertain or inconclusive results. High‑accuracy methods give no definitive verdict: AlphaMissense‑Optimized is uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is a tie and thus unavailable, and Foldetta is uncertain. Overall, the majority of available predictions (five pathogenic vs. four benign) lean toward pathogenicity. Therefore, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.394753 | Structured | 0.407674 | Uncertain | 0.939 | 0.490 | 0.125 | -10.441 | Likely Pathogenic | 0.943 | Likely Pathogenic | Ambiguous | 2.45 | Destabilizing | 0.2 | 1.17 | Ambiguous | 1.81 | Ambiguous | 0.91 | Ambiguous | 0.525 | Likely Pathogenic | -1.84 | Neutral | 0.421 | Benign | 0.054 | Benign | 5.38 | Benign | 0.05 | Affected | 0.1533 | 0.1760 | 0 | -2 | -5.3 | 44.01 | ||||||||||||||||||||||||||||||
| c.1331A>C | K444T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K444T is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect are REVEL and FATHMM. Those that predict a pathogenic effect include FoldX, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools with uncertain or inconclusive results are Foldetta, Rosetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta as uncertain (treated as unavailable for pathogenicity inference). Overall, the majority of reliable predictions indicate a pathogenic impact. This conclusion does not contradict ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.203355 | Structured | 0.262172 | Uncertain | 0.955 | 0.213 | 0.000 | -15.557 | Likely Pathogenic | 0.991 | Likely Pathogenic | Likely Pathogenic | 2.12 | Destabilizing | 0.1 | 1.17 | Ambiguous | 1.65 | Ambiguous | 0.96 | Ambiguous | 0.442 | Likely Benign | -5.73 | Deleterious | 0.999 | Probably Damaging | 1.000 | Probably Damaging | 3.45 | Benign | 0.01 | Affected | 0.1643 | 0.3416 | 0 | -1 | 3.2 | -27.07 | |||||||||||||||||||||||||||||
| c.763G>C | D255H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant D255H is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: benign predictions come from premPS and FATHMM, while pathogenic predictions are made by SGM‑Consensus (Likely Pathogenic), REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Uncertain results are reported by FoldX, Rosetta, and Foldetta. High‑accuracy methods reinforce the pathogenic interpretation: AlphaMissense‑Optimized predicts Pathogenic, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates Likely Pathogenic, and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, remains Uncertain. Overall, the consensus of the available predictions points to a pathogenic effect for D255H, and this conclusion is consistent with the lack of ClinVar annotation (no contradiction). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | 0.501700 | Disordered | 0.219132 | Uncertain | 0.801 | 0.273 | 0.250 | -14.645 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 1.72 | Ambiguous | 0.2 | 1.17 | Ambiguous | 1.45 | Ambiguous | 0.44 | Likely Benign | 0.808 | Likely Pathogenic | -5.93 | Deleterious | 1.000 | Probably Damaging | 0.997 | Probably Damaging | 5.76 | Benign | 0.00 | Affected | 0.1447 | 0.5211 | 1 | -1 | 0.3 | 22.05 | ||||||||||||||||||||||||||||||
| c.883A>G | T295A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 T295A missense variant has no ClinVar entry and is not reported in gnomAD. Prediction tools cluster into benign (REVEL, FoldX, SIFT, AlphaMissense‑Default, AlphaMissense‑Optimized) and pathogenic (PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, FATHMM). Four tools are uncertain (Rosetta, Foldetta, premPS, ESM1b). High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as inconclusive. Overall, the majority of conventional predictors lean toward a benign effect, whereas the SGM Consensus indicates a pathogenic signal. Thus, the variant is most likely benign based on the bulk of evidence, and this assessment does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.401658 | Structured | 0.295548 | Uncertain | 0.881 | 0.288 | 0.125 | -7.276 | In-Between | 0.336 | Likely Benign | Likely Benign | 0.22 | Likely Benign | 0.1 | 1.17 | Ambiguous | 0.70 | Ambiguous | 0.56 | Ambiguous | 0.340 | Likely Benign | -3.51 | Deleterious | 0.997 | Probably Damaging | 0.992 | Probably Damaging | 1.96 | Pathogenic | 0.11 | Tolerated | 0.4623 | 0.4190 | 1 | 0 | 2.5 | -30.03 | ||||||||||||||||||||||||||||||
| c.892C>G | P298A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P298A is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as Likely Benign. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as Likely Benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. No evidence from FoldX or Rosetta alone is conclusive. Based on the preponderance of benign predictions and the lack of pathogenic consensus, the variant is most likely benign, and this assessment does not contradict any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.328603 | Structured | 0.268765 | Uncertain | 0.860 | 0.283 | 0.500 | -6.082 | Likely Benign | 0.074 | Likely Benign | Likely Benign | 1.22 | Ambiguous | 0.1 | 1.17 | Ambiguous | 1.20 | Ambiguous | 0.50 | Likely Benign | 0.191 | Likely Benign | -0.98 | Neutral | 0.885 | Possibly Damaging | 0.589 | Possibly Damaging | 1.94 | Pathogenic | 0.66 | Tolerated | 0.3761 | 0.5787 | 1 | -1 | 3.4 | -26.04 | |||||||||||||||||||||||||||||
| c.1105A>G | T369A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant T369A is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Only FATHMM predicts a pathogenic outcome, while Rosetta and Foldetta are inconclusive. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as Likely Benign. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized is benign, SGM‑Consensus is Likely Benign, and Foldetta remains uncertain. Overall, the preponderance of evidence indicates that T369A is most likely benign, and this conclusion does not contradict any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.468512 | Structured | 0.437011 | Uncertain | 0.417 | 0.707 | 0.500 | -1.957 | Likely Benign | 0.056 | Likely Benign | Likely Benign | 0.09 | Likely Benign | 0.1 | 1.18 | Ambiguous | 0.64 | Ambiguous | 0.26 | Likely Benign | 0.090 | Likely Benign | -1.93 | Neutral | 0.012 | Benign | 0.016 | Benign | 1.72 | Pathogenic | 0.30 | Tolerated | 0.4538 | 0.5053 | 1 | 0 | 2.5 | -30.03 | |||||||||||||||||||||||||||||
| c.1381G>T | A461S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A461S is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, polyPhen‑2 (HumVar), FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized; pathogenic predictions arise from PROVEAN, polyPhen‑2 (HumDiv), SIFT, and ESM1b. Four tools (FoldX, Rosetta, Foldetta, premPS) give uncertain results and are not considered evidence for either side. High‑accuracy assessments show AlphaMissense‑Optimized predicts benign, while the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive due to a 2‑vs‑2 split, and Foldetta’s stability analysis is also unavailable. Overall, the balance of evidence leans toward a benign effect, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.179055 | Structured | 0.292531 | Uncertain | 0.936 | 0.151 | 0.125 | -10.663 | Likely Pathogenic | 0.309 | Likely Benign | Likely Benign | 0.87 | Ambiguous | 0.0 | 1.18 | Ambiguous | 1.03 | Ambiguous | 0.63 | Ambiguous | 0.236 | Likely Benign | -2.74 | Deleterious | 0.600 | Possibly Damaging | 0.289 | Benign | 3.36 | Benign | 0.02 | Affected | 0.2401 | 0.4551 | 1 | 1 | -2.6 | 16.00 | ||||||||||||||||||||||||||||||
| c.1606T>A | L536M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 L536M missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include FoldX, PROVEAN, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are REVEL, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and FATHMM. The remaining tools—Rosetta, Foldetta, premPS, and AlphaMissense‑Default—return uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points toward a pathogenic impact, and this conclusion does not contradict any ClinVar annotation because no ClinVar status exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.137348 | Structured | 0.042188 | Uncertain | 0.931 | 0.341 | 0.000 | -10.124 | Likely Pathogenic | 0.462 | Ambiguous | Likely Benign | 0.37 | Likely Benign | 0.1 | 1.18 | Ambiguous | 0.78 | Ambiguous | 0.93 | Ambiguous | 0.571 | Likely Pathogenic | -1.94 | Neutral | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.32 | Pathogenic | 0.01 | Affected | 0.0937 | 0.3299 | 4 | 2 | -1.9 | 18.03 | ||||||||||||||||||||||||||||||
| c.1738G>T | G580C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G580C is not reported in ClinVar (ClinVar status: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a pathogenic outcome include REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools with uncertain or inconclusive results are Rosetta and premPS. High‑accuracy methods all support a deleterious effect: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. No tool predicts benign. **Based on the consensus of available predictions, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status (which is currently unreported).** Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.104810 | Structured | 0.025952 | Uncertain | 0.853 | 0.236 | 0.000 | -11.608 | Likely Pathogenic | 0.962 | Likely Pathogenic | Likely Pathogenic | 2.94 | Destabilizing | 0.1 | 1.18 | Ambiguous | 2.06 | Destabilizing | 0.60 | Ambiguous | 0.755 | Likely Pathogenic | -8.66 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.18 | Pathogenic | 0.01 | Affected | 0.1422 | 0.2146 | -3 | -3 | 2.9 | 46.09 | |||||||||||||||||||||||||||||
| c.1915T>G | F639V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F639V is not reported in ClinVar and is absent from gnomAD. Prediction tools largely agree on a deleterious effect: FATHMM is the sole benign predictor, while the remaining 12 tools (SGM‑Consensus, REVEL, FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) all predict pathogenicity; Rosetta is uncertain and is therefore not counted. High‑accuracy methods reinforce this view: AlphaMissense‑Optimized reports pathogenic, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. No prediction or stability result is missing or inconclusive. Based on the consensus of these tools, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.038042 | Structured | 0.117665 | Uncertain | 0.942 | 0.284 | 0.000 | -12.910 | Likely Pathogenic | 0.965 | Likely Pathogenic | Likely Pathogenic | 4.17 | Destabilizing | 0.1 | 1.18 | Ambiguous | 2.68 | Destabilizing | 1.47 | Destabilizing | 0.560 | Likely Pathogenic | -6.97 | Deleterious | 0.992 | Probably Damaging | 0.756 | Possibly Damaging | 3.08 | Benign | 0.02 | Affected | 0.2074 | 0.1583 | -1 | -1 | 1.4 | -48.04 | |||||||||||||||||||||||||||||
| c.2089T>A | W697R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant W697R is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL and FATHMM, whereas a majority of tools predict a pathogenic effect: SGM‑Consensus, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, PROVEAN, AlphaMissense‑Default, and premPS. Predictions that are inconclusive or unavailable are FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Overall, the preponderance of evidence from multiple pathogenic‑predicting tools indicates that W697R is most likely pathogenic, and this conclusion does not contradict any ClinVar status because the variant is not yet catalogued there. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.175930 | Structured | 0.400169 | Uncertain | 0.945 | 0.297 | 0.000 | -10.020 | Likely Pathogenic | 0.941 | Likely Pathogenic | Ambiguous | 1.14 | Ambiguous | 0.1 | 1.18 | Ambiguous | 1.16 | Ambiguous | 1.25 | Destabilizing | 0.401 | Likely Benign | -9.50 | Deleterious | 1.000 | Probably Damaging | 0.994 | Probably Damaging | 3.45 | Benign | 0.02 | Affected | 3.46 | 13 | 0.3944 | 0.0612 | 2 | -3 | -3.6 | -30.03 | 254.4 | -41.2 | 0.0 | 0.0 | -0.7 | 0.0 | X | Potentially Benign | The indole ring of Trp697, located on the outer surface of an α-helix (res. Leu685-Val699), is not involved in any long-lasting interactions in the WT simulations. In the variant simulations, the positively charged guanidinium side chain of Arg697 occasionally forms hydrogen bonds with nearby residues, such as Ser722 and Asn719. However, similar to Trp697 in the WT, Arg697 does not form any long-lasting interactions and thus does not induce any negative structural effects in the simulations. | ||||||||||||||||||
| c.2089T>C | W697R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant W697R is listed in ClinVar as Benign (ClinVar ID 703213.0) and is present in the gnomAD database (gnomAD ID 6‑33441348‑T‑C). Functional prediction tools that agree on a benign effect include REVEL and FATHMM, whereas a majority of tools predict a pathogenic impact: premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and the SGM‑Consensus score. Predictions that are inconclusive or unavailable are FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Overall, the preponderance of evidence from multiple pathogenic‑predicting tools suggests that the variant is most likely pathogenic, which contradicts its current ClinVar benign classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.175930 | Structured | 0.400169 | Uncertain | 0.945 | 0.297 | 0.000 | Likely Benign | 1 | 6-33441348-T-C | 1 | 6.20e-7 | -10.020 | Likely Pathogenic | 0.941 | Likely Pathogenic | Ambiguous | 1.14 | Ambiguous | 0.1 | 1.18 | Ambiguous | 1.16 | Ambiguous | 1.25 | Destabilizing | 0.401 | Likely Benign | -9.50 | Deleterious | 1.000 | Probably Damaging | 0.994 | Probably Damaging | 3.45 | Benign | 0.02 | Affected | 3.46 | 13 | 0.3944 | 0.0612 | 2 | -3 | -3.6 | -30.03 | 254.4 | -41.2 | 0.0 | 0.0 | -0.7 | 0.0 | X | Potentially Benign | The indole ring of Trp697, located on the outer surface of an α-helix (res. Leu685-Val699), is not involved in any long-lasting interactions in the WT simulations. In the variant simulations, the positively charged guanidinium side chain of Arg697 occasionally forms hydrogen bonds with nearby residues, such as Ser722 and Asn719. However, similar to Trp697 in the WT, Arg697 does not form any long-lasting interactions and thus does not induce any negative structural effects in the simulations. | |||||||||||||
| c.674C>T | S225L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S225L is reported in gnomAD (6‑33435525‑C‑T) but has no ClinVar entry. In silico predictors that agree on a benign effect include REVEL, FoldX, premPS, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Only PROVEAN predicts a pathogenic outcome. Predictions that are inconclusive are Rosetta and Foldetta. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Benign, and Foldetta as Uncertain. Overall, the majority of evidence points to a benign effect for S225L, and this conclusion does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | PH | 0.150080 | Structured | 0.344191 | Uncertain | 0.817 | 0.319 | 0.250 | 6-33435525-C-T | 1 | 6.20e-7 | -6.644 | Likely Benign | 0.103 | Likely Benign | Likely Benign | 0.29 | Likely Benign | 0.4 | 1.18 | Ambiguous | 0.74 | Ambiguous | 0.18 | Likely Benign | 0.320 | Likely Benign | -3.76 | Deleterious | 0.000 | Benign | 0.001 | Benign | 5.70 | Benign | 0.28 | Tolerated | 3.41 | 13 | 0.1039 | 0.6554 | -2 | -3 | 4.6 | 26.08 | ||||||||||||||||||||||||
| c.1066C>T | R356C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R356C is listed in ClinVar as Benign (ClinVar ID 469145.0) and is present in gnomAD (ID 6‑33437971‑C‑T). Functional prediction tools cluster into two groups: benign predictions from REVEL and AlphaMissense‑Optimized, and pathogenic predictions from PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus score. Uncertain results are reported by FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as inconclusive. Overall, the majority of evidence points to a pathogenic effect, contradicting the ClinVar benign classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.219301 | Structured | 0.395028 | Uncertain | 0.802 | 0.373 | 0.250 | Likely Benign | 1 | 6-33437971-C-T | 5 | 3.10e-6 | -11.827 | Likely Pathogenic | 0.774 | Likely Pathogenic | Likely Benign | 0.76 | Ambiguous | 0.0 | 1.19 | Ambiguous | 0.98 | Ambiguous | 0.84 | Ambiguous | 0.312 | Likely Benign | -7.12 | Deleterious | 1.000 | Probably Damaging | 0.990 | Probably Damaging | 1.67 | Pathogenic | 0.00 | Affected | 3.39 | 22 | 0.3238 | 0.3618 | -4 | -3 | 7.0 | -53.05 | 212.3 | 91.0 | -0.1 | 0.3 | -0.3 | 0.1 | X | Potentially Pathogenic | Arg356 is located in a loop that includes a short helical section and connects two anti-parallel β sheet strands (res. Gly341-Pro349, res. Thr359-Pro364). In the WT simulations, the guanidinium group of Arg356 alternately forms salt bridges with the carboxylate groups of the GAP domain residues, Glu446 and Glu698. Arg356 also forms hydrogen bonds with the hydroxyl group of the GAP domain residue Thr691 and interacts with Met409 at the C2-GAP interface.In the variant simulations, the Cys356 mutation fails to maintain any of the Arg356 interactions and only occasionally forms weak hydrogen bonds with nearby C2 domain residues (e.g., Gln407). Although no negative structural effects are observed during the simulations, Arg356 is located at the C2 and GAP domain interface, making the residue swap potentially detrimental to the tertiary structure assembly. | |||||||||||||
| c.1181A>G | K394R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K394R is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all indicate benign. Only SIFT predicts a pathogenic outcome, while Rosetta and Foldetta are uncertain. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as Likely Benign. High‑accuracy assessments confirm this: AlphaMissense‑Optimized is benign, SGM‑Consensus is Likely Benign, and Foldetta remains uncertain. Overall, the preponderance of evidence supports a benign classification for K394R, and this conclusion does not contradict the absence of a ClinVar report. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.505461 | Disordered | 0.399336 | Uncertain | 0.387 | 0.634 | 0.625 | -4.902 | Likely Benign | 0.097 | Likely Benign | Likely Benign | -0.01 | Likely Benign | 0.1 | 1.19 | Ambiguous | 0.59 | Ambiguous | 0.42 | Likely Benign | 0.335 | Likely Benign | -1.97 | Neutral | 0.141 | Benign | 0.091 | Benign | 5.11 | Benign | 0.03 | Affected | 0.5488 | 0.2075 | Weaken | 3 | 2 | -0.6 | 28.01 | ||||||||||||||||||||||||||||
| c.1387G>A | D463N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D463N missense variant is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, SIFT, and FATHMM. Those that predict a pathogenic impact are SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, ESM1b, and AlphaMissense‑Default. Predictions that are uncertain (Foldetta, AlphaMissense‑Optimized, Rosetta) are treated as unavailable and do not influence the overall assessment. High‑accuracy methods give the following results: AlphaMissense‑Optimized is uncertain; the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates likely pathogenic; Foldetta is uncertain. Overall, the majority of available predictions lean toward a benign effect, and this conclusion does not contradict any ClinVar status (none reported). Thus, the variant is most likely benign based on current computational evidence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.260850 | Structured | 0.305622 | Uncertain | 0.940 | 0.176 | 0.000 | -11.428 | Likely Pathogenic | 0.812 | Likely Pathogenic | Ambiguous | 0.10 | Likely Benign | 0.1 | 1.19 | Ambiguous | 0.65 | Ambiguous | 0.40 | Likely Benign | 0.217 | Likely Benign | -4.37 | Deleterious | 0.880 | Possibly Damaging | 0.430 | Benign | 3.33 | Benign | 0.10 | Tolerated | 0.1152 | 0.5531 | 2 | 1 | 0.0 | -0.98 | |||||||||||||||||||||||||||||
| c.1808T>A | M603K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M603K is not reported in ClinVar and has no entries in gnomAD. Prediction tools largely agree on a deleterious effect: pathogenic calls come from REVEL, SGM Consensus, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized, while only FoldX reports a benign outcome. High‑accuracy assessments reinforce this trend: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates likely pathogenic, and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is inconclusive. No other tools provide a clear benign signal. Consequently, the variant is most likely pathogenic, and this assessment does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.011342 | Structured | 0.197847 | Uncertain | 0.942 | 0.176 | 0.000 | -15.561 | Likely Pathogenic | 0.994 | Likely Pathogenic | Likely Pathogenic | 0.17 | Likely Benign | 0.0 | 1.19 | Ambiguous | 0.68 | Ambiguous | 0.67 | Ambiguous | 0.933 | Likely Pathogenic | -5.64 | Deleterious | 0.923 | Possibly Damaging | 0.922 | Probably Damaging | -1.35 | Pathogenic | 0.00 | Affected | 0.1268 | 0.0488 | 0 | -1 | -5.8 | -3.02 | |||||||||||||||||||||||||||||
| c.1849G>A | E617K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E617K is not reported in ClinVar but is present in gnomAD (6‑33440901‑G‑A). Functional prediction tools cluster into two groups: benign predictions come from FoldX, premPS, and SIFT, while pathogenic predictions arise from SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, and AlphaMissense‑Default. A third set of methods (Foldetta, AlphaMissense‑Optimized, Rosetta) yield uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points to a pathogenic effect for E617K, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.111485 | Structured | 0.155123 | Uncertain | 0.877 | 0.240 | 0.000 | 6-33440901-G-A | 1 | 6.20e-7 | -10.702 | Likely Pathogenic | 0.910 | Likely Pathogenic | Ambiguous | 0.37 | Likely Benign | 0.1 | 1.19 | Ambiguous | 0.78 | Ambiguous | 0.17 | Likely Benign | 0.534 | Likely Pathogenic | -3.32 | Deleterious | 0.997 | Probably Damaging | 0.987 | Probably Damaging | -1.34 | Pathogenic | 0.48 | Tolerated | 3.37 | 35 | 0.1981 | 0.6282 | 1 | 0 | -0.4 | -0.94 | ||||||||||||||||||||||||
| c.2015C>G | T672R 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant T672R is not reported in ClinVar and is absent from gnomAD. Prediction tools that indicate a benign effect include REVEL, FoldX, Foldetta, polyPhen‑2 HumVar, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Uncertain results from Rosetta and premPS are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign, Foldetta as benign, while the SGM Consensus indicates a likely pathogenic outcome. Overall, the majority of individual predictors lean toward pathogenicity, but the high‑accuracy tools provide conflicting benign signals. Therefore, the variant is most likely pathogenic based on the collective predictions, and this assessment does not contradict any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.116183 | Structured | 0.102069 | Uncertain | 0.586 | 0.362 | 0.000 | -12.454 | Likely Pathogenic | 0.626 | Likely Pathogenic | Likely Benign | -0.48 | Likely Benign | 0.4 | 1.19 | Ambiguous | 0.36 | Likely Benign | 0.60 | Ambiguous | 0.084 | Likely Benign | -4.47 | Deleterious | 0.886 | Possibly Damaging | 0.377 | Benign | 3.43 | Benign | 0.02 | Affected | 0.0996 | 0.2919 | -1 | -1 | -3.8 | 55.08 | |||||||||||||||||||||||||||||
| c.610T>A | S204T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S204T is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) all indicate benign or likely benign. Only polyPhen‑2 HumDiv predicts a pathogenic effect. Stability‑based methods (FoldX, Rosetta, Foldetta, premPS) are inconclusive, providing no definitive evidence. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely benign, and Foldetta as uncertain. Taken together, the majority of reliable predictors classify S204T as benign, and this conclusion does not contradict the absence of a ClinVar assertion. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | PH | 0.268042 | Structured | 0.420667 | Uncertain | 0.816 | 0.405 | 0.125 | -4.164 | Likely Benign | 0.167 | Likely Benign | Likely Benign | 0.75 | Ambiguous | 0.1 | 1.19 | Ambiguous | 0.97 | Ambiguous | -0.52 | Ambiguous | 0.104 | Likely Benign | 0.55 | Neutral | 0.462 | Possibly Damaging | 0.173 | Benign | 4.19 | Benign | 1.00 | Tolerated | 0.0857 | 0.5063 | 1 | 1 | 0.1 | 14.03 | |||||||||||||||||||||||||||||
| c.758A>G | N253S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N253S is listed in ClinVar with no submitted interpretation and is present in gnomAD (ID 6‑33435609‑A‑G). Prediction tools that agree on a benign effect include premPS, SIFT, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are REVEL, PROVEAN, polyPhen‑2 HumDiv, and polyPhen‑2 HumVar. Predictions that are uncertain or inconclusive are FoldX, Rosetta, Foldetta, ESM1b, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive, and Foldetta (combining FoldX‑MD and Rosetta outputs) is also inconclusive. Overall, the evidence is mixed, but the single high‑accuracy tool that is available points to a benign effect. Therefore, the variant is most likely benign based on current predictions, and this assessment does not contradict the ClinVar status, which remains unclassified. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.513880 | Disordered | 0.201744 | Uncertain | 0.771 | 0.298 | 0.250 | 6-33435609-A-G | -7.197 | In-Between | 0.541 | Ambiguous | Likely Benign | 0.60 | Ambiguous | 0.1 | 1.19 | Ambiguous | 0.90 | Ambiguous | -0.03 | Likely Benign | 0.716 | Likely Pathogenic | -4.26 | Deleterious | 0.993 | Probably Damaging | 0.956 | Probably Damaging | 5.56 | Benign | 0.09 | Tolerated | 3.39 | 15 | 0.3960 | 0.7764 | 1 | 1 | 2.7 | -27.03 | |||||||||||||||||||||||||||
| c.848A>C | E283A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E283A is reported in gnomAD (ID 6‑33437753‑A‑C) but has no ClinVar entry. Functional prediction tools that agree on a deleterious effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized, all labeling the change as pathogenic. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also classifies the variant as Likely Pathogenic. Predictions that are inconclusive or unavailable—FoldX, Rosetta, Foldetta, and premPS—do not provide evidence for or against pathogenicity. High‑accuracy assessments confirm the deleterious nature: AlphaMissense‑Optimized predicts pathogenic, SGM Consensus indicates Likely Pathogenic, while Foldetta remains uncertain. Taken together, the overwhelming majority of evidence points to a pathogenic effect, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.098513 | Structured | 0.358602 | Uncertain | 0.950 | 0.249 | 0.000 | 6-33437753-A-C | 2 | 1.24e-6 | -12.547 | Likely Pathogenic | 0.991 | Likely Pathogenic | Likely Pathogenic | 1.26 | Ambiguous | 0.1 | 1.19 | Ambiguous | 1.23 | Ambiguous | 0.53 | Ambiguous | 0.529 | Likely Pathogenic | -5.52 | Deleterious | 0.999 | Probably Damaging | 0.995 | Probably Damaging | 1.67 | Pathogenic | 0.01 | Affected | 3.38 | 19 | 0.4104 | 0.5807 | -1 | 0 | 5.3 | -58.04 | ||||||||||||||||||||||||
| c.890A>C | K297T 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant K297T is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect are none, whereas those that predict a pathogenic effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Uncertain predictions come from FoldX, Rosetta, Foldetta, and premPS. High‑accuracy methods specifically show AlphaMissense‑Optimized as pathogenic, SGM Consensus as likely pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points to a pathogenic impact, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.422041 | Structured | 0.272593 | Uncertain | 0.880 | 0.285 | 0.375 | -10.110 | Likely Pathogenic | 0.987 | Likely Pathogenic | Likely Pathogenic | 1.55 | Ambiguous | 0.1 | 1.19 | Ambiguous | 1.37 | Ambiguous | 0.59 | Ambiguous | 0.551 | Likely Pathogenic | -5.28 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 1.62 | Pathogenic | 0.01 | Affected | 0.2294 | 0.4024 | 0 | -1 | 3.2 | -27.07 | |||||||||||||||||||||||||||||
| c.1417G>A | V473I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V473I is listed in ClinVar with an uncertain significance and is present in gnomAD (ID 6‑33438449‑G‑A). Functional prediction tools that agree on benign impact include REVEL, FoldX, premPS, PROVEAN, SIFT, FATHMM, and AlphaMissense‑Optimized. Pathogenic predictions are provided by both polyPhen‑2 HumDiv and HumVar. Predictions that are inconclusive are AlphaMissense‑Default, ESM1b, Foldetta, and Rosetta. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is unavailable due to no majority, and Foldetta is uncertain. Overall, the balance of evidence favors a benign effect for V473I, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.191378 | Structured | 0.362529 | Uncertain | 0.884 | 0.239 | 0.000 | Uncertain | 1 | 6-33438449-G-A | 1 | 6.20e-7 | -7.481 | In-Between | 0.418 | Ambiguous | Likely Benign | -0.12 | Likely Benign | 0.0 | 1.20 | Ambiguous | 0.54 | Ambiguous | -0.06 | Likely Benign | 0.203 | Likely Benign | -0.91 | Neutral | 0.929 | Possibly Damaging | 0.917 | Probably Damaging | 3.74 | Benign | 0.18 | Tolerated | 3.37 | 34 | 0.0568 | 0.3335 | 3 | 4 | 0.3 | 14.03 | |||||||||||||||||||||||
| c.1819C>T | L607F 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L607F is catalogued in gnomAD (6‑33440871‑C‑T) but has no ClinVar entry. Functional prediction tools largely agree on a deleterious effect: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) all report pathogenic or likely pathogenic. Only FoldX predicts a benign outcome, while Rosetta, Foldetta, premPS, and AlphaMissense‑Optimized are uncertain. High‑accuracy assessments show the SGM‑Consensus as likely pathogenic, AlphaMissense‑Optimized as uncertain, and Foldetta as uncertain. Overall, the preponderance of evidence points to a pathogenic effect for L607F, and this conclusion is not contradicted by ClinVar status (none). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.048328 | Structured | 0.194229 | Uncertain | 0.869 | 0.250 | 0.000 | 6-33440871-C-T | 1 | 6.19e-7 | -13.654 | Likely Pathogenic | 0.948 | Likely Pathogenic | Ambiguous | 0.23 | Likely Benign | 0.1 | 1.20 | Ambiguous | 0.72 | Ambiguous | 0.61 | Ambiguous | 0.758 | Likely Pathogenic | -3.98 | Deleterious | 0.998 | Probably Damaging | 0.997 | Probably Damaging | -1.54 | Pathogenic | 0.01 | Affected | 3.37 | 35 | 0.0872 | 0.2816 | 0 | 2 | -1.0 | 34.02 | ||||||||||||||||||||||||
| c.2029A>G | S677G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S677G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the substitution as benign. No tool predicts pathogenicity. Predictions that are inconclusive—FoldX, Rosetta, Foldetta, and premPS—are treated as unavailable. High‑accuracy assessments reinforce the benign interpretation: AlphaMissense‑Optimized is benign; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also indicates benign; Foldetta, which integrates FoldX‑MD and Rosetta outputs, remains uncertain. Overall, the evidence supports a benign classification for S677G, and this conclusion does not contradict the absence of a ClinVar entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.194234 | Structured | 0.115685 | Uncertain | 0.555 | 0.338 | 0.125 | -5.126 | Likely Benign | 0.060 | Likely Benign | Likely Benign | 0.98 | Ambiguous | 0.1 | 1.20 | Ambiguous | 1.09 | Ambiguous | 0.64 | Ambiguous | 0.126 | Likely Benign | -1.38 | Neutral | 0.002 | Benign | 0.001 | Benign | 3.30 | Benign | 0.26 | Tolerated | 0.2885 | 0.5325 | 1 | 0 | 0.4 | -30.03 | |||||||||||||||||||||||||||||
| c.643G>A | G215S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G215S is not reported in ClinVar and is absent from gnomAD. Prediction tools largely agree on a deleterious effect: FATHMM is the sole benign predictor, while the remaining pathogenic predictors (SGM‑Consensus, REVEL, FoldX, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) all indicate pathogenicity. Uncertain results come from Rosetta, Foldetta, and premPS. High‑accuracy methods reinforce this view: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic, and Foldetta remains uncertain. Overall, the variant is most likely pathogenic based on the consensus of predictive tools, and this assessment does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.155435 | Structured | 0.382818 | Uncertain | 0.791 | 0.291 | 0.000 | -9.067 | Likely Pathogenic | 0.992 | Likely Pathogenic | Likely Pathogenic | 2.30 | Destabilizing | 0.3 | 1.20 | Ambiguous | 1.75 | Ambiguous | 0.55 | Ambiguous | 0.864 | Likely Pathogenic | -5.05 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 5.66 | Benign | 0.02 | Affected | 0.2701 | 0.5761 | 1 | 0 | -0.4 | 30.03 | |||||||||||||||||||||||||||||
| c.657T>G | C219W 2D 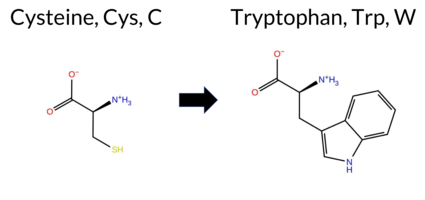 3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C219W is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include only FATHMM, whereas the majority of tools (SGM‑Consensus, REVEL, FoldX, Foldetta, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic impact; Rosetta and premPS are inconclusive. High‑accuracy methods give consistent pathogenic results: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. Taken together, the overwhelming evidence from both general and high‑accuracy predictors indicates that the variant is most likely pathogenic, and this assessment does not contradict any existing ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.254060 | Structured | 0.426845 | Uncertain | 0.903 | 0.279 | 0.000 | -16.565 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 10.10 | Destabilizing | 3.6 | 1.20 | Ambiguous | 5.65 | Destabilizing | 0.52 | Ambiguous | 0.706 | Likely Pathogenic | -9.23 | Deleterious | 0.997 | Probably Damaging | 0.808 | Possibly Damaging | 5.77 | Benign | 0.00 | Affected | 0.1131 | 0.3285 | -8 | -2 | -3.4 | 83.07 | |||||||||||||||||||||||||||||
| c.1298C>G | A433G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A433G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that reach a consensus all indicate a benign effect: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all score the substitution as tolerated. No tool predicts pathogenicity. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized classifies the variant as benign, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) reports it as Likely Benign. The protein‑folding stability method Foldetta, which integrates FoldX‑MD and Rosetta outputs, yields an uncertain result and is therefore treated as unavailable evidence. Overall, the collective predictions strongly suggest that the variant is most likely benign, and this conclusion does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.098513 | Structured | 0.352258 | Uncertain | 0.938 | 0.302 | 0.000 | -5.044 | Likely Benign | 0.107 | Likely Benign | Likely Benign | 0.59 | Ambiguous | 0.0 | 1.21 | Ambiguous | 0.90 | Ambiguous | 0.70 | Ambiguous | 0.038 | Likely Benign | -1.54 | Neutral | 0.035 | Benign | 0.014 | Benign | 3.38 | Benign | 0.15 | Tolerated | 0.1629 | 0.2919 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.1771G>T | A591S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A591S is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool predicts a pathogenic outcome. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized scores the variant as benign, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates a likely benign effect, and the Foldetta stability analysis is inconclusive (unavailable). Consequently, the variant is most likely benign based on the available predictions, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.018787 | Structured | 0.093848 | Uncertain | 0.882 | 0.185 | 0.000 | -7.535 | In-Between | 0.126 | Likely Benign | Likely Benign | 0.58 | Ambiguous | 0.1 | 1.21 | Ambiguous | 0.90 | Ambiguous | 0.49 | Likely Benign | 0.083 | Likely Benign | -2.11 | Neutral | 0.034 | Benign | 0.082 | Benign | 3.52 | Benign | 0.19 | Tolerated | 0.2405 | 0.3304 | 1 | 1 | -2.6 | 16.00 | |||||||||||||||||||||||||||||
| c.2009T>A | L670Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L670Q is not reported in ClinVar and has no entry in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Benign). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and ESM1b. Predictions that are inconclusive (FoldX, Rosetta, Foldetta, premPS) are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus also as Likely Benign, while Foldetta remains uncertain. Overall, the majority of available evidence points to a benign impact. This conclusion is not contradicted by ClinVar, which contains no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.161087 | Structured | 0.090855 | Uncertain | 0.812 | 0.385 | 0.000 | -8.217 | Likely Pathogenic | 0.306 | Likely Benign | Likely Benign | 1.02 | Ambiguous | 0.1 | 1.21 | Ambiguous | 1.12 | Ambiguous | 0.96 | Ambiguous | 0.152 | Likely Benign | -1.12 | Neutral | 1.000 | Probably Damaging | 0.988 | Probably Damaging | 3.40 | Benign | 0.36 | Tolerated | 0.1088 | 0.1303 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.2015C>A | T672K 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant T672K is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, FoldX, polyPhen‑2 HumVar, SIFT, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, ESM1b, and AlphaMissense‑Default. Uncertain predictions come from Foldetta, premPS, and Rosetta. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as inconclusive. Overall, the majority of tools lean toward a benign interpretation, but the high‑accuracy consensus is split, leaving the variant’s impact uncertain. Thus, the variant is most likely benign based on the bulk of predictions, and this does not contradict its ClinVar status of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.116183 | Structured | 0.102069 | Uncertain | 0.586 | 0.362 | 0.000 | Uncertain | 1 | -12.192 | Likely Pathogenic | 0.698 | Likely Pathogenic | Likely Benign | 0.20 | Likely Benign | 0.5 | 1.21 | Ambiguous | 0.71 | Ambiguous | 0.72 | Ambiguous | 0.065 | Likely Benign | -4.31 | Deleterious | 0.745 | Possibly Damaging | 0.051 | Benign | 3.40 | Benign | 0.07 | Tolerated | 3.40 | 25 | 0.1152 | 0.3250 | 0 | -1 | -3.2 | 27.07 | 195.1 | 7.0 | 0.4 | 0.7 | 0.4 | 0.1 | X | X | Potentially Pathogenic | The hydroxyl group of Thr672, located in an entangled α-α loop connecting the two α-helices (res. Ser641-Glu666 and res. Leu685-Val699), is involved in a highly coordinated hydrogen-bonding network between residues from two α-helices (res. Ser641-Glu666 and res. Arg563-Glu578) and from the α-α loop itself, such as Lys566, Glu666, and Asn669. In the variant simulations, Lys672 can only form a hydrogen bond with the amino group of the Lys566 side chain via its backbone carbonyl group. Consequently, it cannot maintain the Lys566-Glu666 salt bridge through hydrogen bonding. However, the amino group of Lys periodically forms a salt bridge with the carboxylate group of Glu666, which prevents a drastic disruption of the hydrogen-bond network that keeps the loop close to the helices. | |||||||||||||||
| c.601G>A | D201N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D201N missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, PROVEAN, SIFT, and FATHMM. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv and polyPhen‑2 HumVar. The remaining tools—Rosetta, Foldetta, ESM1b, and AlphaMissense‑Default—return uncertain results and are treated as unavailable for pathogenicity inference. High‑accuracy assessments show AlphaMissense‑Optimized predicts a benign outcome. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is unavailable because two of the four inputs are uncertain. Foldetta also yields an uncertain result and is unavailable. Overall, the preponderance of evidence points to a benign impact for D201N, and this conclusion does not contradict the lack of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.366687 | Structured | 0.428570 | Uncertain | 0.698 | 0.447 | 0.125 | -7.686 | In-Between | 0.525 | Ambiguous | Likely Benign | 0.31 | Likely Benign | 0.0 | 1.21 | Ambiguous | 0.76 | Ambiguous | -0.10 | Likely Benign | 0.160 | Likely Benign | -2.36 | Neutral | 0.996 | Probably Damaging | 0.877 | Possibly Damaging | 4.08 | Benign | 0.13 | Tolerated | 0.0911 | 0.5413 | 2 | 1 | 0.0 | -0.98 | ||||||||||||||||||||||||||||||
| c.618C>G | I206M 2D  AIThe SynGAP1 I206M missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, PROVEAN, and FATHMM, whereas tools that predict a pathogenic effect are premPS, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as uncertain, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. With six pathogenic predictions versus four benign and three uncertain, the overall evidence leans toward pathogenicity. This conclusion does not contradict ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.298791 | Structured | 0.405123 | Uncertain | 0.863 | 0.391 | 0.125 | -10.008 | Likely Pathogenic | 0.878 | Likely Pathogenic | Ambiguous | 0.42 | Likely Benign | 0.6 | 1.21 | Ambiguous | 0.82 | Ambiguous | 1.08 | Destabilizing | 0.085 | Likely Benign | -2.42 | Neutral | 0.838 | Possibly Damaging | 0.467 | Possibly Damaging | 3.64 | Benign | 0.01 | Affected | 0.0576 | 0.2716 | 2 | 1 | -2.6 | 18.03 | ||||||||||||||||||||||||||||||
| c.619A>G | K207E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K207E is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, SIFT, and FATHMM, whereas a larger group predicts pathogenicity: premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized is pathogenic, the SGM Consensus is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is inconclusive. The remaining stability predictions (FoldX and Rosetta) are uncertain. Overall, the majority of evidence points to a pathogenic effect for K207E, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.374039 | Structured | 0.406823 | Uncertain | 0.847 | 0.359 | 0.125 | -14.387 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 1.39 | Ambiguous | 0.1 | 1.21 | Ambiguous | 1.30 | Ambiguous | 1.09 | Destabilizing | 0.265 | Likely Benign | -3.00 | Deleterious | 0.982 | Probably Damaging | 0.679 | Possibly Damaging | 4.02 | Benign | 0.07 | Tolerated | 0.3504 | 0.1557 | 0 | 1 | 0.4 | 0.94 | |||||||||||||||||||||||||||||
| c.913A>T | T305S 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant T305S is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, FoldX, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic outcome are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. Predictions that are inconclusive or uncertain are provided by Rosetta, Foldetta, and premPS. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized scores the variant as benign, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates “Likely Benign,” and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, yields an uncertain result. Overall, the majority of evidence points to a benign effect, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.359901 | Structured | 0.299706 | Uncertain | 0.872 | 0.274 | 0.125 | -2.899 | Likely Benign | 0.107 | Likely Benign | Likely Benign | 0.45 | Likely Benign | 0.4 | 1.21 | Ambiguous | 0.83 | Ambiguous | 0.55 | Ambiguous | 0.135 | Likely Benign | -0.60 | Neutral | 0.760 | Possibly Damaging | 0.484 | Possibly Damaging | 1.86 | Pathogenic | 0.54 | Tolerated | 0.3579 | 0.4496 | 1 | 1 | -0.1 | -14.03 | |||||||||||||||||||||||||||||
| c.914C>G | T305S 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant T305S is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, FoldX, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic outcome are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. Predictions that are inconclusive or uncertain are provided by Rosetta, Foldetta, and premPS. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized scores the variant as benign, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates “Likely Benign,” and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, yields an uncertain result. Overall, the majority of evidence points to a benign effect, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.359901 | Structured | 0.299706 | Uncertain | 0.872 | 0.274 | 0.125 | -2.899 | Likely Benign | 0.107 | Likely Benign | Likely Benign | 0.45 | Likely Benign | 0.4 | 1.21 | Ambiguous | 0.83 | Ambiguous | 0.55 | Ambiguous | 0.104 | Likely Benign | -0.60 | Neutral | 0.760 | Possibly Damaging | 0.484 | Possibly Damaging | 1.86 | Pathogenic | 0.54 | Tolerated | 0.3579 | 0.4496 | 1 | 1 | -0.1 | -14.03 | |||||||||||||||||||||||||||||
| c.1549C>A | L517M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L517M is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: pathogenic predictions come from REVEL, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus score. Benign predictions are provided by FoldX and PROVEAN. High‑accuracy assessments are mixed: the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates pathogenic, whereas AlphaMissense‑Optimized and Foldetta yield uncertain results and are treated as unavailable. No evidence from ClinVar contradicts these findings. Overall, the preponderance of evidence supports a pathogenic classification for L517M, with no conflict from existing database annotations. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.118441 | Structured | 0.147645 | Uncertain | 0.938 | 0.296 | 0.000 | -11.613 | Likely Pathogenic | 0.799 | Likely Pathogenic | Ambiguous | 0.38 | Likely Benign | 0.2 | 1.22 | Ambiguous | 0.80 | Ambiguous | 0.93 | Ambiguous | 0.665 | Likely Pathogenic | -1.74 | Neutral | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.47 | Pathogenic | 0.01 | Affected | 0.0775 | 0.2558 | 4 | 2 | -1.9 | 18.03 | |||||||||||||||||||||||||||||
| c.1606T>G | L536V 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L536V is listed in ClinVar (ID 1690714.0) with an uncertain significance designation and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from SIFT, AlphaMissense‑Default, and AlphaMissense‑Optimized; pathogenic predictions arise from REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and FATHMM. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a likely pathogenic verdict. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) remains uncertain. No evidence from FoldX or Rosetta alone is available. Overall, the majority of evidence points toward a pathogenic effect, which does not contradict the ClinVar uncertain status but suggests a higher likelihood of pathogenicity. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.137348 | Structured | 0.042188 | Uncertain | 0.931 | 0.341 | 0.000 | Uncertain | 1 | -9.014 | Likely Pathogenic | 0.269 | Likely Benign | Likely Benign | 1.25 | Ambiguous | 0.3 | 1.22 | Ambiguous | 1.24 | Ambiguous | 1.20 | Destabilizing | 0.586 | Likely Pathogenic | -2.81 | Deleterious | 0.998 | Probably Damaging | 0.992 | Probably Damaging | -1.34 | Pathogenic | 0.09 | Tolerated | 3.37 | 34 | 0.1591 | 0.3565 | 2 | 1 | 0.4 | -14.03 | 204.7 | 26.4 | 0.2 | 0.0 | -0.2 | 0.2 | X | Potentially Benign | Leu536 is located on an α-helix (res. Ala533-Val560) at the membrane interface. The iso-butyl group of Leu536 interacts with nearby hydrophobic residues in the preceding loop (e.g., Val526, Pro528, Cys531). In the variant simulations, the iso-propyl side chain of Val536 forms similar hydrophobic interactions as Leu536 in the WT, causing no negative structural effects. | ||||||||||||||||
| c.1685C>A | P562Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P562Q is recorded in gnomAD (variant ID 6‑33440737‑C‑A) but has no ClinVar entry. All available in silico predictors classify the substitution as pathogenic: SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool reports a benign outcome, so the benign group is empty. High‑accuracy assessments reinforce this: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. Consequently, the variant is most likely pathogenic based on the consensus of predictive tools, and this assessment is not contradicted by any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.022306 | Structured | 0.023606 | Uncertain | 0.893 | 0.200 | 0.000 | 6-33440737-C-A | -15.705 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 4.18 | Destabilizing | 0.4 | 1.22 | Ambiguous | 2.70 | Destabilizing | 1.24 | Destabilizing | 0.778 | Likely Pathogenic | -7.96 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 0.57 | Pathogenic | 0.00 | Affected | 3.37 | 35 | 0.1634 | 0.3184 | -1 | 0 | -1.9 | 31.01 | ||||||||||||||||||||||||||
| c.749T>C | V250A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 V250A missense change is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely converge on a pathogenic effect: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and AlphaMissense‑Default all indicate pathogenicity, whereas ESM1b and FATHMM predict a benign outcome. Stability‑based methods (FoldX, Rosetta, Foldetta) and AlphaMissense‑Optimized return uncertain results and are treated as unavailable. High‑accuracy assessments are likewise inconclusive: AlphaMissense‑Optimized is uncertain, the SGM consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is balanced and therefore unavailable, and Foldetta is uncertain. Overall, the preponderance of evidence points to a pathogenic effect for V250A, and this assessment does not contradict any ClinVar annotation, as none exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.447574 | Structured | 0.244075 | Uncertain | 0.778 | 0.324 | 0.125 | -6.385 | Likely Benign | 0.852 | Likely Pathogenic | Ambiguous | 0.82 | Ambiguous | 0.1 | 1.22 | Ambiguous | 1.02 | Ambiguous | 1.48 | Destabilizing | 0.818 | Likely Pathogenic | -3.11 | Deleterious | 0.930 | Possibly Damaging | 0.584 | Possibly Damaging | 5.82 | Benign | 0.02 | Affected | 0.2491 | 0.2151 | 0 | 0 | -2.4 | -28.05 | ||||||||||||||||||||||||||||||
| c.824C>T | P275L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P275L is not reported in ClinVar and is absent from gnomAD. In silico predictions cluster into two groups: benign predictions come from REVEL, premPS, AlphaMissense‑Default, and AlphaMissense‑Optimized; pathogenic predictions arise from PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and FATHMM. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, labels the variant as Likely Pathogenic. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. No evidence from FoldX or Rosetta alone is conclusive. Overall, the majority of tools predict pathogenicity, and the high‑accuracy consensus supports a likely pathogenic classification. This prediction does not contradict any ClinVar status, as the variant is currently unreported in that database. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.059222 | Structured | 0.353469 | Uncertain | 0.811 | 0.208 | 0.250 | -9.785 | Likely Pathogenic | 0.304 | Likely Benign | Likely Benign | 1.63 | Ambiguous | 0.2 | 1.22 | Ambiguous | 1.43 | Ambiguous | 0.29 | Likely Benign | 0.430 | Likely Benign | -6.81 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.83 | Pathogenic | 0.00 | Affected | 0.2139 | 0.5056 | -3 | -3 | 5.4 | 16.04 | |||||||||||||||||||||||||||||
| c.850C>A | L284I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 L284I missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Benign). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. No evidence from FoldX or Rosetta alone is conclusive. Overall, the majority of predictions, including the high‑accuracy methods, support a benign classification. This consensus does not contradict any ClinVar status, as no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.094817 | Structured | 0.371601 | Uncertain | 0.950 | 0.255 | 0.000 | -5.853 | Likely Benign | 0.293 | Likely Benign | Likely Benign | 0.50 | Ambiguous | 0.0 | 1.22 | Ambiguous | 0.86 | Ambiguous | 0.49 | Likely Benign | 0.364 | Likely Benign | -1.51 | Neutral | 0.999 | Probably Damaging | 0.994 | Probably Damaging | 1.88 | Pathogenic | 0.13 | Tolerated | 0.0738 | 0.3006 | 2 | 2 | 0.7 | 0.00 | |||||||||||||||||||||||||||||
| c.920T>C | F307S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F307S is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. In silico predictors that provide a definitive call all classify the variant as pathogenic: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also reports a likely pathogenic verdict. Predictions that are inconclusive or uncertain include FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, SGM‑Consensus is likely pathogenic, while Foldetta (combining FoldX‑MD and Rosetta outputs) remains uncertain. Overall, the preponderance of evidence indicates that the variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.298791 | Structured | 0.327302 | Uncertain | 0.900 | 0.315 | 0.125 | -12.282 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 1.58 | Ambiguous | 0.2 | 1.22 | Ambiguous | 1.40 | Ambiguous | 0.99 | Ambiguous | 0.766 | Likely Pathogenic | -7.32 | Deleterious | 0.999 | Probably Damaging | 0.996 | Probably Damaging | 1.97 | Pathogenic | 0.01 | Affected | 0.4576 | 0.0758 | -3 | -2 | -3.6 | -60.10 | |||||||||||||||||||||||||||||
| c.628C>G | H210D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant H210D is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, and FATHMM. In contrast, the majority of tools predict a pathogenic impact: premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Foldetta, which integrates FoldX‑MD and Rosetta outputs, is uncertain, and Rosetta alone is also uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as Likely Pathogenic, and Foldetta as inconclusive. Overall, the preponderance of evidence points to a pathogenic effect for H210D, and this conclusion does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.144935 | Structured | 0.390904 | Uncertain | 0.872 | 0.298 | 0.125 | -16.440 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 0.13 | Likely Benign | 0.4 | 1.23 | Ambiguous | 0.68 | Ambiguous | 1.23 | Destabilizing | 0.489 | Likely Benign | -7.73 | Deleterious | 0.895 | Possibly Damaging | 0.533 | Possibly Damaging | 3.18 | Benign | 0.00 | Affected | 0.2025 | 0.1255 | 1 | -1 | -0.3 | -22.05 | |||||||||||||||||||||||||||||
| c.755C>G | P252R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P252R is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Computational predictors that classify the variant as benign are FoldX and FATHMM, whereas the majority of tools—REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—predict it to be pathogenic. Predictions that are inconclusive are Rosetta, Foldetta, and premPS. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is likely pathogenic; Foldetta remains uncertain. Overall, the preponderance of evidence indicates that P252R is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.461924 | Structured | 0.211606 | Uncertain | 0.753 | 0.304 | 0.250 | -14.648 | Likely Pathogenic | 0.994 | Likely Pathogenic | Likely Pathogenic | 0.40 | Likely Benign | 0.1 | 1.23 | Ambiguous | 0.82 | Ambiguous | 0.81 | Ambiguous | 0.890 | Likely Pathogenic | -8.27 | Deleterious | 0.997 | Probably Damaging | 0.916 | Probably Damaging | 5.80 | Benign | 0.00 | Affected | 0.1413 | 0.2848 | 0 | -2 | -2.9 | 59.07 | |||||||||||||||||||||||||||||
| c.844T>A | C282S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C282S is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that indicate a benign effect include REVEL, whereas the majority of other in‑silico predictors (premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus) all classify the variant as pathogenic. Stability‑based methods (FoldX, Rosetta, and Foldetta) return uncertain results and are therefore not considered evidence for or against pathogenicity. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. Based on the preponderance of pathogenic predictions, the variant is most likely pathogenic, which does not contradict its current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.098513 | Structured | 0.348535 | Uncertain | 0.942 | 0.250 | 0.000 | Uncertain | 1 | -11.846 | Likely Pathogenic | 0.958 | Likely Pathogenic | Likely Pathogenic | 1.55 | Ambiguous | 0.1 | 1.23 | Ambiguous | 1.39 | Ambiguous | 1.62 | Destabilizing | 0.460 | Likely Benign | -9.19 | Deleterious | 0.997 | Probably Damaging | 0.994 | Probably Damaging | 1.64 | Pathogenic | 0.03 | Affected | 3.39 | 18 | 0.5009 | 0.1286 | Weaken | 0 | -1 | -3.3 | -16.06 | 233.2 | 14.8 | -0.1 | 0.0 | -0.2 | 0.3 | X | Potentially Benign | The thiol-containing side chain of Cys282, located at the beginning of an anti-parallel β sheet strand (res. Arg279-Leu286), packs against multiple hydrophobic residues (e.g., Ile268, Leu284, Trp308, Leu327). In the variant simulations, the hydroxyl-containing side chain of Ser282 is more hydrophilic and, hence, not as favorable as Cys282 for this hydrophobic niche. Due to this polarity difference, the residue swap could potentially weaken the hydrophobic packing of the C2 domain during the folding process.Moreover, because the C2 domain interacts with the membrane, there could also be a negative effect on the stability of the SynGAP-membrane association. However, no large-scale structural changes were observed during the variant simulations. The hydroxyl group of Ser282 forms a hydrogen bond with the backbone carbonyl group of His326 in another β strand (res. Ala322-Arg329), which competes directly with the backbone amide group of Glu283 within the secondary structure element. | |||||||||||||||
| c.845G>C | C282S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C282S is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only REVEL, which scores the variant as benign. All other evaluated algorithms predict a pathogenic impact: premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Predictions from FoldX, Rosetta, and Foldetta are inconclusive and are treated as unavailable. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus is likely pathogenic, while Foldetta remains uncertain. Based on the preponderance of pathogenic predictions and the lack of benign evidence, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.098513 | Structured | 0.348535 | Uncertain | 0.942 | 0.250 | 0.000 | -11.846 | Likely Pathogenic | 0.958 | Likely Pathogenic | Likely Pathogenic | 1.55 | Ambiguous | 0.1 | 1.23 | Ambiguous | 1.39 | Ambiguous | 1.62 | Destabilizing | 0.436 | Likely Benign | -9.19 | Deleterious | 0.997 | Probably Damaging | 0.994 | Probably Damaging | 1.64 | Pathogenic | 0.03 | Affected | 3.39 | 18 | 0.5009 | 0.1286 | Weaken | 0 | -1 | -3.3 | -16.06 | 233.2 | 14.8 | -0.1 | 0.0 | -0.2 | 0.3 | X | Potentially Benign | The thiol-containing side chain of Cys282, located at the beginning of an anti-parallel β sheet strand (res. Arg279-Leu286), packs against multiple hydrophobic residues (e.g., Ile268, Leu284, Trp308, Leu327). In the variant simulations, the hydroxyl-containing side chain of Ser282 is more hydrophilic and, hence, not as favorable as Cys282 for this hydrophobic niche. Due to this polarity difference, the residue swap could potentially weaken the hydrophobic packing of the C2 domain during the folding process.Moreover, because the C2 domain interacts with the membrane, there could also be a negative effect on the stability of the SynGAP-membrane association. However, no large-scale structural changes were observed during the variant simulations. The hydroxyl group of Ser282 forms a hydrogen bond with the backbone carbonyl group of His326 in another β strand (res. Ala322-Arg329), which competes directly with the backbone amide group of Glu283 within the secondary structure element. | |||||||||||||||||
| c.1061C>G | A354G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A354G is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are SIFT and FATHMM. Predictions marked uncertain (Rosetta, Foldetta, premPS) are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely benign, while Foldetta’s stability analysis is inconclusive. Overall, the majority of evidence points to a benign impact for A354G, and this conclusion does not contradict any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.203355 | Structured | 0.381329 | Uncertain | 0.882 | 0.335 | 0.125 | -4.285 | Likely Benign | 0.113 | Likely Benign | Likely Benign | 0.41 | Likely Benign | 0.0 | 1.24 | Ambiguous | 0.83 | Ambiguous | 0.65 | Ambiguous | 0.037 | Likely Benign | -1.87 | Neutral | 0.146 | Benign | 0.038 | Benign | 1.75 | Pathogenic | 0.05 | Affected | 0.2563 | 0.4777 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.1522G>A | D508N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D508N missense variant is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions (REVEL, FoldX, premPS, FATHMM, AlphaMissense‑Optimized) and pathogenic predictions (PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b). Three tools remain uncertain (Rosetta, Foldetta, AlphaMissense‑Default). High‑accuracy assessments further clarify the picture: AlphaMissense‑Optimized classifies the change as benign; the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—returns a pathogenic verdict; Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, yields an inconclusive result. Because the consensus of the high‑accuracy methods is split, the overall prediction is ambiguous, but the balance of evidence leans toward pathogenicity. This assessment does not contradict ClinVar, as the variant has no ClinVar entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.019401 | Structured | 0.255890 | Uncertain | 0.890 | 0.228 | 0.000 | -9.909 | Likely Pathogenic | 0.411 | Ambiguous | Likely Benign | 0.11 | Likely Benign | 0.1 | 1.24 | Ambiguous | 0.68 | Ambiguous | -0.12 | Likely Benign | 0.265 | Likely Benign | -4.62 | Deleterious | 0.870 | Possibly Damaging | 0.615 | Possibly Damaging | 3.32 | Benign | 0.04 | Affected | 0.1577 | 0.4599 | 2 | 1 | 0.0 | -0.98 | ||||||||||||||||||||||||||||||
| c.1745A>G | E582G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E582G is not reported in ClinVar (ClinVar ID: None) and has no entry in gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include REVEL, premPS, SIFT, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Default. FoldX, Rosetta, and Foldetta give uncertain or inconclusive results. High‑accuracy methods give mixed signals: AlphaMissense‑Optimized predicts benign, the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts pathogenic, and Foldetta’s stability assessment is uncertain. Overall, six tools predict pathogenicity while five predict benign, and the high‑accuracy consensus is split. Thus, the variant is most likely pathogenic based on the preponderance of evidence, and this assessment does not contradict ClinVar status, which currently has no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.102787 | Structured | 0.033838 | Uncertain | 0.845 | 0.235 | 0.000 | -9.621 | Likely Pathogenic | 0.630 | Likely Pathogenic | Likely Benign | 1.35 | Ambiguous | 0.2 | 1.24 | Ambiguous | 1.30 | Ambiguous | 0.37 | Likely Benign | 0.224 | Likely Benign | -3.95 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.13 | Benign | 0.13 | Tolerated | 0.2835 | 0.3325 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.592C>G | L198V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 L198V variant has no ClinVar record and is not listed in gnomAD. Prediction tools that agree on a benign effect include REVEL, premPS, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is labeled Likely Pathogenic. FoldX, Rosetta, and Foldetta provide uncertain or inconclusive stability results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while the SGM‑Consensus (3 pathogenic vs. 1 benign) indicates pathogenicity; Foldetta remains uncertain. Overall, the majority of evaluated tools predict a pathogenic impact, and this is not contradicted by any ClinVar annotation (none exists). Thus, the variant is most likely pathogenic based on current computational predictions. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | 0.444081 | Structured | 0.431715 | Uncertain | 0.572 | 0.485 | 0.125 | -9.343 | Likely Pathogenic | 0.747 | Likely Pathogenic | Likely Benign | 1.10 | Ambiguous | 0.2 | 1.24 | Ambiguous | 1.17 | Ambiguous | 0.34 | Likely Benign | 0.265 | Likely Benign | -2.54 | Deleterious | 0.990 | Probably Damaging | 0.675 | Possibly Damaging | 3.44 | Benign | 0.00 | Affected | 0.1520 | 0.3017 | 2 | 1 | 0.4 | -14.03 | ||||||||||||||||||||||||||||||
| c.628C>A | H210N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 H210N missense variant is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, FoldX, and FATHMM, while pathogenic predictions are made by premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus score (Likely Pathogenic). Two tools give uncertain results: Foldetta (combining FoldX‑MD and Rosetta outputs) and Rosetta alone. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus as Likely Pathogenic, and Foldetta as inconclusive. Overall, the majority of evidence points toward a pathogenic effect for H210N. This conclusion is not contradicted by ClinVar, which contains no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.144935 | Structured | 0.390904 | Uncertain | 0.872 | 0.298 | 0.125 | -13.699 | Likely Pathogenic | 0.991 | Likely Pathogenic | Likely Pathogenic | 0.19 | Likely Benign | 0.3 | 1.24 | Ambiguous | 0.72 | Ambiguous | 1.12 | Destabilizing | 0.375 | Likely Benign | -6.01 | Deleterious | 0.895 | Possibly Damaging | 0.533 | Possibly Damaging | 3.11 | Benign | 0.00 | Affected | 0.1166 | 0.1839 | 2 | 1 | -0.3 | -23.04 | |||||||||||||||||||||||||||||
| c.706G>T | A236S 2D  AIThe SynGAP1 A236S variant has no ClinVar entry and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include FoldX, PROVEAN, SIFT, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are REVEL, polyPhen‑2 HumDiv, and polyPhen‑2 HumVar. Predictions that are inconclusive or uncertain are Rosetta, Foldetta, premPS, and ESM1b. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is “Likely Benign.” High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus as benign, and Foldetta as uncertain. Overall, the majority of evidence points to a benign impact for A236S, and this conclusion is not contradicted by any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | PH | 0.185198 | Structured | 0.329926 | Uncertain | 0.775 | 0.330 | 0.000 | -7.845 | In-Between | 0.106 | Likely Benign | Likely Benign | 0.40 | Likely Benign | 0.5 | 1.24 | Ambiguous | 0.82 | Ambiguous | 0.65 | Ambiguous | 0.648 | Likely Pathogenic | -2.30 | Neutral | 0.948 | Possibly Damaging | 0.588 | Possibly Damaging | 5.83 | Benign | 0.13 | Tolerated | 0.2641 | 0.5547 | 1 | 1 | -2.6 | 16.00 | |||||||||||||||||||||||||||||
| c.905C>G | S302C 2D  AIThe SynGAP1 missense variant S302C is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include SGM‑Consensus (Likely Benign), REVEL, FoldX, premPS, PROVEAN, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT. Uncertain predictions come from Rosetta, Foldetta, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely benign, and Foldetta as uncertain. Overall, the majority of evidence points to a benign impact for S302C, and this conclusion does not contradict any ClinVar status, as none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.414856 | Structured | 0.263489 | Uncertain | 0.616 | 0.258 | 0.375 | -7.290 | In-Between | 0.105 | Likely Benign | Likely Benign | 0.32 | Likely Benign | 0.5 | 1.24 | Ambiguous | 0.78 | Ambiguous | -0.04 | Likely Benign | 0.070 | Likely Benign | -0.83 | Neutral | 0.833 | Possibly Damaging | 0.455 | Possibly Damaging | 4.05 | Benign | 0.02 | Affected | 0.1221 | 0.6514 | 0 | -1 | 3.3 | 16.06 | |||||||||||||||||||||||||||||
| c.1265A>G | E422G 2D  3DClick to see structure in 3D Viewer AISynGAP1 E422G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split assessment: benign predictions come from REVEL, premPS, and FATHMM, while pathogenic predictions are reported by PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. Uncertain results are provided by FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy analyses indicate that the SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, predicts a likely pathogenic effect, whereas AlphaMissense‑Optimized remains inconclusive and Foldetta shows no definitive stability change. Overall, the majority of evidence points toward a pathogenic impact for E422G, and this conclusion does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.067594 | Structured | 0.426709 | Uncertain | 0.965 | 0.255 | 0.000 | -11.468 | Likely Pathogenic | 0.952 | Likely Pathogenic | Ambiguous | 1.13 | Ambiguous | 0.0 | 1.25 | Ambiguous | 1.19 | Ambiguous | 0.39 | Likely Benign | 0.488 | Likely Benign | -6.38 | Deleterious | 1.000 | Probably Damaging | 0.997 | Probably Damaging | 3.31 | Benign | 0.01 | Affected | 0.2440 | 0.4184 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.1318A>T | N440Y 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N440Y is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, premPS, SIFT, FATHMM, AlphaMissense‑Optimized, and polyPhen2_HumVar. Tools that predict a pathogenic effect are SGM‑Consensus, PROVEAN, polyPhen2_HumDiv, AlphaMissense‑Default, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized classifying the variant as benign, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicating pathogenic, and Foldetta, which integrates FoldX‑MD and Rosetta outputs, yielding an uncertain stability change. Overall, the majority of predictions lean toward a benign interpretation, and this is consistent with the lack of ClinVar annotation. Therefore, the variant is most likely benign, with no contradiction to ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.191378 | Structured | 0.267204 | Uncertain | 0.929 | 0.245 | 0.000 | -10.586 | Likely Pathogenic | 0.674 | Likely Pathogenic | Likely Benign | 0.81 | Ambiguous | 0.1 | 1.25 | Ambiguous | 1.03 | Ambiguous | 0.20 | Likely Benign | 0.135 | Likely Benign | -3.81 | Deleterious | 0.931 | Possibly Damaging | 0.230 | Benign | 3.43 | Benign | 0.07 | Tolerated | 0.0535 | 0.3624 | -2 | -2 | 2.2 | 49.07 | |||||||||||||||||||||||||||||
| c.1322T>G | V441G 2D  3DClick to see structure in 3D Viewer AISynGAP1 variant V441G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into three groups: benign predictions come from REVEL, FoldX, SIFT, FATHMM, and AlphaMissense‑Optimized; pathogenic predictions arise from PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and ESM1b; the remaining tools (Rosetta, Foldetta, premPS, AlphaMissense‑Default) give uncertain results. High‑accuracy assessments further refine the picture: AlphaMissense‑Optimized remains benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) favors pathogenic, and Foldetta is inconclusive. Taken together, the majority of evidence points to a benign effect, but the SGM Consensus and several individual pathogenic predictors introduce uncertainty. Therefore, the variant is most likely benign based on the overall prediction landscape, and this assessment does not contradict the lack of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.161087 | Structured | 0.259875 | Uncertain | 0.918 | 0.249 | 0.000 | -12.380 | Likely Pathogenic | 0.478 | Ambiguous | Likely Benign | 0.18 | Likely Benign | 0.0 | 1.25 | Ambiguous | 0.72 | Ambiguous | 0.95 | Ambiguous | 0.273 | Likely Benign | -5.88 | Deleterious | 0.841 | Possibly Damaging | 0.997 | Probably Damaging | 3.41 | Benign | 0.24 | Tolerated | 0.1664 | 0.1715 | -1 | -3 | -4.6 | -42.08 | ||||||||||||||||||||||||||||||
| c.1819C>A | L607I 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L607I is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: pathogenic predictions come from SGM‑Consensus, REVEL, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default, while benign calls are made by PROVEAN and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Stability predictions from FoldX, Rosetta, and premPS are inconclusive. Overall, the preponderance of evidence points to a pathogenic effect for L607I. This conclusion is not contradicted by ClinVar, which contains no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.048328 | Structured | 0.194229 | Uncertain | 0.869 | 0.250 | 0.000 | -12.061 | Likely Pathogenic | 0.644 | Likely Pathogenic | Likely Benign | 0.63 | Ambiguous | 0.1 | 1.25 | Ambiguous | 0.94 | Ambiguous | 0.82 | Ambiguous | 0.727 | Likely Pathogenic | -1.99 | Neutral | 0.992 | Probably Damaging | 0.997 | Probably Damaging | -1.54 | Pathogenic | 0.01 | Affected | 0.1079 | 0.3767 | 2 | 2 | 0.7 | 0.00 | |||||||||||||||||||||||||||||
| c.1933T>C | F645L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F645L is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM; pathogenic predictions arise from SGM‑Consensus (Likely Pathogenic), PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenicity, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates Likely Pathogenic, while Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is inconclusive. No evidence from FoldX, Rosetta, or premPS is available to alter this view. Overall, the majority of computational evidence points to a pathogenic impact for F645L, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.046336 | Structured | 0.276445 | Uncertain | 0.921 | 0.325 | 0.000 | -10.624 | Likely Pathogenic | 0.987 | Likely Pathogenic | Likely Pathogenic | 0.98 | Ambiguous | 0.1 | 1.25 | Ambiguous | 1.12 | Ambiguous | 0.99 | Ambiguous | 0.264 | Likely Benign | -4.24 | Deleterious | 0.116 | Benign | 0.008 | Benign | 3.44 | Benign | 0.03 | Affected | 0.2214 | 0.3873 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1935T>A | F645L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F645L is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM; pathogenic predictions arise from SGM‑Consensus (Likely Pathogenic), PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenicity, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates Likely Pathogenic, while Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is inconclusive. No evidence from FoldX, Rosetta, or premPS is available to alter this view. Overall, the majority of computational evidence points to a pathogenic impact for F645L, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.046336 | Structured | 0.276445 | Uncertain | 0.921 | 0.325 | 0.000 | -10.624 | Likely Pathogenic | 0.987 | Likely Pathogenic | Likely Pathogenic | 0.98 | Ambiguous | 0.1 | 1.25 | Ambiguous | 1.12 | Ambiguous | 0.99 | Ambiguous | 0.159 | Likely Benign | -4.24 | Deleterious | 0.116 | Benign | 0.008 | Benign | 3.44 | Benign | 0.03 | Affected | 0.2214 | 0.3873 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1935T>G | F645L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F645L is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that assess sequence conservation and structural impact fall into two groups: benign predictions are made by REVEL, polyPhen‑2 (HumDiv and HumVar), and FATHMM; pathogenic predictions are made by SGM‑Consensus (Likely Pathogenic), PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenicity, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates Likely Pathogenic, while Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, yields an uncertain result. No evidence from FoldX‑MD, Rosetta, or premPS is available to alter this conclusion. Overall, the majority of computational evidence points to a pathogenic impact for F645L, and this assessment is consistent with the absence of a ClinVar entry, so there is no contradiction with existing clinical annotations. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.046336 | Structured | 0.276445 | Uncertain | 0.921 | 0.325 | 0.000 | -10.624 | Likely Pathogenic | 0.987 | Likely Pathogenic | Likely Pathogenic | 0.98 | Ambiguous | 0.1 | 1.25 | Ambiguous | 1.12 | Ambiguous | 0.99 | Ambiguous | 0.159 | Likely Benign | -4.24 | Deleterious | 0.116 | Benign | 0.008 | Benign | 3.44 | Benign | 0.03 | Affected | 0.2214 | 0.3873 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.2093A>G | E698G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E698G missense variant is not reported in ClinVar (ClinVar ID None) and has no entries in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, and FATHMM. In contrast, a majority of tools predict a pathogenic impact: SGM‑Consensus, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default all indicate deleteriousness. FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized are inconclusive, providing no definitive evidence. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as Likely Pathogenic (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN), and Foldetta as uncertain. Overall, the preponderance of evidence points to a pathogenic effect for E698G. This prediction aligns with the absence of ClinVar annotation, so there is no contradiction with existing clinical data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.120615 | Structured | 0.417514 | Uncertain | 0.922 | 0.315 | 0.000 | -10.617 | Likely Pathogenic | 0.786 | Likely Pathogenic | Ambiguous | 0.57 | Ambiguous | 0.0 | 1.25 | Ambiguous | 0.91 | Ambiguous | 0.29 | Likely Benign | 0.439 | Likely Benign | -6.37 | Deleterious | 1.000 | Probably Damaging | 0.993 | Probably Damaging | 3.31 | Benign | 0.01 | Affected | 0.2529 | 0.3286 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.2149C>G | L717V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L717V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Benign). Two tools predict a damaging effect: polyPhen‑2 HumDiv and polyPhen‑2 HumVar. Stability‑based methods (FoldX, Rosetta, Foldetta) return uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus as Likely Benign, and Foldetta as inconclusive. Taken together, the majority of evidence points to a benign impact. This conclusion is consistent with the absence of a ClinVar assertion, so there is no contradiction with existing clinical data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.239899 | Structured | 0.429342 | Uncertain | 0.969 | 0.397 | 0.000 | -6.164 | Likely Benign | 0.308 | Likely Benign | Likely Benign | 1.53 | Ambiguous | 0.0 | 1.25 | Ambiguous | 1.39 | Ambiguous | 0.09 | Likely Benign | 0.119 | Likely Benign | -0.25 | Neutral | 0.995 | Probably Damaging | 0.970 | Probably Damaging | 3.38 | Benign | 0.51 | Tolerated | 0.1226 | 0.2489 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.643G>C | G215R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G215R is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only FATHMM, whereas the majority of algorithms (SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic impact. Four methods (FoldX, Rosetta, Foldetta, premPS) return uncertain results and are not considered evidence for either side. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized classifies the variant as pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates likely pathogenic, while Foldetta’s stability prediction is inconclusive. Taken together, the preponderance of evidence points to a pathogenic effect for G215R. This conclusion is consistent with the absence of a ClinVar entry and does not contradict any existing database annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.155435 | Structured | 0.382818 | Uncertain | 0.791 | 0.291 | 0.000 | -12.654 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 1.75 | Ambiguous | 0.1 | 1.25 | Ambiguous | 1.50 | Ambiguous | 0.56 | Ambiguous | 0.823 | Likely Pathogenic | -6.84 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 5.71 | Benign | 0.01 | Affected | 0.1048 | 0.4890 | -3 | -2 | -4.1 | 99.14 | |||||||||||||||||||||||||||||
| c.694G>C | A232P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A232P is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include FoldX, SIFT, and FATHMM. Tools that agree on a pathogenic effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Uncertain or inconclusive results come from Rosetta, premPS, and Foldetta. High‑accuracy assessments show AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates a likely pathogenic outcome, and Foldetta remains uncertain. Overall, the majority of evaluated tools predict a pathogenic impact, and this conclusion does not contradict the ClinVar status, which is currently unreported. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.254060 | Structured | 0.307228 | Uncertain | 0.878 | 0.305 | 0.000 | -12.697 | Likely Pathogenic | 0.977 | Likely Pathogenic | Likely Pathogenic | 0.05 | Likely Benign | 0.8 | 1.25 | Ambiguous | 0.65 | Ambiguous | 0.71 | Ambiguous | 0.748 | Likely Pathogenic | -2.75 | Deleterious | 0.917 | Possibly Damaging | 0.502 | Possibly Damaging | 5.78 | Benign | 0.06 | Tolerated | 0.2163 | 0.5259 | 1 | -1 | -3.4 | 26.04 | |||||||||||||||||||||||||||||
| c.697T>A | C233S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C233S is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. Those that predict a pathogenic effect are REVEL, premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. FoldX, Rosetta, and Foldetta give uncertain results and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also favors pathogenicity (3 pathogenic vs 1 benign). Foldetta remains uncertain. Overall, the majority of evidence points to a pathogenic impact. This conclusion does not contradict ClinVar status, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.239899 | Structured | 0.306787 | Uncertain | 0.868 | 0.322 | 0.000 | -10.862 | Likely Pathogenic | 0.993 | Likely Pathogenic | Likely Pathogenic | 0.61 | Ambiguous | 0.1 | 1.25 | Ambiguous | 0.93 | Ambiguous | 1.50 | Destabilizing | 0.764 | Likely Pathogenic | -8.89 | Deleterious | 0.421 | Benign | 0.080 | Benign | 5.79 | Benign | 0.03 | Affected | 0.4528 | 0.2833 | 0 | -1 | -3.3 | -16.06 | |||||||||||||||||||||||||||||
| c.698G>C | C233S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C233S is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions come from polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM, while pathogenic predictions arise from SGM‑Consensus, REVEL, premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenicity, and the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates “Likely Pathogenic.” Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, yields an uncertain result and is treated as unavailable evidence. Overall, the preponderance of evidence points to a pathogenic impact for C233S, and this conclusion does not contradict any ClinVar annotation, as none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.239899 | Structured | 0.306787 | Uncertain | 0.868 | 0.322 | 0.000 | -10.862 | Likely Pathogenic | 0.993 | Likely Pathogenic | Likely Pathogenic | 0.61 | Ambiguous | 0.1 | 1.25 | Ambiguous | 0.93 | Ambiguous | 1.50 | Destabilizing | 0.830 | Likely Pathogenic | -8.89 | Deleterious | 0.421 | Benign | 0.080 | Benign | 5.79 | Benign | 0.03 | Affected | 0.4528 | 0.2833 | 0 | -1 | -3.3 | -16.06 | |||||||||||||||||||||||||||||
| c.1172G>A | G391D 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant G391D is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include premPS, PROVEAN, SIFT, ESM1b, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are REVEL, FoldX, polyPhen‑2 (HumDiv and HumVar), FATHMM, and AlphaMissense‑Default. Two tools, Rosetta and Foldetta, return uncertain results. High‑accuracy methods give a benign call from AlphaMissense‑Optimized; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive, and Foldetta is also inconclusive. Overall, six tools favor pathogenicity while five favor benignity, with two uncertain. Thus, the variant is most likely pathogenic based on the current computational evidence, and this assessment does not contradict any ClinVar annotation because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.637480 | Disordered | 0.409509 | Uncertain | 0.279 | 0.741 | 0.750 | -4.651 | Likely Benign | 0.674 | Likely Pathogenic | Likely Benign | 2.59 | Destabilizing | 1.1 | 1.26 | Ambiguous | 1.93 | Ambiguous | 0.22 | Likely Benign | 0.562 | Likely Pathogenic | -0.95 | Neutral | 0.999 | Probably Damaging | 0.960 | Probably Damaging | 1.32 | Pathogenic | 0.29 | Tolerated | 0.1900 | 0.1305 | 1 | -1 | -3.1 | 58.04 | ||||||||||||||||||||||||||||||
| c.1279C>G | H427D 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant H427D is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, premPS, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, polyPhen‑2 HumVar, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) all indicate benign or likely benign. Only polyPhen‑2 HumDiv predicts a pathogenic outcome. High‑accuracy assessments reinforce the benign consensus: AlphaMissense‑Optimized scores benign, SGM‑Consensus is likely benign, while Foldetta (combining FoldX‑MD and Rosetta outputs) is inconclusive. No evidence from the available tools suggests pathogenicity, and the absence of a ClinVar classification means there is no contradiction. Therefore, based on the current predictions, the variant is most likely benign. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.081712 | Structured | 0.394261 | Uncertain | 0.962 | 0.287 | 0.000 | -3.684 | Likely Benign | 0.500 | Ambiguous | Likely Benign | 0.76 | Ambiguous | 0.1 | 1.26 | Ambiguous | 1.01 | Ambiguous | 0.10 | Likely Benign | 0.163 | Likely Benign | -1.43 | Neutral | 0.677 | Possibly Damaging | 0.236 | Benign | 3.58 | Benign | 0.12 | Tolerated | 0.2241 | 0.1678 | 1 | -1 | -0.3 | -22.05 | |||||||||||||||||||||||||||||
| c.1492A>G | M498V 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant M498V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign (REVEL, PROVEAN, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) and pathogenic (FoldX, Foldetta, premPS, polyPhen‑2 HumDiv, SIFT, FATHMM). Rosetta is uncertain. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is “Likely Benign.” High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely benign, while Foldetta predicts a pathogenic effect on protein stability. Overall, the majority of predictions lean toward a benign impact, and this is consistent with the lack of ClinVar evidence; there is no contradiction with ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.092881 | Structured | 0.399612 | Uncertain | 0.932 | 0.158 | 0.000 | -3.229 | Likely Benign | 0.317 | Likely Benign | Likely Benign | 2.74 | Destabilizing | 0.1 | 1.26 | Ambiguous | 2.00 | Destabilizing | 1.17 | Destabilizing | 0.483 | Likely Benign | -1.82 | Neutral | 0.752 | Possibly Damaging | 0.279 | Benign | -1.28 | Pathogenic | 0.04 | Affected | 0.2737 | 0.2737 | 2 | 1 | 2.3 | -32.06 | |||||||||||||||||||||||||||||
| c.1579G>C | D527H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant D527H is not reported in ClinVar and is absent from gnomAD. Prediction tools that classify the variant as benign include FoldX and premPS, whereas the majority of tools—REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized—predict it to be pathogenic. Uncertain results come from Rosetta and Foldetta. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is also pathogenic. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, SGM Consensus as pathogenic, and Foldetta as inconclusive. Based on the preponderance of pathogenic predictions and the lack of benign consensus, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.139895 | Structured | 0.021908 | Uncertain | 0.913 | 0.408 | 0.000 | -13.334 | Likely Pathogenic | 0.986 | Likely Pathogenic | Likely Pathogenic | 0.40 | Likely Benign | 1.2 | 1.26 | Ambiguous | 0.83 | Ambiguous | 0.49 | Likely Benign | 0.901 | Likely Pathogenic | -6.80 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | -2.39 | Pathogenic | 0.00 | Affected | 0.1092 | 0.4346 | 1 | -1 | 0.3 | 22.05 | |||||||||||||||||||||||||||||
| c.1589A>T | K530M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 K530M missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only premPS. Tools that agree on a pathogenic effect are REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. Predictions that are uncertain or inconclusive are FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments: AlphaMissense‑Optimized is uncertain; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic; Foldetta (combining FoldX‑MD and Rosetta outputs) is uncertain. Based on the predominance of pathogenic predictions and the SGM Consensus result, the variant is most likely pathogenic. This assessment does not contradict ClinVar status, as no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.308712 | Structured | 0.018455 | Uncertain | 0.891 | 0.409 | 0.000 | -12.235 | Likely Pathogenic | 0.953 | Likely Pathogenic | Ambiguous | 0.51 | Ambiguous | 0.0 | 1.26 | Ambiguous | 0.89 | Ambiguous | 0.24 | Likely Benign | 0.671 | Likely Pathogenic | -5.17 | Deleterious | 0.999 | Probably Damaging | 0.988 | Probably Damaging | -1.69 | Pathogenic | 0.00 | Affected | 0.0745 | 0.3123 | 0 | -1 | 5.8 | 3.02 | |||||||||||||||||||||||||||||
| c.1738G>C | G580R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G580R is not reported in ClinVar and is absent from gnomAD. Prediction tools that assess pathogenicity all converge on a deleterious effect: REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) all predict a pathogenic outcome. No tool predicts a benign effect. Several methods return uncertain results—Rosetta, Foldetta (combining FoldX‑MD and Rosetta outputs), and premPS—so these do not influence the overall assessment. High‑accuracy evaluations reinforce the pathogenic prediction: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus is likely pathogenic, while Foldetta remains uncertain. Based on the collective evidence, the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.104810 | Structured | 0.025952 | Uncertain | 0.853 | 0.236 | 0.000 | -11.459 | Likely Pathogenic | 0.977 | Likely Pathogenic | Likely Pathogenic | 2.20 | Destabilizing | 0.1 | 1.26 | Ambiguous | 1.73 | Ambiguous | 0.81 | Ambiguous | 0.623 | Likely Pathogenic | -7.33 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.26 | Pathogenic | 0.03 | Affected | 0.1009 | 0.3197 | -3 | -2 | -4.1 | 99.14 | |||||||||||||||||||||||||||||
| c.691T>C | F231L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F231L is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. Tools that agree on a pathogenic effect include REVEL, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized; the SGM‑Consensus score is “Likely Pathogenic.” Predictions that are uncertain or inconclusive are FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Overall, the majority of evidence points to a pathogenic impact for F231L, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.366687 | Structured | 0.306467 | Uncertain | 0.895 | 0.300 | 0.000 | -9.482 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 1.07 | Ambiguous | 0.3 | 1.26 | Ambiguous | 1.17 | Ambiguous | 0.85 | Ambiguous | 0.801 | Likely Pathogenic | -5.08 | Deleterious | 0.390 | Benign | 0.142 | Benign | 6.04 | Benign | 0.01 | Affected | 0.2338 | 0.3794 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.693T>A | F231L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F231L is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. Tools that agree on a pathogenic effect include REVEL, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized; the SGM‑Consensus score is “Likely Pathogenic.” Predictions that are uncertain or inconclusive are FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Overall, the majority of evidence points to a pathogenic impact for F231L, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.366687 | Structured | 0.306467 | Uncertain | 0.895 | 0.300 | 0.000 | -9.482 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 1.07 | Ambiguous | 0.3 | 1.26 | Ambiguous | 1.17 | Ambiguous | 0.85 | Ambiguous | 0.662 | Likely Pathogenic | -5.08 | Deleterious | 0.390 | Benign | 0.142 | Benign | 6.04 | Benign | 0.01 | Affected | 0.2338 | 0.3794 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.693T>G | F231L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F231L is not reported in ClinVar and is absent from gnomAD. Prediction tools that classify it as benign include polyPhen‑2 HumDiv, polyPhen‑2 HumVar and FATHMM, whereas REVEL, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default and AlphaMissense‑Optimized all predict a pathogenic effect. The SGM‑Consensus score is labeled Likely Pathogenic, and the remaining tools (FoldX, Rosetta, Foldetta, premPS) are inconclusive. High‑accuracy methods give a pathogenic verdict from AlphaMissense‑Optimized, a Likely Pathogenic result from the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN), and an uncertain outcome from Foldetta. Overall, the majority of evidence points to a pathogenic impact for F231L, and this assessment does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.366687 | Structured | 0.306467 | Uncertain | 0.895 | 0.300 | 0.000 | -9.482 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 1.07 | Ambiguous | 0.3 | 1.26 | Ambiguous | 1.17 | Ambiguous | 0.85 | Ambiguous | 0.661 | Likely Pathogenic | -5.08 | Deleterious | 0.390 | Benign | 0.142 | Benign | 6.04 | Benign | 0.01 | Affected | 0.2338 | 0.3794 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1075A>C | T359P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant T359P is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv and FATHMM. Uncertain or inconclusive results come from FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as benign, and Foldetta as uncertain. Overall, the majority of evidence points to a benign impact. This conclusion is consistent with the lack of ClinVar annotation, so there is no contradiction with existing clinical data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.281712 | Structured | 0.414952 | Uncertain | 0.939 | 0.480 | 0.250 | -6.884 | Likely Benign | 0.319 | Likely Benign | Likely Benign | 0.97 | Ambiguous | 0.1 | 1.27 | Ambiguous | 1.12 | Ambiguous | 0.68 | Ambiguous | 0.248 | Likely Benign | -2.36 | Neutral | 0.627 | Possibly Damaging | 0.091 | Benign | 1.78 | Pathogenic | 0.14 | Tolerated | 0.2352 | 0.5611 | 0 | -1 | -0.9 | -3.99 | |||||||||||||||||||||||||||||
| c.1129A>G | M377V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M377V is not reported in ClinVar (ClinVar ID None) but is present in gnomAD (gnomAD ID 6‑33438034‑A‑G). Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool predicts a pathogenic outcome; the only inconclusive results come from FoldX, Rosetta, and Foldetta, which are treated as unavailable. High‑accuracy assessments confirm the benign prediction: AlphaMissense‑Optimized is benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is benign, while Foldetta remains uncertain. Overall, the variant is most likely benign, and this assessment does not contradict any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.675549 | Disordered | 0.431183 | Uncertain | 0.324 | 0.884 | 0.625 | 6-33438034-A-G | -1.507 | Likely Benign | 0.073 | Likely Benign | Likely Benign | 0.92 | Ambiguous | 0.3 | 1.27 | Ambiguous | 1.10 | Ambiguous | 0.48 | Likely Benign | 0.161 | Likely Benign | -0.31 | Neutral | 0.000 | Benign | 0.000 | Benign | 5.46 | Benign | 0.15 | Tolerated | 4.32 | 12 | 0.4530 | 0.4096 | 1 | 2 | 2.3 | -32.06 | ||||||||||||||||||||||||||
| c.1363C>A | L455M 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L455M is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, and FATHMM, while those that predict a pathogenic outcome are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Default. High‑accuracy assessments are inconclusive: AlphaMissense‑Optimized is uncertain, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is a tie, and Foldetta is uncertain. No folding‑stability method provides a definitive result. Consequently, the computational evidence does not favor either benign or pathogenic classification. The variant’s status is therefore inconclusive, and this lack of consensus does not contradict any ClinVar annotation, as none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.188120 | Structured | 0.310377 | Uncertain | 0.963 | 0.168 | 0.000 | -10.086 | Likely Pathogenic | 0.802 | Likely Pathogenic | Ambiguous | 0.88 | Ambiguous | 0.0 | 1.27 | Ambiguous | 1.08 | Ambiguous | 0.59 | Ambiguous | 0.147 | Likely Benign | -1.64 | Neutral | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.25 | Benign | 0.11 | Tolerated | 0.0606 | 0.3362 | 4 | 2 | -1.9 | 18.03 | ||||||||||||||||||||||||||||||
| c.1393C>A | L465I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 L465I missense variant is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic outcome are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Default. The high‑accuracy AlphaMissense‑Optimized score classifies the variant as benign, whereas the Foldetta stability assessment is uncertain. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is inconclusive due to a 2‑to‑2 split. Overall, the evidence is mixed; the balance of predictions leans toward a benign interpretation, and this does not contradict the lack of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.346032 | Structured | 0.319240 | Uncertain | 0.956 | 0.202 | 0.000 | -9.672 | Likely Pathogenic | 0.770 | Likely Pathogenic | Likely Benign | 1.21 | Ambiguous | 0.1 | 1.27 | Ambiguous | 1.24 | Ambiguous | 0.78 | Ambiguous | 0.258 | Likely Benign | -1.99 | Neutral | 0.998 | Probably Damaging | 0.997 | Probably Damaging | 2.54 | Benign | 0.08 | Tolerated | 0.0967 | 0.3539 | 2 | 2 | 0.7 | 0.00 | ||||||||||||||||||||||||||||||
| c.1471A>T | T491S 2D  AIThe SynGAP1 missense variant T491S is not reported in ClinVar (ClinVar status: none) and is absent from gnomAD (gnomAD ID: none). Prediction tools that agree on a benign effect include only SIFT, whereas the majority of tools predict a pathogenic impact: REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus (which is a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. No evidence from FoldX or Rosetta is available to support a stability change. Overall, the preponderance of evidence points to a pathogenic effect for T491S, and this conclusion does not contradict any ClinVar annotation because no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.064632 | Structured | 0.325158 | Uncertain | 0.929 | 0.188 | 0.125 | -7.273 | In-Between | 0.924 | Likely Pathogenic | Ambiguous | 0.93 | Ambiguous | 0.7 | 1.27 | Ambiguous | 1.10 | Ambiguous | 1.00 | Destabilizing | 0.704 | Likely Pathogenic | -3.90 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | -1.25 | Pathogenic | 0.19 | Tolerated | 0.3119 | 0.2815 | 1 | 1 | -0.1 | -14.03 | |||||||||||||||||||||||||||||
| c.1472C>G | T491S 2D  AIThe SynGAP1 missense variant T491S is not reported in ClinVar (ClinVar status: none) and is absent from gnomAD (gnomAD ID: none). Prediction tools that agree on a benign effect include only SIFT, whereas the majority of tools predict a pathogenic impact: REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus (which is a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. No evidence from FoldX or Rosetta is available to support a stability change. Overall, the preponderance of evidence points to a pathogenic effect for T491S, and this conclusion does not contradict any ClinVar annotation because no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.064632 | Structured | 0.325158 | Uncertain | 0.929 | 0.188 | 0.125 | -7.273 | In-Between | 0.924 | Likely Pathogenic | Ambiguous | 0.93 | Ambiguous | 0.7 | 1.27 | Ambiguous | 1.10 | Ambiguous | 1.00 | Destabilizing | 0.666 | Likely Pathogenic | -3.90 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | -1.25 | Pathogenic | 0.19 | Tolerated | 0.3119 | 0.2815 | 1 | 1 | -0.1 | -14.03 | |||||||||||||||||||||||||||||
| c.1588A>G | K530E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K530E is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include premPS and polyPhen‑2 HumVar, whereas a majority of tools predict a pathogenic impact: REVEL, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus score (Likely Pathogenic). High‑accuracy assessments are mixed: AlphaMissense‑Optimized is uncertain, the SGM‑Consensus remains Likely Pathogenic, and Foldetta is uncertain. Overall, the preponderance of evidence points to a pathogenic effect for K530E. This conclusion is consistent with the lack of a ClinVar entry, so there is no contradiction with existing clinical annotations. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.308712 | Structured | 0.018455 | Uncertain | 0.891 | 0.409 | 0.000 | -14.450 | Likely Pathogenic | 0.951 | Likely Pathogenic | Ambiguous | 0.79 | Ambiguous | 0.2 | 1.27 | Ambiguous | 1.03 | Ambiguous | 0.43 | Likely Benign | 0.581 | Likely Pathogenic | -3.45 | Deleterious | 0.703 | Possibly Damaging | 0.276 | Benign | -1.57 | Pathogenic | 0.00 | Affected | 0.2505 | 0.0810 | 0 | 1 | 0.4 | 0.94 | |||||||||||||||||||||||||||||
| c.2182C>A | P728T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P728T has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include only REVEL, while the majority of tools (PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default) predict a pathogenic impact. Predictions that are inconclusive or uncertain are FoldX, Rosetta, Foldetta, premPS, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Overall, the preponderance of evidence points to a pathogenic effect for P728T, and this conclusion does not contradict any ClinVar status because no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | 0.632174 | Disordered | 0.434760 | Uncertain | 0.725 | 0.567 | 0.625 | -9.605 | Likely Pathogenic | 0.863 | Likely Pathogenic | Ambiguous | 1.06 | Ambiguous | 0.0 | 1.27 | Ambiguous | 1.17 | Ambiguous | 0.62 | Ambiguous | 0.298 | Likely Benign | -6.21 | Deleterious | 0.999 | Probably Damaging | 0.993 | Probably Damaging | 0.67 | Pathogenic | 0.00 | Affected | 0.1843 | 0.3917 | 0 | -1 | 0.9 | 3.99 | ||||||||||||||||||||||||||||||
| c.622C>A | P208T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P208T missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect comprise SGM‑Consensus, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and Foldetta; premPS and Rosetta are uncertain. High‑accuracy methods give a consistent pathogenic signal: AlphaMissense‑Optimized predicts benign, but the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic, and Foldetta also predicts pathogenic. With the majority of tools, including the high‑accuracy ones, indicating pathogenicity, the variant is most likely pathogenic. This assessment does not contradict ClinVar status, as the variant is currently unreported there. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.271506 | Structured | 0.399506 | Uncertain | 0.864 | 0.345 | 0.125 | -9.016 | Likely Pathogenic | 0.727 | Likely Pathogenic | Likely Benign | 3.20 | Destabilizing | 0.5 | 1.27 | Ambiguous | 2.24 | Destabilizing | 0.94 | Ambiguous | 0.270 | Likely Benign | -6.80 | Deleterious | 1.000 | Probably Damaging | 0.994 | Probably Damaging | 3.76 | Benign | 0.01 | Affected | 0.1857 | 0.5197 | 0 | -1 | 0.9 | 3.99 | |||||||||||||||||||||||||||||
| c.863A>C | D288A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D288A missense variant is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, and SIFT. Those that agree on a pathogenic effect are SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, and AlphaMissense‑Default. AlphaMissense‑Optimized, Foldetta, and Rosetta give uncertain results and are treated as unavailable for pathogenicity inference. High‑accuracy assessments show that the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts pathogenicity, AlphaMissense‑Optimized is uncertain, and Foldetta (combining FoldX‑MD and Rosetta outputs) is also uncertain. Overall, seven tools predict pathogenicity while four predict benign, with no conflicting ClinVar evidence. Therefore, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.125101 | Structured | 0.395525 | Uncertain | 0.873 | 0.261 | 0.000 | -13.470 | Likely Pathogenic | 0.908 | Likely Pathogenic | Ambiguous | 0.34 | Likely Benign | 0.1 | 1.27 | Ambiguous | 0.81 | Ambiguous | 0.10 | Likely Benign | 0.451 | Likely Benign | -6.09 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 1.71 | Pathogenic | 0.07 | Tolerated | 0.4060 | 0.5788 | 0 | -2 | 5.3 | -44.01 | |||||||||||||||||||||||||||||
| c.947A>C | N316T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N316T is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, SIFT, and AlphaMissense‑Optimized. Those that agree on a pathogenic effect are FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), and FATHMM. The remaining tools (AlphaMissense‑Default, ESM1b, Foldetta, premPS, Rosetta) give uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized predicts a benign outcome, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive, and Foldetta also yields an uncertain stability change. Overall, the majority of available predictions lean toward pathogenicity, and this assessment does not contradict any ClinVar annotation (none is present). Thus, the variant is most likely pathogenic based on the current computational evidence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.118441 | Structured | 0.385187 | Uncertain | 0.817 | 0.246 | 0.125 | -7.538 | In-Between | 0.550 | Ambiguous | Likely Benign | 2.71 | Destabilizing | 0.2 | 1.27 | Ambiguous | 1.99 | Ambiguous | 0.57 | Ambiguous | 0.214 | Likely Benign | -3.68 | Deleterious | 0.999 | Probably Damaging | 0.995 | Probably Damaging | 1.79 | Pathogenic | 0.08 | Tolerated | 0.1482 | 0.8474 | 0 | 0 | 2.8 | -13.00 | ||||||||||||||||||||||||||||||
| c.973C>G | L325V 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L325V is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are FoldX, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. Rosetta and Foldetta give uncertain results. The high‑accuracy consensus from AlphaMissense‑Optimized is benign, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely benign, and Foldetta remains uncertain. Overall, the majority of evidence points to a benign impact, and this assessment does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.352862 | Structured | 0.424577 | Uncertain | 0.955 | 0.436 | 0.000 | -3.104 | Likely Benign | 0.162 | Likely Benign | Likely Benign | 2.26 | Destabilizing | 0.1 | 1.27 | Ambiguous | 1.77 | Ambiguous | -0.39 | Likely Benign | 0.183 | Likely Benign | 0.16 | Neutral | 0.898 | Possibly Damaging | 0.472 | Possibly Damaging | 1.68 | Pathogenic | 1.00 | Tolerated | 0.1571 | 0.3957 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.1027G>T | V343F 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V343F is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions are made by REVEL and premPS, whereas pathogenic predictions are made by SIFT, polyPhen‑2 (HumDiv and HumVar), PROVEAN, ESM1b, FATHMM, and AlphaMissense‑Default. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, labels the variant as Likely Pathogenic. High‑accuracy assessments are mixed: AlphaMissense‑Optimized is uncertain; Foldetta, which integrates FoldX‑MD and Rosetta outputs, is also uncertain. No evidence from FoldX or Rosetta alone is available. Overall, the majority of predictions support a pathogenic effect, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.291804 | Structured | 0.383911 | Uncertain | 0.882 | 0.497 | 0.250 | -10.709 | Likely Pathogenic | 0.799 | Likely Pathogenic | Ambiguous | 1.51 | Ambiguous | 0.4 | 1.28 | Ambiguous | 1.40 | Ambiguous | 0.23 | Likely Benign | 0.324 | Likely Benign | -3.37 | Deleterious | 0.976 | Probably Damaging | 0.759 | Possibly Damaging | 1.61 | Pathogenic | 0.01 | Affected | 0.0915 | 0.4552 | -1 | -1 | -1.4 | 48.04 | |||||||||||||||||||||||||||||
| c.1057C>A | L353M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L353M has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include SGM‑Consensus (Likely Benign), REVEL, FoldX, PROVEAN, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, SIFT, and FATHMM. Uncertain results come from Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as Benign, SGM‑Consensus as Likely Benign, and Foldetta as Uncertain. Overall, the majority of evidence points to a benign impact. This conclusion is consistent with the absence of a ClinVar classification; there is no contradiction with existing clinical data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.137348 | Structured | 0.373584 | Uncertain | 0.926 | 0.315 | 0.000 | -6.943 | Likely Benign | 0.206 | Likely Benign | Likely Benign | 0.10 | Likely Benign | 0.0 | 1.28 | Ambiguous | 0.69 | Ambiguous | 0.60 | Ambiguous | 0.117 | Likely Benign | -0.47 | Neutral | 0.744 | Possibly Damaging | 0.289 | Benign | 1.33 | Pathogenic | 0.03 | Affected | 0.0847 | 0.4080 | 4 | 2 | -1.9 | 18.03 | |||||||||||||||||||||||||||||
| c.1156G>T | G386W 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G386W is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Computational predictors that classify the change as benign include REVEL, premPS, PROVEAN, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are FoldX, polyPhen‑2 (HumDiv and HumVar), SIFT, and ESM1b. High‑accuracy assessments give a benign verdict from AlphaMissense‑Optimized, a benign consensus from the SGM method (majority of the four contributing tools are benign), and a pathogenic result from Foldetta. Uncertain calls from AlphaMissense‑Default and Rosetta are treated as unavailable. Overall, the majority of evidence points to a benign impact, and this conclusion does not conflict with the lack of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.733139 | Disordered | 0.424156 | Uncertain | 0.334 | 0.898 | 0.750 | -10.389 | Likely Pathogenic | 0.519 | Ambiguous | Likely Benign | 6.07 | Destabilizing | 5.5 | 1.28 | Ambiguous | 3.68 | Destabilizing | -0.23 | Likely Benign | 0.471 | Likely Benign | -0.85 | Neutral | 0.996 | Probably Damaging | 0.920 | Probably Damaging | 3.90 | Benign | 0.00 | Affected | 0.1058 | 0.4676 | -7 | -2 | -0.5 | 129.16 | ||||||||||||||||||||||||||||||
| c.1461C>A | N487K 2D  AIThe SynGAP1 missense variant N487K lies in the GAP domain. ClinVar has no entry for this variant, and it is not reported in gnomAD. Prediction tools that agree on a benign effect are REVEL and FATHMM. The remaining tools—PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—consistently predict a pathogenic impact, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is labeled “Likely Pathogenic.” High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, SGM Consensus as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. No other stability predictions are available. Based on the preponderance of pathogenic predictions, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which is currently absent. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.209395 | Structured | 0.338511 | Uncertain | 0.890 | 0.243 | 0.125 | -13.520 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 1.10 | Ambiguous | 0.9 | 1.28 | Ambiguous | 1.19 | Ambiguous | 0.80 | Ambiguous | 0.489 | Likely Benign | -5.97 | Deleterious | 0.998 | Probably Damaging | 0.994 | Probably Damaging | 2.72 | Benign | 0.01 | Affected | 0.1953 | 0.2721 | 1 | 0 | -0.4 | 14.07 | |||||||||||||||||||||||||||||
| c.1461C>G | N487K 2D  AIThe SynGAP1 missense variant N487K is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely disagree: benign predictions come from REVEL and FATHMM, while pathogenic predictions are made by PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Stability‑related methods (FoldX, Rosetta, premPS, Foldetta) yield uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as likely pathogenic, and Foldetta as unavailable. Overall, the majority of evidence points to a pathogenic effect, and this conclusion does not contradict any ClinVar annotation (none is available). Thus, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.209395 | Structured | 0.338511 | Uncertain | 0.890 | 0.243 | 0.125 | -13.520 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 1.10 | Ambiguous | 0.9 | 1.28 | Ambiguous | 1.19 | Ambiguous | 0.80 | Ambiguous | 0.488 | Likely Benign | -5.97 | Deleterious | 0.998 | Probably Damaging | 0.994 | Probably Damaging | 2.72 | Benign | 0.01 | Affected | 0.1953 | 0.2721 | 1 | 0 | -0.4 | 14.07 | |||||||||||||||||||||||||||||
| c.1582C>T | P528S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P528S missense variant is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: pathogenic calls are made by REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and the SGM‑Consensus score (Likely Pathogenic). Only FATHMM predicts a benign outcome. High‑accuracy assessments are mixed: AlphaMissense‑Optimized is inconclusive, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) remains Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta) is inconclusive. No folding‑stability metrics (FoldX, Rosetta, premPS) provide definitive evidence. Overall, the preponderance of pathogenic predictions suggests the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.225814 | Structured | 0.020396 | Uncertain | 0.909 | 0.403 | 0.000 | -11.729 | Likely Pathogenic | 0.819 | Likely Pathogenic | Ambiguous | 1.88 | Ambiguous | 0.1 | 1.28 | Ambiguous | 1.58 | Ambiguous | 0.80 | Ambiguous | 0.617 | Likely Pathogenic | -7.65 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 2.52 | Benign | 0.01 | Affected | 0.3115 | 0.3982 | 1 | -1 | 0.8 | -10.04 | |||||||||||||||||||||||||||||
| c.1684C>G | P562A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P562A is not reported in ClinVar and is absent from gnomAD. Prediction tools largely agree on a deleterious effect: only premPS classifies it as benign, whereas REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict pathogenicity; Rosetta remains uncertain. High‑accuracy methods reinforce this view: AlphaMissense‑Optimized reports a pathogenic change, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenicity. Consequently, the variant is most likely pathogenic. This assessment does not contradict ClinVar, as no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.022306 | Structured | 0.023606 | Uncertain | 0.893 | 0.200 | 0.000 | -13.554 | Likely Pathogenic | 0.989 | Likely Pathogenic | Likely Pathogenic | 2.87 | Destabilizing | 0.0 | 1.28 | Ambiguous | 2.08 | Destabilizing | 0.39 | Likely Benign | 0.633 | Likely Pathogenic | -7.96 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 0.59 | Pathogenic | 0.01 | Affected | 0.3418 | 0.2883 | 1 | -1 | 3.4 | -26.04 | |||||||||||||||||||||||||||||
| c.1768A>G | S590G 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant S590G is listed in ClinVar (ID 1721675.0) with an uncertain significance status and is present in gnomAD (6‑33440820‑A‑G). Functional prediction tools that report a benign effect include REVEL, SIFT, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, and ESM1b. The high‑accuracy consensus (SGM Consensus) derived from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN yields a pathogenic majority. Foldetta, which integrates FoldX‑MD and Rosetta outputs, is inconclusive, as are FoldX, Rosetta, and premPS. Overall, the majority of evidence points toward a pathogenic impact, which does not contradict the ClinVar uncertain status but suggests a higher likelihood of pathogenicity. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.022667 | Structured | 0.088943 | Uncertain | 0.918 | 0.199 | 0.000 | Conflicting | 2 | 6-33440820-A-G | 14 | 8.67e-6 | -14.277 | Likely Pathogenic | 0.574 | Likely Pathogenic | Likely Benign | 0.67 | Ambiguous | 0.1 | 1.28 | Ambiguous | 0.98 | Ambiguous | 0.71 | Ambiguous | 0.379 | Likely Benign | -3.92 | Deleterious | 1.000 | Probably Damaging | 0.922 | Probably Damaging | 3.42 | Benign | 0.06 | Tolerated | 3.37 | 35 | 0.2627 | 0.4118 | 1 | 0 | 0.4 | -30.03 | 186.7 | 49.4 | 0.0 | 0.0 | 0.1 | 0.0 | X | Potentially Pathogenic | In the WT simulations, the hydroxyl group of Ser590, located on an α helix (res. Glu582-Met603), forms hydrogen bonds with the backbone carbonyl of Ala634 and/or the carboxamide group of the Asn635 side chain at the end of the opposing α helix (res. Thr619-Ala634).The residue swap could weaken the integrity of the α helix, as glycine is known as an “α helix breaker.” However, no discernible difference was observed between the WT and variant simulations in this regard. Importantly, Gly590 cannot form hydrogen bonds with the opposing helix in the same way that serine can, which could weaken the tertiary structure assembly between the two helices. | |||||||||||||
| c.1843C>T | P615S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P615S is not reported in ClinVar (ClinVar status: not reported) and is absent from gnomAD (gnomAD ID: none). Prediction tools that agree on a pathogenic effect include REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized; the SGM Consensus also indicates a likely pathogenic outcome. No tools predict a benign effect. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, while Foldetta’s stability prediction is uncertain and therefore not taken as evidence. Overall, the preponderance of evidence points to the variant being most likely pathogenic, with no contradiction to the ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.236433 | Structured | 0.179032 | Uncertain | 0.879 | 0.255 | 0.000 | -12.566 | Likely Pathogenic | 0.996 | Likely Pathogenic | Likely Pathogenic | 2.46 | Destabilizing | 0.3 | 1.28 | Ambiguous | 1.87 | Ambiguous | 0.98 | Ambiguous | 0.780 | Likely Pathogenic | -7.97 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.19 | Pathogenic | 0.04 | Affected | 0.3191 | 0.3310 | 1 | -1 | 0.8 | -10.04 | |||||||||||||||||||||||||||||
| c.1844C>A | P615Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense change P615Q is not listed in ClinVar and has no allele record in gnomAD. Functional prediction tools that assess sequence conservation and structural impact uniformly classify the variant as pathogenic: REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Only Rosetta and the combined Foldetta stability assessment are inconclusive. Grouping the predictions, the benign category contains no tools, while the pathogenic category includes all the above. High‑accuracy methods reinforce this view: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) reports Likely Pathogenic, and Foldetta remains uncertain. Consequently, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.236433 | Structured | 0.179032 | Uncertain | 0.879 | 0.255 | 0.000 | -13.247 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 2.16 | Destabilizing | 0.3 | 1.28 | Ambiguous | 1.72 | Ambiguous | 1.33 | Destabilizing | 0.742 | Likely Pathogenic | -7.97 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | -1.28 | Pathogenic | 0.01 | Affected | 0.1395 | 0.3445 | 0 | -1 | -1.9 | 31.01 | |||||||||||||||||||||||||||||
| c.1900G>T | A634S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 A634S variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are FATHMM and AlphaMissense‑Optimized; those that agree on a pathogenic effect are REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and ESM1b. The remaining tools (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Default) are inconclusive. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points to a pathogenic impact, and this conclusion does not contradict the ClinVar status, which simply lacks an entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.085092 | Structured | 0.052058 | Uncertain | 0.932 | 0.242 | 0.000 | -9.706 | Likely Pathogenic | 0.434 | Ambiguous | Likely Benign | 0.91 | Ambiguous | 0.1 | 1.28 | Ambiguous | 1.10 | Ambiguous | 0.77 | Ambiguous | 0.506 | Likely Pathogenic | -2.99 | Deleterious | 0.953 | Possibly Damaging | 0.985 | Probably Damaging | 2.67 | Benign | 0.05 | Affected | 0.2607 | 0.4231 | 1 | 1 | -2.6 | 16.00 | ||||||||||||||||||||||||||||||
| c.2078A>G | H693R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant H693R is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect are limited to FATHMM, while the majority of algorithms (SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic impact; FoldX, Rosetta, and Foldetta are inconclusive. High‑accuracy assessments further support a deleterious outcome: AlphaMissense‑Optimized classifies the variant as pathogenic, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates a likely pathogenic effect, and Foldetta’s stability analysis is unavailable. Based on the preponderance of pathogenic predictions and the lack of benign evidence, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar annotation (none is present). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.073402 | Structured | 0.323991 | Uncertain | 0.964 | 0.260 | 0.000 | -14.326 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 1.39 | Ambiguous | 0.2 | 1.28 | Ambiguous | 1.34 | Ambiguous | 1.03 | Destabilizing | 0.593 | Likely Pathogenic | -7.97 | Deleterious | 0.998 | Probably Damaging | 0.646 | Possibly Damaging | 3.13 | Benign | 0.01 | Affected | 0.1839 | 0.1670 | 2 | 0 | -1.3 | 19.05 | |||||||||||||||||||||||||||||
| c.781G>A | D261N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant D261N is not reported in ClinVar (ClinVar status: not listed) and is absent from gnomAD (no gnomAD ID). Prediction tools that indicate a benign effect include premPS, FATHMM, and AlphaMissense‑Optimized, while the majority of tools predict a pathogenic outcome: SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. Uncertain results are reported by FoldX, Rosetta, and Foldetta. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely pathogenic, and Foldetta as uncertain. Overall, the balance of evidence favors a pathogenic classification, and this assessment does not contradict the ClinVar status, which simply lacks an entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.284882 | Structured | 0.422514 | Uncertain | 0.883 | 0.264 | 0.125 | -11.804 | Likely Pathogenic | 0.746 | Likely Pathogenic | Likely Benign | 1.58 | Ambiguous | 0.7 | 1.28 | Ambiguous | 1.43 | Ambiguous | 0.23 | Likely Benign | 0.579 | Likely Pathogenic | -2.94 | Deleterious | 0.997 | Probably Damaging | 0.989 | Probably Damaging | 5.82 | Benign | 0.02 | Affected | 0.0767 | 0.4745 | 2 | 1 | 0.0 | -0.98 | |||||||||||||||||||||||||||||
| c.823C>T | P275S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P275S is not reported in ClinVar (ClinVar ID None) but is present in gnomAD (ID 6‑33437728‑C‑T). Prediction tools that agree on a benign effect include REVEL, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that agree on a pathogenic effect are FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM. The remaining tools—Rosetta, Foldetta, premPS, and ESM1b—return uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points to a pathogenic impact for P275S, and this conclusion does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.059222 | Structured | 0.353469 | Uncertain | 0.811 | 0.208 | 0.250 | 6-33437728-C-T | 1 | 6.20e-7 | -7.886 | In-Between | 0.312 | Likely Benign | Likely Benign | 2.11 | Destabilizing | 0.3 | 1.28 | Ambiguous | 1.70 | Ambiguous | 0.77 | Ambiguous | 0.388 | Likely Benign | -5.24 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.78 | Pathogenic | 0.03 | Affected | 3.38 | 19 | 0.3489 | 0.3339 | -1 | 1 | 0.8 | -10.04 | |||||||||||||||||||||||||
| c.1644G>C | E548D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E548D variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Functional prediction tools that agree on a benign effect include REVEL, premPS, SIFT, and FATHMM, whereas a separate group predicts pathogenicity: PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, and AlphaMissense‑Optimized. Predictions that are uncertain or unavailable are FoldX, Rosetta, Foldetta, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also yields a pathogenic verdict (2 pathogenic, 1 benign, 1 uncertain). Foldetta’s stability prediction is unavailable. Overall, the majority of evidence points to a pathogenic effect, and this conclusion does not contradict the ClinVar status, which simply lacks an entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.054297 | Structured | 0.008632 | Uncertain | 0.965 | 0.288 | 0.000 | -7.359 | In-Between | 0.992 | Likely Pathogenic | Likely Pathogenic | 0.74 | Ambiguous | 0.1 | 1.29 | Ambiguous | 1.02 | Ambiguous | 0.32 | Likely Benign | 0.254 | Likely Benign | -2.85 | Deleterious | 0.998 | Probably Damaging | 0.989 | Probably Damaging | 3.51 | Benign | 0.09 | Tolerated | 0.1374 | 0.2524 | 3 | 2 | 0.0 | -14.03 | ||||||||||||||||||||||||||||||
| c.1644G>T | E548D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E548D variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Functional prediction tools that agree on a benign effect include REVEL, premPS, SIFT, and FATHMM, whereas a separate group predicts a pathogenic effect: PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, and AlphaMissense‑Optimized. Predictions that are uncertain or unavailable are FoldX, Rosetta, Foldetta, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also yields a pathogenic verdict (2 pathogenic, 1 benign, 1 uncertain). Foldetta’s stability prediction is unavailable. Overall, the majority of evidence points to a pathogenic impact for E548D, and this conclusion does not contradict the lack of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.054297 | Structured | 0.008632 | Uncertain | 0.965 | 0.288 | 0.000 | -7.359 | In-Between | 0.992 | Likely Pathogenic | Likely Pathogenic | 0.74 | Ambiguous | 0.1 | 1.29 | Ambiguous | 1.02 | Ambiguous | 0.32 | Likely Benign | 0.254 | Likely Benign | -2.85 | Deleterious | 0.998 | Probably Damaging | 0.989 | Probably Damaging | 3.51 | Benign | 0.09 | Tolerated | 0.1374 | 0.2524 | 3 | 2 | 0.0 | -14.03 | ||||||||||||||||||||||||||||||
| c.2005A>T | N669Y 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N669Y is not reported in ClinVar and has no entries in gnomAD. Prediction tools that agree on a benign effect include FoldX, FATHMM, and premPS. Those that predict a pathogenic effect comprise SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. With eight pathogenic predictions versus three benign, the overall evidence points to a pathogenic impact. This conclusion is not contradicted by ClinVar status, which currently has no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.142424 | Structured | 0.086615 | Uncertain | 0.872 | 0.380 | 0.000 | -13.548 | Likely Pathogenic | 0.802 | Likely Pathogenic | Ambiguous | 0.45 | Likely Benign | 0.4 | 1.29 | Ambiguous | 0.87 | Ambiguous | 0.21 | Likely Benign | 0.546 | Likely Pathogenic | -7.01 | Deleterious | 1.000 | Probably Damaging | 0.997 | Probably Damaging | 3.34 | Benign | 0.00 | Affected | 0.0666 | 0.4151 | -2 | -2 | 2.2 | 49.07 | |||||||||||||||||||||||||||||
| c.772C>A | R258S 2D  AIThe SynGAP1 missense variant R258S is not reported in ClinVar and has no entries in gnomAD. Prediction tools that indicate a benign effect are limited to FATHMM, whereas the majority of algorithms—SGM‑Consensus, REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—classify the change as pathogenic. High‑accuracy assessments further support a deleterious outcome: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) reports “Likely Pathogenic,” and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, yields an uncertain result. Overall, the preponderance of evidence points to a pathogenic effect for R258S, and this conclusion is consistent with the absence of ClinVar annotation or gnomAD frequency data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.295083 | Structured | 0.293667 | Uncertain | 0.894 | 0.260 | 0.250 | -14.336 | Likely Pathogenic | 0.990 | Likely Pathogenic | Likely Pathogenic | 2.11 | Destabilizing | 0.8 | 1.29 | Ambiguous | 1.70 | Ambiguous | 1.14 | Destabilizing | 0.796 | Likely Pathogenic | -4.92 | Deleterious | 0.997 | Probably Damaging | 0.987 | Probably Damaging | 5.89 | Benign | 0.01 | Affected | 0.3057 | 0.3881 | 0 | -1 | 3.7 | -69.11 | |||||||||||||||||||||||||||||
| c.953C>A | P318Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P318Q is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely converge on a deleterious effect: SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all indicate pathogenicity. Only FoldX, Rosetta, and Foldetta provide uncertain results and are therefore not considered evidence for benign impact. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized predicts pathogenicity, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta remains uncertain. Consequently, the variant is most likely pathogenic based on the consensus of predictive algorithms, and this assessment does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.111485 | Structured | 0.400936 | Uncertain | 0.858 | 0.234 | 0.000 | -11.403 | Likely Pathogenic | 0.976 | Likely Pathogenic | Likely Pathogenic | 1.64 | Ambiguous | 0.2 | 1.29 | Ambiguous | 1.47 | Ambiguous | 1.18 | Destabilizing | 0.638 | Likely Pathogenic | -7.05 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 1.83 | Pathogenic | 0.01 | Affected | 0.1467 | 0.4731 | 0 | -1 | -1.9 | 31.01 | |||||||||||||||||||||||||||||
| c.1255G>A | E419K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E419K missense variant is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, SIFT, and FATHMM. Tools that agree on a pathogenic effect include SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Foldetta and Rosetta give uncertain results and are not counted in either group. High‑accuracy assessments show AlphaMissense‑Optimized predicts pathogenic, SGM‑Consensus predicts likely pathogenic, and Foldetta remains uncertain. Overall, the majority of evidence points to a pathogenic impact for E419K. This conclusion is not contradicted by ClinVar, which contains no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.102787 | Structured | 0.371949 | Uncertain | 0.961 | 0.261 | 0.000 | -12.257 | Likely Pathogenic | 0.989 | Likely Pathogenic | Likely Pathogenic | 0.01 | Likely Benign | 0.1 | 1.30 | Ambiguous | 0.66 | Ambiguous | -0.03 | Likely Benign | 0.399 | Likely Benign | -3.75 | Deleterious | 0.998 | Probably Damaging | 0.975 | Probably Damaging | 3.36 | Benign | 0.07 | Tolerated | 0.2759 | 0.6899 | 0 | 1 | -0.4 | -0.94 | |||||||||||||||||||||||||||||
| c.1993T>G | Y665D 2D 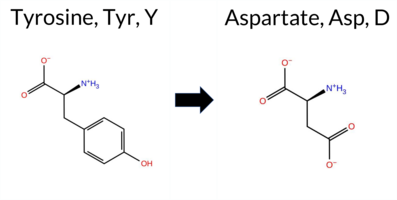 3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant Y665D is not reported in ClinVar and is absent from gnomAD. Prediction tools cluster into two groups: benign predictions come from REVEL, premPS, SIFT, FATHMM, and AlphaMissense‑Optimized; pathogenic predictions come from SGM‑Consensus, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as Likely Pathogenic (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN), and Foldetta as Uncertain. No evidence from these tools contradicts the ClinVar status, which is currently unreported. Overall, the balance of evidence—seven pathogenic versus four benign predictions, with a pathogenic consensus from SGM‑Consensus—suggests the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.098513 | Structured | 0.086641 | Uncertain | 0.922 | 0.361 | 0.000 | -10.101 | Likely Pathogenic | 0.737 | Likely Pathogenic | Likely Benign | 1.15 | Ambiguous | 0.2 | 1.30 | Ambiguous | 1.23 | Ambiguous | 0.15 | Likely Benign | 0.227 | Likely Benign | -3.67 | Deleterious | 1.000 | Probably Damaging | 0.996 | Probably Damaging | 3.75 | Benign | 1.00 | Tolerated | 0.3838 | 0.0573 | -4 | -3 | -2.2 | -48.09 | |||||||||||||||||||||||||||||
| c.2096T>C | V699A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V699A has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, and AlphaMissense‑Optimized, while premPS, polyPhen‑2 HumDiv, and polyPhen‑2 HumVar predict a pathogenic impact. Tools with inconclusive results are FoldX, Rosetta, Foldetta, and AlphaMissense‑Default. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is Benign, and the high‑accuracy AlphaMissense‑Optimized also reports Benign. Foldetta, which integrates FoldX‑MD and Rosetta outputs, is Uncertain. Overall, the majority of evidence points to a benign effect, and this assessment does not contradict any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.069024 | Structured | 0.432975 | Uncertain | 0.935 | 0.315 | 0.000 | -6.017 | Likely Benign | 0.426 | Ambiguous | Likely Benign | 1.37 | Ambiguous | 0.0 | 1.30 | Ambiguous | 1.34 | Ambiguous | 1.21 | Destabilizing | 0.236 | Likely Benign | -2.39 | Neutral | 0.861 | Possibly Damaging | 0.625 | Possibly Damaging | 3.42 | Benign | 0.54 | Tolerated | 0.2450 | 0.2124 | 0 | 0 | -2.4 | -28.05 | |||||||||||||||||||||||||||||
| c.776G>A | R259Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R259Q is catalogued in gnomAD (6‑33437681‑G‑A) but has no entry in ClinVar. In silico assessment shows a consensus of pathogenicity: 9 of 11 evaluated tools (REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a deleterious effect, while only FATHMM indicates a benign outcome. Predictions of protein‑stability change are inconclusive, with FoldX, Rosetta and the combined Foldetta method returning uncertain results. High‑accuracy predictors reinforce the pathogenic signal: AlphaMissense‑Optimized is pathogenic, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also classifies the variant as likely pathogenic; Foldetta remains uncertain. Overall, the computational evidence strongly favors a pathogenic interpretation, and this is consistent with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.222385 | Structured | 0.338208 | Uncertain | 0.885 | 0.255 | 0.250 | 6-33437681-G-A | 1 | 6.20e-7 | -12.598 | Likely Pathogenic | 0.966 | Likely Pathogenic | Likely Pathogenic | 1.19 | Ambiguous | 0.3 | 1.30 | Ambiguous | 1.25 | Ambiguous | 1.40 | Destabilizing | 0.851 | Likely Pathogenic | -3.68 | Deleterious | 0.999 | Probably Damaging | 0.978 | Probably Damaging | 5.81 | Benign | 0.01 | Affected | 3.39 | 15 | 0.3329 | 0.2703 | 1 | 1 | 1.0 | -28.06 | 258.7 | 52.8 | 0.1 | 0.1 | -0.3 | 0.4 | X | X | Potentially Pathogenic | The guanidinium group of Arg259, located at the beginning of an anti-parallel β sheet strand (res. Arg259-Arg272), forms salt bridges with the carboxylate groups of Asp684 at the end of an α helix (res. Ile683-Gln702, GAP domain) and Asp261 on the same β strand. The Arg259 side chain also frequently forms hydrogen bonds with the backbone carbonyl groups of Ser257, Asn256, and Asp255. In the variant simulations, the carboxamide group of the Gln259 side chain cannot form salt bridges or maintain hydrogen bonding with the carboxylate group of Asp684, which could affect the tertiary structure assembly between the C2 and GAP domains. Notably, the amino group of the Lys254 side chain maintains a salt bridge with Asp684 and Glu244 throughout the variant simulations, but this interaction is not maintained in the WT simulations. Thus, the partially or loosely α helical conformation of a lysine-containing loop (res. Lys251-Ser257), which extends to a nearby α helix (res. Met414-Asn426), could be stabilized due to the residue swap. | ||||||||||||||
| c.877C>A | R293S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R293S is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a deleterious effect include REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote). Tools that are inconclusive or uncertain are Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized classifying the variant as pathogenic, the SGM‑Consensus indicating a likely pathogenic outcome, while Foldetta’s stability analysis remains uncertain. Taken together, the overwhelming majority of evidence points to a pathogenic impact for R293S. This conclusion is consistent with the lack of ClinVar annotation, as there is no conflicting status to contradict the prediction. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.335645 | Structured | 0.338192 | Uncertain | 0.924 | 0.269 | 0.125 | -14.103 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 2.66 | Destabilizing | 0.2 | 1.30 | Ambiguous | 1.98 | Ambiguous | 0.71 | Ambiguous | 0.561 | Likely Pathogenic | -5.52 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | 1.51 | Pathogenic | 0.01 | Affected | 0.2794 | 0.5045 | 0 | -1 | 3.7 | -69.11 | |||||||||||||||||||||||||||||
| c.1445T>G | L482R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L482R is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely converge on a deleterious effect: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all indicate pathogenicity, while FoldX, Rosetta, and Foldetta return uncertain results. In a consensus framework, the SGM‑Consensus score is “Likely Pathogenic,” reflecting the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN. High‑accuracy assessments further support a damaging outcome: AlphaMissense‑Optimized predicts pathogenicity, SGM‑Consensus is Likely Pathogenic, and Foldetta remains inconclusive. Taken together, the overwhelming majority of evidence points to a pathogenic effect. Thus, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.254060 | Structured | 0.426236 | Uncertain | 0.795 | 0.248 | 0.000 | -14.684 | Likely Pathogenic | 0.976 | Likely Pathogenic | Likely Pathogenic | 0.74 | Ambiguous | 0.1 | 1.31 | Ambiguous | 1.03 | Ambiguous | 1.53 | Destabilizing | 0.878 | Likely Pathogenic | -5.92 | Deleterious | 0.998 | Probably Damaging | 0.996 | Probably Damaging | -1.27 | Pathogenic | 0.01 | Affected | 0.1255 | 0.0679 | -3 | -2 | -8.3 | 43.03 | |||||||||||||||||||||||||||||
| c.1468G>C | A490P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A490P is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Among the available in‑silico predictors, 10 tools (REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus) uniformly predict a pathogenic effect, whereas only Foldetta predicts a benign outcome; FoldX, Rosetta, and AlphaMissense‑Optimized are inconclusive. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is uncertain, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is pathogenic, and Foldetta (combining FoldX‑MD and Rosetta stability outputs) is benign. Overall, the preponderance of evidence indicates that the variant is most likely pathogenic, which does not contradict the current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.120615 | Structured | 0.322979 | Uncertain | 0.938 | 0.210 | 0.125 | Uncertain | 1 | -12.905 | Likely Pathogenic | 0.941 | Likely Pathogenic | Ambiguous | -1.27 | Ambiguous | 0.1 | 1.31 | Ambiguous | 0.02 | Likely Benign | 1.07 | Destabilizing | 0.878 | Likely Pathogenic | -4.81 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | -1.42 | Pathogenic | 0.01 | Affected | 3.37 | 35 | 0.1755 | 0.3071 | -1 | 1 | -3.4 | 26.04 | |||||||||||||||||||||||||
| c.1607T>G | L536W 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L536W is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a deleterious effect include REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized; all of these classify the variant as pathogenic. Tools that are inconclusive (FoldX, Rosetta, Foldetta) do not provide evidence for or against pathogenicity. High‑accuracy assessments further support a damaging impact: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates likely pathogenic, while Foldetta’s stability analysis is uncertain. Based on the collective evidence, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.137348 | Structured | 0.042188 | Uncertain | 0.931 | 0.341 | 0.000 | -14.169 | Likely Pathogenic | 0.970 | Likely Pathogenic | Likely Pathogenic | 1.07 | Ambiguous | 0.6 | 1.31 | Ambiguous | 1.19 | Ambiguous | 1.33 | Destabilizing | 0.897 | Likely Pathogenic | -5.92 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | -1.50 | Pathogenic | 0.00 | Affected | 0.0843 | 0.2577 | -2 | -2 | -4.7 | 73.05 | |||||||||||||||||||||||||||||
| c.622C>G | P208A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P208A is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic outcome are FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), and SIFT. Predictions that are inconclusive are Rosetta and Foldetta. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely benign, and Foldetta as uncertain. Overall, the majority of evidence points toward a benign effect, and this is consistent with the lack of ClinVar annotation; there is no contradiction with ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | PH | 0.271506 | Structured | 0.399506 | Uncertain | 0.864 | 0.345 | 0.125 | -5.623 | Likely Benign | 0.172 | Likely Benign | Likely Benign | 2.19 | Destabilizing | 0.3 | 1.31 | Ambiguous | 1.75 | Ambiguous | 1.03 | Destabilizing | 0.245 | Likely Benign | -6.80 | Deleterious | 0.999 | Probably Damaging | 0.991 | Probably Damaging | 3.80 | Benign | 0.04 | Affected | 0.3727 | 0.4390 | 1 | -1 | 3.4 | -26.04 | |||||||||||||||||||||||||||||
| c.800G>T | W267L 2D 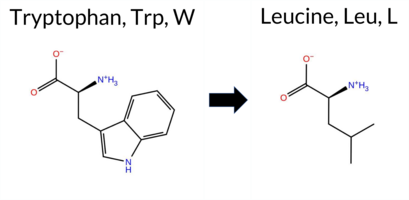 AIThe SynGAP1 missense variant W267L is not reported in ClinVar and is absent from gnomAD. Prediction tools that assess pathogenicity unanimously classify the variant as deleterious: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict pathogenicity. No tool in the dataset predicts a benign effect. Uncertain or inconclusive results come from FoldX, Rosetta, Foldetta, and premPS, which are treated as unavailable evidence. High‑accuracy assessments further support a damaging effect: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is “Likely Pathogenic,” while Foldetta’s stability prediction is uncertain. Overall, the evidence strongly indicates that W267L is most likely pathogenic, and this conclusion does not contradict any ClinVar status because no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.216401 | Structured | 0.298060 | Uncertain | 0.943 | 0.274 | 0.000 | -13.670 | Likely Pathogenic | 0.990 | Likely Pathogenic | Likely Pathogenic | 1.19 | Ambiguous | 0.7 | 1.31 | Ambiguous | 1.25 | Ambiguous | 0.75 | Ambiguous | 0.628 | Likely Pathogenic | -11.95 | Deleterious | 0.999 | Probably Damaging | 0.996 | Probably Damaging | 1.93 | Pathogenic | 0.01 | Affected | 0.2122 | 0.2843 | -2 | -2 | 4.7 | -73.05 | |||||||||||||||||||||||||||||
| c.1450T>C | F484L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F484L is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split opinion: benign predictions come from REVEL, polyPhen‑2 (HumDiv and HumVar), and FATHMM, while pathogenic predictions arise from SGM‑Consensus (Likely Pathogenic), premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The high‑accuracy consensus, SGM‑Consensus, is derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN (3 pathogenic vs. 1 benign) and therefore also indicates pathogenicity. AlphaMissense‑Optimized independently predicts pathogenicity. Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is inconclusive. Overall, seven tools predict pathogenicity versus four predicting benign, and the high‑accuracy consensus supports a pathogenic interpretation. Thus, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.182256 | Structured | 0.403079 | Uncertain | 0.798 | 0.245 | 0.125 | -11.052 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 1.52 | Ambiguous | 0.0 | 1.32 | Ambiguous | 1.42 | Ambiguous | 1.20 | Destabilizing | 0.275 | Likely Benign | -5.64 | Deleterious | 0.054 | Benign | 0.022 | Benign | 3.20 | Benign | 0.04 | Affected | 0.1737 | 0.2956 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1452C>A | F484L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F484L is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split opinion: benign predictions come from REVEL, polyPhen‑2 (HumDiv and HumVar), and FATHMM, while pathogenic predictions arise from SGM‑Consensus (Likely Pathogenic), premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The high‑accuracy consensus, SGM‑Consensus, is derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN (3 pathogenic vs. 1 benign) and therefore also indicates pathogenicity. AlphaMissense‑Optimized independently predicts pathogenicity. Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is inconclusive. Overall, seven tools predict pathogenicity versus four predicting benign, and the high‑accuracy consensus supports a pathogenic interpretation. Thus, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.182256 | Structured | 0.403079 | Uncertain | 0.798 | 0.245 | 0.125 | -11.052 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 1.52 | Ambiguous | 0.0 | 1.32 | Ambiguous | 1.42 | Ambiguous | 1.20 | Destabilizing | 0.214 | Likely Benign | -5.64 | Deleterious | 0.054 | Benign | 0.022 | Benign | 3.20 | Benign | 0.04 | Affected | 0.1737 | 0.2956 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1452C>G | F484L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F484L is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split opinion: benign predictions come from REVEL, polyPhen‑2 (HumDiv and HumVar), and FATHMM, while pathogenic predictions arise from SGM‑Consensus (Likely Pathogenic), premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenic, and the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates likely pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, yields an uncertain result. Overall, seven tools predict pathogenicity versus four predicting benign, and the high‑accuracy methods reinforce the pathogenic signal. Thus, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.182256 | Structured | 0.403079 | Uncertain | 0.798 | 0.245 | 0.125 | -11.052 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 1.52 | Ambiguous | 0.0 | 1.32 | Ambiguous | 1.42 | Ambiguous | 1.20 | Destabilizing | 0.214 | Likely Benign | -5.64 | Deleterious | 0.054 | Benign | 0.022 | Benign | 3.20 | Benign | 0.04 | Affected | 0.1737 | 0.2956 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1531G>A | G511R 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant G511R is listed in ClinVar as Pathogenic (ClinVar ID 1774641.0) and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, and FATHMM, while pathogenic predictions are made by PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts Pathogenic; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also indicates Pathogenic; Foldetta, which integrates FoldX‑MD and Rosetta outputs, is inconclusive. Stability calculations from FoldX and Rosetta are uncertain, and premPS is unavailable. Overall, the majority of evidence points to a pathogenic impact, aligning with the ClinVar classification and not contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.048328 | Structured | 0.244404 | Uncertain | 0.924 | 0.287 | 0.000 | Likely Pathogenic | 1 | -11.327 | Likely Pathogenic | 0.991 | Likely Pathogenic | Likely Pathogenic | 1.94 | Ambiguous | 0.3 | 1.32 | Ambiguous | 1.63 | Ambiguous | 0.94 | Ambiguous | 0.416 | Likely Benign | -7.72 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.26 | Benign | 0.06 | Tolerated | 3.37 | 35 | 0.1307 | 0.4104 | -3 | -2 | -4.1 | 99.14 | 279.4 | -159.9 | 0.0 | 0.0 | 0.7 | 0.1 | X | X | Potentially Pathogenic | Gly511 is located in an α-helix (res. Gly502-Tyr518), facing hydrophobic residues in an inter-helix space (e.g., Leu610, Ile514) in the WT simulations. In contrast, in the variant simulations, the bulkier and positively charged guanidinium side chain of Arg511 forms a salt bridge with the carboxylate group of Glu217 or hydrogen bonds with the backbone carbonyl group of Leu610. Although the residue swap introduces a third positively charged residue in close vicinity (Arg511, Lys507, Arg515), the protein structure seems to remain stable in the variant simulations. Importantly, according to ClinVar, the residue swap alters the last nucleotide of an exon and is predicted to destroy the splice donor site, resulting in aberrant splicing and pathogenic status. | 10.1016/j.ajhg.2020.11.011 | ||||||||||||||
| c.1531G>C | G511R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G511R is listed in ClinVar (ID 452818.0) as Pathogenic and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, SIFT, and FATHMM, whereas those that predict a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). High‑accuracy assessments show AlphaMissense‑Optimized as Pathogenic, the SGM‑Consensus as Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as Uncertain, which is treated as unavailable evidence. Overall, the majority of available predictions support a pathogenic impact, aligning with the ClinVar classification. Thus, the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.048328 | Structured | 0.244404 | Uncertain | 0.924 | 0.287 | 0.000 | Pathogenic | 1 | -11.327 | Likely Pathogenic | 0.991 | Likely Pathogenic | Likely Pathogenic | 1.94 | Ambiguous | 0.3 | 1.32 | Ambiguous | 1.63 | Ambiguous | 0.94 | Ambiguous | 0.415 | Likely Benign | -7.72 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.26 | Benign | 0.06 | Tolerated | 3.37 | 35 | 0.1307 | 0.4104 | -3 | -2 | -4.1 | 99.14 | 279.4 | -159.9 | 0.0 | 0.0 | 0.7 | 0.1 | X | X | Potentially Pathogenic | Gly511 is located in an α-helix (res. Gly502-Tyr518), facing hydrophobic residues in an inter-helix space (e.g., Leu610, Ile514) in the WT simulations. In contrast, in the variant simulations, the bulkier and positively charged guanidinium side chain of Arg511 forms a salt bridge with the carboxylate group of Glu217 or hydrogen bonds with the backbone carbonyl group of Leu610. Although the residue swap introduces a third positively charged residue in close vicinity (Arg511, Lys507, Arg515), the protein structure seems to remain stable in the variant simulations. Importantly, according to ClinVar, the residue swap alters the last nucleotide of an exon and is predicted to destroy the splice donor site, resulting in aberrant splicing and pathogenic status. | 10.1016/j.ajhg.2020.11.011 | ||||||||||||||
| c.1846G>T | D616Y 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant D616Y is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that indicate a benign effect include REVEL, premPS, and FATHMM, whereas a majority of tools predict a pathogenic outcome: SIFT, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also classifies the variant as likely pathogenic. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, SGM Consensus as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. No evidence from FoldX or Rosetta alone is conclusive. Overall, the balance of evidence favors a pathogenic interpretation, and this conclusion does not contradict any existing ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.129801 | Structured | 0.166689 | Uncertain | 0.867 | 0.252 | 0.000 | -12.638 | Likely Pathogenic | 0.957 | Likely Pathogenic | Likely Pathogenic | 1.70 | Ambiguous | 0.3 | 1.32 | Ambiguous | 1.51 | Ambiguous | 0.35 | Likely Benign | 0.374 | Likely Benign | -7.43 | Deleterious | 0.999 | Probably Damaging | 0.970 | Probably Damaging | 3.28 | Benign | 0.01 | Affected | 0.0465 | 0.4069 | -4 | -3 | 2.2 | 48.09 | |||||||||||||||||||||||||||||
| c.1948A>T | N650Y 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N650Y is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include premPS and FATHMM, whereas the majority of tools (SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic impact; FoldX, Rosetta, and Foldetta are inconclusive. High‑accuracy assessments reinforce this trend: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also predicts pathogenic, and Foldetta remains uncertain. Overall, the evidence strongly favors a pathogenic classification for the variant, and this conclusion does not contradict the ClinVar status, which currently contains no entry for this mutation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.086953 | Structured | 0.361944 | Uncertain | 0.961 | 0.357 | 0.000 | -16.923 | Likely Pathogenic | 0.994 | Likely Pathogenic | Likely Pathogenic | 1.56 | Ambiguous | 0.3 | 1.32 | Ambiguous | 1.44 | Ambiguous | 0.16 | Likely Benign | 0.587 | Likely Pathogenic | -7.98 | Deleterious | 1.000 | Probably Damaging | 0.984 | Probably Damaging | 3.03 | Benign | 0.03 | Affected | 0.0852 | 0.3865 | -2 | -2 | 2.2 | 49.07 | |||||||||||||||||||||||||||||
| c.2111G>T | S704I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S704I lies in the GAP domain. ClinVar has no entry for this variant, and it is not reported in gnomAD. Prediction tools that indicate a benign effect include REVEL, premPS, and FATHMM, whereas the majority of other in silico predictors (PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus) report a pathogenic or likely pathogenic outcome. FoldX, Rosetta, and Foldetta provide uncertain results. High‑accuracy methods specifically give AlphaMissense‑Optimized as pathogenic, the SGM Consensus as likely pathogenic, and Foldetta as uncertain. Based on the overall consensus, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.096677 | Structured | 0.383620 | Uncertain | 0.928 | 0.363 | 0.000 | -14.222 | Likely Pathogenic | 0.978 | Likely Pathogenic | Likely Pathogenic | 1.63 | Ambiguous | 0.1 | 1.32 | Ambiguous | 1.48 | Ambiguous | 0.29 | Likely Benign | 0.232 | Likely Benign | -4.05 | Deleterious | 0.997 | Probably Damaging | 0.758 | Possibly Damaging | 3.49 | Benign | 0.02 | Affected | 0.0727 | 0.4798 | -1 | -2 | 5.3 | 26.08 | |||||||||||||||||||||||||||||
| c.2117A>G | E706G 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant E706G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, premPS, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and polyPhen2_HumVar all classify the substitution as benign or tolerated. Only polyPhen2_HumDiv predicts a pathogenic effect. Tools with uncertain outcomes—AlphaMissense‑Default, FoldX, Rosetta, and Foldetta—do not provide a definitive assessment. High‑accuracy predictors reinforce the benign consensus: AlphaMissense‑Optimized reports a benign change; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, indicates a likely benign effect; and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, yields an uncertain result. Overall, the majority of evidence supports a benign impact, and this conclusion is consistent with the absence of a ClinVar pathogenic classification. Thus, the variant is most likely benign, and this is not contradictory to ClinVar. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.200174 | Structured | 0.377033 | Uncertain | 0.929 | 0.363 | 0.000 | -5.289 | Likely Benign | 0.535 | Ambiguous | Likely Benign | 1.22 | Ambiguous | 0.0 | 1.32 | Ambiguous | 1.27 | Ambiguous | 0.12 | Likely Benign | 0.071 | Likely Benign | -1.71 | Neutral | 0.931 | Possibly Damaging | 0.138 | Benign | 4.07 | Benign | 0.23 | Tolerated | 0.2781 | 0.3614 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.638T>C | I213T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 I213T missense variant is not reported in ClinVar and is absent from gnomAD. Prediction tools largely agree on a deleterious effect: FATHMM is the sole benign predictor, while the remaining evidence—SGM‑Consensus (Likely Pathogenic), REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—consistently indicates pathogenicity. FoldX, Rosetta, and Foldetta provide uncertain results. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta remains uncertain. Overall, the preponderance of evidence supports a pathogenic classification for I213T, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.158265 | Structured | 0.372201 | Uncertain | 0.850 | 0.295 | 0.125 | -11.080 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 1.46 | Ambiguous | 0.8 | 1.32 | Ambiguous | 1.39 | Ambiguous | 1.49 | Destabilizing | 0.882 | Likely Pathogenic | -3.99 | Deleterious | 0.948 | Possibly Damaging | 0.588 | Possibly Damaging | 5.82 | Benign | 0.00 | Affected | 0.0905 | 0.0708 | 0 | -1 | -5.2 | -12.05 | |||||||||||||||||||||||||||||
| c.988G>C | D330H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D330H missense variant is not reported in ClinVar and is absent from gnomAD. Prediction tools cluster into three groups: benign predictions are limited to REVEL; pathogenic predictions include PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, FoldX, and the SGM‑Consensus (Likely Pathogenic). Uncertain or inconclusive results come from Rosetta, Foldetta, and premPS. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenicity, the SGM‑Consensus also indicates likely pathogenic, while Foldetta’s stability analysis is inconclusive. Taken together, the overwhelming majority of evidence points to a pathogenic impact for D330H. This conclusion is not contradicted by any ClinVar annotation, as the variant is currently unreported in that database. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.380708 | Structured | 0.360008 | Uncertain | 0.805 | 0.488 | 0.250 | -13.926 | Likely Pathogenic | 0.980 | Likely Pathogenic | Likely Pathogenic | 2.29 | Destabilizing | 0.6 | 1.32 | Ambiguous | 1.81 | Ambiguous | 0.61 | Ambiguous | 0.425 | Likely Benign | -4.67 | Deleterious | 0.998 | Probably Damaging | 0.961 | Probably Damaging | 0.96 | Pathogenic | 0.01 | Affected | 0.1608 | 0.4843 | 1 | -1 | 0.3 | 22.05 | |||||||||||||||||||||||||||||
| c.1010A>C | K337T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K337T is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two consensus groups: benign predictions come from REVEL, FoldX, and premPS, whereas pathogenic predictions are made by PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. Three tools report uncertainty: Rosetta, Foldetta, and AlphaMissense‑Optimized. The high‑accuracy consensus, SGM‑Consensus, classifies the variant as Likely Pathogenic. In the high‑accuracy subset, AlphaMissense‑Optimized remains uncertain, SGM‑Consensus is Likely Pathogenic, and Foldetta is uncertain. Taken together, the majority of evidence points toward a deleterious effect. Therefore, K337T is most likely pathogenic, and this assessment does not conflict with the absence of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.321458 | Structured | 0.348540 | Uncertain | 0.449 | 0.438 | 0.500 | -10.896 | Likely Pathogenic | 0.953 | Likely Pathogenic | Ambiguous | 0.45 | Likely Benign | 0.2 | 1.33 | Ambiguous | 0.89 | Ambiguous | 0.25 | Likely Benign | 0.338 | Likely Benign | -5.32 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | 1.70 | Pathogenic | 0.01 | Affected | 0.1741 | 0.3354 | 0 | -1 | 3.2 | -27.07 | |||||||||||||||||||||||||||||
| c.1024T>C | Y342H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant Y342H is reported in gnomAD (ID 6‑33437929‑T‑C) but has no ClinVar entry. Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, and ESM1b, while pathogenic predictions are made by PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, FATHMM, and AlphaMissense‑Default. Five tools predict pathogenicity versus three predicting benign, with the remaining five (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Optimized) yielding uncertain or inconclusive results. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is uncertain, SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is uncertain. Overall, the preponderance of evidence indicates that Y342H is most likely pathogenic, and this conclusion is not contradicted by ClinVar status, which currently contains no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.366687 | Structured | 0.408200 | Uncertain | 0.866 | 0.487 | 0.250 | 6-33437929-T-C | 1 | 6.20e-7 | -6.459 | Likely Benign | 0.944 | Likely Pathogenic | Ambiguous | 1.63 | Ambiguous | 0.1 | 1.33 | Ambiguous | 1.48 | Ambiguous | 0.73 | Ambiguous | 0.453 | Likely Benign | -3.61 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.72 | Pathogenic | 0.06 | Tolerated | 3.37 | 25 | 0.2491 | 0.0862 | 2 | 0 | -1.9 | -26.03 | ||||||||||||||||||||||||
| c.1187G>A | G396D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G396D is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Optimized all classify the substitution as benign, whereas only AlphaMissense‑Default predicts a pathogenic outcome. Tools that assess protein stability (FoldX, Rosetta, Foldetta) yield uncertain or inconclusive results. High‑accuracy consensus methods reinforce the benign assessment: the SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports “Likely Benign”; AlphaMissense‑Optimized also predicts benign; Foldetta, which integrates FoldX‑MD and Rosetta outputs, remains uncertain. Based on the preponderance of evidence, the variant is most likely benign, and this conclusion is consistent with the absence of a ClinVar pathogenic classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.414856 | Structured | 0.394626 | Uncertain | 0.640 | 0.584 | 0.500 | -4.148 | Likely Benign | 0.678 | Likely Pathogenic | Likely Benign | 1.92 | Ambiguous | 0.8 | 1.33 | Ambiguous | 1.63 | Ambiguous | 0.17 | Likely Benign | 0.272 | Likely Benign | -1.49 | Neutral | 0.421 | Benign | 0.080 | Benign | 3.91 | Benign | 0.35 | Tolerated | 0.1702 | 0.1122 | 1 | -1 | -3.1 | 58.04 | |||||||||||||||||||||||||||||
| c.1204C>A | L402M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L402M is not reported in ClinVar (ClinVar status: not listed) and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, FoldX, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT. Uncertain results come from Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Overall, the majority of evidence points to a benign impact for this variant, and this conclusion does not contradict any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.243554 | Structured | 0.431978 | Uncertain | 0.961 | 0.383 | 0.000 | -6.991 | Likely Benign | 0.305 | Likely Benign | Likely Benign | 0.12 | Likely Benign | 0.0 | 1.33 | Ambiguous | 0.73 | Ambiguous | 0.80 | Ambiguous | 0.071 | Likely Benign | -0.99 | Neutral | 0.994 | Probably Damaging | 0.938 | Probably Damaging | 3.73 | Benign | 0.02 | Affected | 0.1064 | 0.4095 | 4 | 2 | -1.9 | 18.03 | |||||||||||||||||||||||||||||
| c.1228A>G | S410G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S410G is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, polyPhen‑2 HumVar, SIFT, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are PROVEAN and polyPhen‑2 HumDiv. The remaining tools (FoldX, Rosetta, Foldetta, premPS, ESM1b) give uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as benign, and Foldetta as uncertain. Overall, the majority of evidence points to a benign impact. This conclusion does not contradict ClinVar status, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | 0.098513 | Structured | 0.349627 | Uncertain | 0.908 | 0.206 | 0.000 | -7.147 | In-Between | 0.137 | Likely Benign | Likely Benign | 0.58 | Ambiguous | 0.1 | 1.33 | Ambiguous | 0.96 | Ambiguous | 0.85 | Ambiguous | 0.117 | Likely Benign | -2.54 | Deleterious | 0.952 | Possibly Damaging | 0.145 | Benign | 4.13 | Benign | 0.19 | Tolerated | 0.2651 | 0.5143 | 1 | 0 | 0.4 | -30.03 | |||||||||||||||||||||||||||||||
| c.1679T>C | V560A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V560A is not reported in ClinVar (ClinVar status: none) and has no entry in gnomAD (gnomAD ID: none). Prediction tools that agree on a benign effect include REVEL, SIFT, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect comprise SGM‑Consensus, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, and AlphaMissense‑Default. FoldX, Rosetta, and Foldetta provide uncertain or inconclusive results and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic; Foldetta remains uncertain. Overall, the majority of predictions (eight pathogenic vs. three benign) indicate that the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status because no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.021381 | Structured | 0.013872 | Uncertain | 0.853 | 0.204 | 0.000 | -8.260 | Likely Pathogenic | 0.701 | Likely Pathogenic | Likely Benign | 0.54 | Ambiguous | 0.1 | 1.33 | Ambiguous | 0.94 | Ambiguous | 1.19 | Destabilizing | 0.447 | Likely Benign | -3.15 | Deleterious | 0.911 | Possibly Damaging | 0.657 | Possibly Damaging | -1.20 | Pathogenic | 0.31 | Tolerated | 0.2549 | 0.2212 | 0 | 0 | -2.4 | -28.05 | |||||||||||||||||||||||||||||
| c.1698G>C | K566N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K566N is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that assess pathogenicity uniformly indicate a deleterious effect: REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict pathogenicity, while Rosetta is uncertain. No tool predicts a benign outcome. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. Consequently, the variant is most likely pathogenic based on the available predictions, and this assessment does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.027463 | Structured | 0.047887 | Uncertain | 0.924 | 0.219 | 0.000 | -11.255 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 2.83 | Destabilizing | 0.3 | 1.33 | Ambiguous | 2.08 | Destabilizing | 1.32 | Destabilizing | 0.683 | Likely Pathogenic | -4.04 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.43 | Pathogenic | 0.03 | Affected | 3.37 | 35 | 0.3042 | 0.1567 | 0 | 1 | 0.4 | -14.07 | |||||||||||||||||||||||||||
| c.1698G>T | K566N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K566N is listed in gnomAD (variant ID 6‑33440750‑G‑T) but has no ClinVar entry. All available in‑silico predictors that provide a definitive call classify the substitution as pathogenic: SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. The only tool with an inconclusive result is Rosetta (Uncertain), which is treated as unavailable evidence. High‑accuracy methods reinforce this assessment: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. Consequently, the variant is most likely pathogenic based on current predictions, and this conclusion does not contradict any ClinVar status because none is reported. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.027463 | Structured | 0.047887 | Uncertain | 0.924 | 0.219 | 0.000 | 6-33440750-G-T | -11.255 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 2.83 | Destabilizing | 0.3 | 1.33 | Ambiguous | 2.08 | Destabilizing | 1.32 | Destabilizing | 0.683 | Likely Pathogenic | -4.04 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.43 | Pathogenic | 0.03 | Affected | 3.37 | 35 | 0.3042 | 0.1567 | 0 | 1 | 0.4 | -14.07 | ||||||||||||||||||||||||||
| c.2110A>G | S704G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S704G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv and polyPhen‑2 HumVar. Predictions that are inconclusive or unavailable are FoldX, Rosetta, Foldetta, premPS, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus as likely benign, and Foldetta as uncertain. Taken together, the majority of reliable predictors indicate a benign impact, and this conclusion does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.096677 | Structured | 0.383620 | Uncertain | 0.928 | 0.363 | 0.000 | -7.827 | In-Between | 0.169 | Likely Benign | Likely Benign | 1.05 | Ambiguous | 0.1 | 1.33 | Ambiguous | 1.19 | Ambiguous | 0.53 | Ambiguous | 0.091 | Likely Benign | -2.25 | Neutral | 0.981 | Probably Damaging | 0.514 | Possibly Damaging | 3.42 | Benign | 0.07 | Tolerated | 0.2224 | 0.3378 | 1 | 0 | 0.4 | -30.03 | |||||||||||||||||||||||||||||
| c.881C>T | T294I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant T294I is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on pathogenicity include REVEL, FoldX, Foldetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools with uncertain or inconclusive results are Rosetta and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as pathogenic. Based on the overwhelming agreement among pathogenic predictions and the high‑accuracy tool results, the variant is most likely pathogenic. This conclusion does not contradict ClinVar status, as no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.328603 | Structured | 0.316932 | Uncertain | 0.919 | 0.267 | 0.125 | -15.302 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 4.09 | Destabilizing | 0.2 | 1.33 | Ambiguous | 2.71 | Destabilizing | 0.54 | Ambiguous | 0.768 | Likely Pathogenic | -5.52 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | -0.19 | Pathogenic | 0.01 | Affected | 0.0808 | 0.5485 | 0 | -1 | 5.2 | 12.05 | |||||||||||||||||||||||||||||
| c.958G>C | V320L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V320L is listed in ClinVar with an uncertain significance and is present in gnomAD (ID 6‑33437863‑G‑C). Functional prediction tools that agree on benign impact include REVEL, FoldX, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, and AlphaMissense‑Optimized. Pathogenic predictions come from polyPhen‑2 HumDiv and FATHMM, while Rosetta, Foldetta, premPS, and AlphaMissense‑Default are inconclusive. The high‑accuracy consensus (SGM Consensus) derived from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN yields a benign verdict. AlphaMissense‑Optimized also predicts benign, whereas Foldetta remains uncertain. Overall, the majority of evidence points to a benign effect, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.185198 | Structured | 0.419626 | Uncertain | 0.905 | 0.266 | 0.125 | Uncertain | 2 | 6-33437863-G-C | 6 | 3.72e-6 | -6.207 | Likely Benign | 0.362 | Ambiguous | Likely Benign | -0.26 | Likely Benign | 0.2 | 1.33 | Ambiguous | 0.54 | Ambiguous | 0.51 | Ambiguous | 0.096 | Likely Benign | -1.02 | Neutral | 0.900 | Possibly Damaging | 0.373 | Benign | 1.78 | Pathogenic | 0.92 | Tolerated | 3.38 | 23 | 0.0661 | 0.3863 | 2 | 1 | -0.4 | 14.03 | 245.8 | -10.2 | 0.3 | 0.9 | 0.1 | 0.3 | X | Potentially Benign | The isopropyl side chain of Val310, located in a β hairpin loop linking two anti-parallel β sheet strands (res. Thr305-Asn315, res. Ala322-Asp330), hydrophobically packs with the side chains of nearby residues (e.g., Leu286, Val350, Pro318). The hydrophobic Leu320 side chain mostly forms the same interactions; hence, the residue swap does not seem to negatively affect the protein structure based on the variant simulations. | ||||||||||||||
| c.1012G>A | D338N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D338N missense variant is not reported in ClinVar (ClinVar ID None) and has no entries in gnomAD (gnomAD ID None). Prediction tools that indicate a benign effect include REVEL, premPS, and polyPhen‑2 HumVar, whereas a majority of tools predict pathogenicity: SGM‑Consensus (Likely Pathogenic), PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. No evidence from FoldX, Rosetta, or Foldetta supports a benign outcome. Overall, the balance of evidence favors a pathogenic interpretation; this is consistent with the absence of ClinVar annotation, so there is no contradiction with existing clinical data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.335645 | Structured | 0.363354 | Uncertain | 0.460 | 0.438 | 0.375 | -9.520 | Likely Pathogenic | 0.809 | Likely Pathogenic | Ambiguous | 0.95 | Ambiguous | 0.4 | 1.34 | Ambiguous | 1.15 | Ambiguous | 0.06 | Likely Benign | 0.442 | Likely Benign | -3.62 | Deleterious | 0.801 | Possibly Damaging | 0.315 | Benign | 1.71 | Pathogenic | 0.02 | Affected | 0.1399 | 0.5970 | 2 | 1 | 0.0 | -0.98 | |||||||||||||||||||||||||||||
| c.1238C>G | P413R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P413R is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only FATHMM, whereas the remaining tools—SGM‑Consensus, REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—consistently predict a pathogenic impact. The remaining methods, Foldetta and Rosetta, yield uncertain results and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, SGM‑Consensus as likely pathogenic (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN), and Foldetta as uncertain. Overall, the majority of evidence points to a pathogenic effect, and this conclusion does not contradict any ClinVar annotation because no ClinVar status exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.113710 | Structured | 0.332472 | Uncertain | 0.927 | 0.201 | 0.000 | -15.523 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 2.40 | Destabilizing | 0.2 | 1.34 | Ambiguous | 1.87 | Ambiguous | 1.07 | Destabilizing | 0.562 | Likely Pathogenic | -8.29 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 3.32 | Benign | 0.01 | Affected | 0.1550 | 0.2199 | 0 | -2 | -2.9 | 59.07 | |||||||||||||||||||||||||||||
| c.1238C>T | P413L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P413L is not reported in ClinVar (ClinVar status: none) and is absent from gnomAD (gnomAD ID: none). Prediction tools that agree on a benign effect include REVEL, premPS, and FATHMM. Those that predict a pathogenic effect comprise SGM‑Consensus, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Rosetta and Foldetta give uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points to a pathogenic impact. This conclusion does not contradict ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.113710 | Structured | 0.332472 | Uncertain | 0.927 | 0.201 | 0.000 | -12.735 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 2.61 | Destabilizing | 0.4 | 1.34 | Ambiguous | 1.98 | Ambiguous | 0.30 | Likely Benign | 0.461 | Likely Benign | -9.21 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 3.19 | Benign | 0.00 | Affected | 0.2056 | 0.5614 | -3 | -3 | 5.4 | 16.04 | |||||||||||||||||||||||||||||
| c.1315C>A | L439M 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L439M is reported in gnomAD (variant ID 6‑33438220‑C‑A) but has no ClinVar entry. Functional prediction tools that agree on benign impact include REVEL, FoldX, PROVEAN, ESM1b, FATHMM, and AlphaMissense‑Optimized, while pathogenic predictions come from premPS, polyPhen‑2 (HumDiv and HumVar) and SIFT. Uncertain results are reported by Rosetta, Foldetta, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely benign, and Foldetta as inconclusive. Overall, the majority of evidence points to a benign effect; there is no ClinVar classification to contradict this assessment. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.222385 | Structured | 0.281542 | Uncertain | 0.942 | 0.265 | 0.000 | 6-33438220-C-A | 1 | 6.20e-7 | -5.840 | Likely Benign | 0.363 | Ambiguous | Likely Benign | -0.33 | Likely Benign | 0.1 | 1.34 | Ambiguous | 0.51 | Ambiguous | 1.01 | Destabilizing | 0.187 | Likely Benign | -1.43 | Neutral | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.24 | Benign | 0.02 | Affected | 3.38 | 25 | 0.0720 | 0.2753 | 2 | 4 | -1.9 | 18.03 | ||||||||||||||||||||||||
| c.1410G>A | M470I 2D  AIThe SynGAP1 missense variant M470I is not reported in ClinVar and has no entry in gnomAD. Prediction tools that agree on a benign effect include only SIFT, whereas the remaining tools (REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default) all predict a pathogenic impact. Predictions marked as uncertain (AlphaMissense‑Optimized, FoldX, Rosetta, Foldetta, premPS) are treated as unavailable. High‑accuracy assessments show the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, while AlphaMissense‑Optimized and Foldetta are uncertain. Overall, the preponderance of evidence points to a pathogenic effect. This conclusion is not contradicted by ClinVar status, which currently has no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.298791 | Structured | 0.351497 | Uncertain | 0.908 | 0.272 | 0.000 | -9.474 | Likely Pathogenic | 0.936 | Likely Pathogenic | Ambiguous | 1.53 | Ambiguous | 0.7 | 1.34 | Ambiguous | 1.44 | Ambiguous | 0.84 | Ambiguous | 0.747 | Likely Pathogenic | -3.55 | Deleterious | 0.833 | Possibly Damaging | 0.886 | Possibly Damaging | -1.26 | Pathogenic | 0.07 | Tolerated | 0.1061 | 0.2827 | 2 | 1 | 2.6 | -18.03 | |||||||||||||||||||||||||||||
| c.1410G>C | M470I 2D  AIThe SynGAP1 missense variant M470I is not reported in ClinVar and is absent from gnomAD. Prediction tools that classify it as benign include only SIFT, whereas the remaining evidence—REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus—predict pathogenicity. Results from high‑accuracy methods are mixed: AlphaMissense‑Optimized is uncertain, the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates likely pathogenic, and Foldetta is uncertain. Overall, the preponderance of predictions supports a pathogenic effect for M470I, and this assessment does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.298791 | Structured | 0.351497 | Uncertain | 0.908 | 0.272 | 0.000 | -9.474 | Likely Pathogenic | 0.936 | Likely Pathogenic | Ambiguous | 1.53 | Ambiguous | 0.7 | 1.34 | Ambiguous | 1.44 | Ambiguous | 0.84 | Ambiguous | 0.747 | Likely Pathogenic | -3.55 | Deleterious | 0.833 | Possibly Damaging | 0.886 | Possibly Damaging | -1.26 | Pathogenic | 0.07 | Tolerated | 0.1061 | 0.2827 | 2 | 1 | 2.6 | -18.03 | |||||||||||||||||||||||||||||
| c.1410G>T | M470I 2D  AIThe SynGAP1 missense variant M470I is not reported in ClinVar and is absent from gnomAD. Prediction tools that classify it as benign include only SIFT, whereas the remaining evidence—REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus—predict pathogenicity. Results from high‑accuracy methods are mixed: AlphaMissense‑Optimized is uncertain, the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates likely pathogenic, and Foldetta is uncertain. Overall, the preponderance of predictions supports a pathogenic effect for M470I, and this assessment does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.298791 | Structured | 0.351497 | Uncertain | 0.908 | 0.272 | 0.000 | -9.474 | Likely Pathogenic | 0.936 | Likely Pathogenic | Ambiguous | 1.53 | Ambiguous | 0.7 | 1.34 | Ambiguous | 1.44 | Ambiguous | 0.84 | Ambiguous | 0.747 | Likely Pathogenic | -3.55 | Deleterious | 0.833 | Possibly Damaging | 0.886 | Possibly Damaging | -1.26 | Pathogenic | 0.07 | Tolerated | 0.1061 | 0.2827 | 2 | 1 | 2.6 | -18.03 | |||||||||||||||||||||||||||||
| c.1838A>G | E613G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E613G is not reported in ClinVar and has no entry in gnomAD. Prediction tools that agree on a benign effect include only premPS, whereas the remaining tools (SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default) uniformly predict a pathogenic impact. The high‑accuracy methods give the following results: AlphaMissense‑Optimized is uncertain; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, is uncertain. No prediction or stability assessment is missing or inconclusive beyond the uncertain labels. Overall, the preponderance of evidence points to a pathogenic effect for E613G, and this assessment does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.275179 | Structured | 0.193489 | Uncertain | 0.816 | 0.254 | 0.000 | -12.417 | Likely Pathogenic | 0.911 | Likely Pathogenic | Ambiguous | 1.49 | Ambiguous | 0.3 | 1.34 | Ambiguous | 1.42 | Ambiguous | 0.08 | Likely Benign | 0.641 | Likely Pathogenic | -6.56 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | -1.26 | Pathogenic | 0.01 | Affected | 0.3422 | 0.5266 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.2186A>G | N729S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N729S is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). No tool in the dataset predicts a pathogenic outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus also as benign, while Foldetta (combining FoldX‑MD and Rosetta outputs) is inconclusive. Based on the collective predictions, the variant is most likely benign, and this assessment does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | 0.750527 | Disordered | 0.426547 | Uncertain | 0.651 | 0.583 | 0.625 | Uncertain | 1 | -1.578 | Likely Benign | 0.066 | Likely Benign | Likely Benign | 0.14 | Likely Benign | 0.1 | 1.34 | Ambiguous | 0.74 | Ambiguous | -0.36 | Likely Benign | 0.063 | Likely Benign | -0.42 | Neutral | 0.221 | Benign | 0.027 | Benign | 3.38 | Benign | 0.93 | Tolerated | 3.59 | 7 | 0.3411 | 0.4854 | 1 | 1 | 2.7 | -27.03 | ||||||||||||||||||||||||||
| c.764A>G | D255G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D255G missense variant is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that agree on a benign effect include premPS and FATHMM, while the majority of tools (REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus) predict a pathogenic impact. FoldX, Rosetta, and Foldetta provide uncertain or inconclusive results and are therefore not used as evidence. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta as uncertain. Based on the collective predictions, the variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | 0.501700 | Disordered | 0.219132 | Uncertain | 0.801 | 0.273 | 0.250 | -12.652 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 1.72 | Ambiguous | 0.1 | 1.34 | Ambiguous | 1.53 | Ambiguous | 0.31 | Likely Benign | 0.885 | Likely Pathogenic | -6.13 | Deleterious | 0.997 | Probably Damaging | 0.989 | Probably Damaging | 5.86 | Benign | 0.01 | Affected | 0.3211 | 0.5185 | 1 | -1 | 3.1 | -58.04 | ||||||||||||||||||||||||||||||
| c.1193C>G | P398R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P398R variant has no ClinVar entry and is not reported in gnomAD. Prediction tools that classify it as benign include only FATHMM. All other evaluated predictors—SGM‑Consensus, REVEL, FoldX, Foldetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default—indicate a pathogenic effect, while Rosetta, premPS, and AlphaMissense‑Optimized are uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as pathogenic. Based on the preponderance of pathogenic predictions and the high‑accuracy consensus, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.436924 | Structured | 0.401041 | Uncertain | 0.891 | 0.525 | 0.250 | -9.575 | Likely Pathogenic | 0.889 | Likely Pathogenic | Ambiguous | 3.01 | Destabilizing | 0.5 | 1.35 | Ambiguous | 2.18 | Destabilizing | 0.98 | Ambiguous | 0.755 | Likely Pathogenic | -6.55 | Deleterious | 0.988 | Probably Damaging | 0.724 | Possibly Damaging | 5.49 | Benign | 0.00 | Affected | 0.1479 | 0.3472 | 0 | -2 | -2.9 | 59.07 | |||||||||||||||||||||||||||||
| c.1348G>T | A450S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 A450S missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, SIFT, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and ESM1b. The remaining methods—FoldX, Rosetta, Foldetta, and premPS—return uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is a 2‑vs‑2 tie and thus unavailable; Foldetta is uncertain. Overall, the balance of evidence (five benign versus four pathogenic predictions, with three uncertain) suggests the variant is most likely benign. This conclusion does not contradict ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.321458 | Structured | 0.306281 | Uncertain | 0.963 | 0.234 | 0.000 | -9.257 | Likely Pathogenic | 0.274 | Likely Benign | Likely Benign | 0.81 | Ambiguous | 0.0 | 1.35 | Ambiguous | 1.08 | Ambiguous | 0.69 | Ambiguous | 0.268 | Likely Benign | -2.70 | Deleterious | 0.965 | Probably Damaging | 0.972 | Probably Damaging | 3.47 | Benign | 0.10 | Tolerated | 0.2003 | 0.4322 | 1 | 1 | -2.6 | 16.00 | ||||||||||||||||||||||||||||||
| c.1855A>T | T619S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant T619S is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Prediction tools that agree on a benign effect include only AlphaMissense‑Optimized. All other evaluated algorithms—SGM‑Consensus (Likely Pathogenic), REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default—consistently predict a pathogenic impact. High‑accuracy assessments further support this view: AlphaMissense‑Optimized reports a benign outcome, whereas the SGM Consensus, derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, indicates pathogenicity. Foldetta, which integrates FoldX‑MD and Rosetta stability predictions, yields an uncertain result. Overall, the majority of evidence points to a pathogenic effect for T619S, and this conclusion does not contradict the ClinVar designation of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.219301 | Structured | 0.119723 | Uncertain | 0.929 | 0.237 | 0.000 | Uncertain | 1 | -8.608 | Likely Pathogenic | 0.677 | Likely Pathogenic | Likely Benign | 1.09 | Ambiguous | 0.2 | 1.35 | Ambiguous | 1.22 | Ambiguous | 0.85 | Ambiguous | 0.602 | Likely Pathogenic | -3.42 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | -1.30 | Pathogenic | 0.05 | Affected | 3.37 | 35 | 0.3255 | 0.2860 | 1 | 1 | -0.1 | -14.03 | |||||||||||||||||||||||||
| c.1856C>G | T619S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 T619S missense variant has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include only AlphaMissense‑Optimized. All other evaluated predictors (SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default) uniformly predict a pathogenic impact. High‑accuracy assessments further support this: AlphaMissense‑Optimized remains benign, while the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—labels the variant as Likely Pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, yields an uncertain result. No other high‑confidence stability predictions are available. Overall, the preponderance of evidence indicates that the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status, as none is assigned. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.219301 | Structured | 0.119723 | Uncertain | 0.929 | 0.237 | 0.000 | -8.608 | Likely Pathogenic | 0.677 | Likely Pathogenic | Likely Benign | 1.09 | Ambiguous | 0.2 | 1.35 | Ambiguous | 1.22 | Ambiguous | 0.85 | Ambiguous | 0.523 | Likely Pathogenic | -3.42 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | -1.30 | Pathogenic | 0.05 | Affected | 3.37 | 35 | 0.3255 | 0.2860 | 1 | 1 | -0.1 | -14.03 | |||||||||||||||||||||||||||
| c.1990T>A | L664M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L664M is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, PROVEAN, and FATHMM, while those that agree on a pathogenic effect are polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. High‑accuracy assessments are mixed: AlphaMissense‑Optimized is uncertain; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is also uncertain due to a 2‑vs‑2 split; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, is uncertain. Overall, the majority of conventional tools predict a pathogenic impact, whereas the high‑accuracy methods do not provide a definitive verdict. Consequently, the variant is most likely pathogenic based on the available predictions, and this assessment does not contradict any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.100716 | Structured | 0.089318 | Uncertain | 0.937 | 0.339 | 0.000 | -11.819 | Likely Pathogenic | 0.807 | Likely Pathogenic | Ambiguous | 0.49 | Likely Benign | 0.1 | 1.35 | Ambiguous | 0.92 | Ambiguous | 0.96 | Ambiguous | 0.429 | Likely Benign | -2.00 | Neutral | 1.000 | Probably Damaging | 0.996 | Probably Damaging | 2.92 | Benign | 0.01 | Affected | 0.0620 | 0.2541 | 4 | 2 | -1.9 | 18.03 | ||||||||||||||||||||||||||||||
| c.2048T>C | I683T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I683T has no ClinVar record and is not reported in gnomAD. Prediction tools that agree on a benign effect include SIFT, FATHMM, and AlphaMissense‑Optimized, whereas a majority of tools predict pathogenicity: SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and AlphaMissense‑Default. High‑accuracy assessments further support this pattern: AlphaMissense‑Optimized classifies the variant as benign, while the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates it is likely pathogenic; the Foldetta stability analysis is inconclusive and therefore unavailable. Taken together, the preponderance of evidence points to a pathogenic effect for I683T. This conclusion does not contradict ClinVar status, which currently contains no classification for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.200174 | Structured | 0.143268 | Uncertain | 0.848 | 0.314 | 0.000 | -9.891 | Likely Pathogenic | 0.775 | Likely Pathogenic | Likely Benign | 1.67 | Ambiguous | 0.1 | 1.35 | Ambiguous | 1.51 | Ambiguous | 1.25 | Destabilizing | 0.548 | Likely Pathogenic | -4.77 | Deleterious | 0.999 | Probably Damaging | 0.981 | Probably Damaging | 3.29 | Benign | 0.08 | Tolerated | 0.1090 | 0.0880 | 0 | -1 | -5.2 | -12.05 | |||||||||||||||||||||||||||||
| c.629A>T | H210L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 H210L missense variant is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools cluster into two groups: benign predictions come from REVEL, Foldetta, premPS, and FATHMM, while pathogenic predictions arise from SGM‑Consensus (Likely Pathogenic), PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. FoldX and Rosetta provide uncertain results. High‑accuracy assessments further highlight the discrepancy: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates Likely Pathogenic, whereas Foldetta (a protein‑folding stability method combining FoldX‑MD and Rosetta outputs) predicts benign. Overall, the majority of evidence points to a pathogenic effect, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.144935 | Structured | 0.390904 | Uncertain | 0.872 | 0.298 | 0.125 | -14.516 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | -0.71 | Ambiguous | 0.1 | 1.35 | Ambiguous | 0.32 | Likely Benign | 0.49 | Likely Benign | 0.421 | Likely Benign | -9.41 | Deleterious | 0.895 | Possibly Damaging | 0.614 | Possibly Damaging | 3.09 | Benign | 0.00 | Affected | 0.0617 | 0.4452 | -2 | -3 | 7.0 | -23.98 | |||||||||||||||||||||||||||||
| c.644G>C | G215A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G215A is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a pathogenic effect include SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The only tool predicting a benign outcome is FATHMM. Predictions that are uncertain or inconclusive are Foldetta, Rosetta, and premPS. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; and Foldetta is uncertain. Overall, the majority of evidence indicates that G215A is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.155435 | Structured | 0.382818 | Uncertain | 0.791 | 0.291 | 0.000 | -8.930 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 2.36 | Destabilizing | 0.2 | 1.35 | Ambiguous | 1.86 | Ambiguous | 0.59 | Ambiguous | 0.874 | Likely Pathogenic | -5.08 | Deleterious | 0.999 | Probably Damaging | 0.995 | Probably Damaging | 5.61 | Benign | 0.02 | Affected | 0.3897 | 0.5525 | 1 | 0 | 2.2 | 14.03 | |||||||||||||||||||||||||||||
| c.965C>A | A322D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 A322D missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and AlphaMissense‑Optimized. Only FATHMM predicts a pathogenic outcome. The remaining tools—FoldX, Rosetta, ESM1b, and AlphaMissense‑Default—return uncertain results and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive, and Foldetta indicates no significant stability change (uncertain). Overall, the preponderance of evidence (seven benign predictions versus one pathogenic) suggests the variant is most likely benign, and this conclusion does not contradict any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.175930 | Structured | 0.425745 | Uncertain | 0.938 | 0.334 | 0.000 | -7.184 | In-Between | 0.411 | Ambiguous | Likely Benign | 0.65 | Ambiguous | 0.1 | 1.35 | Ambiguous | 1.00 | Ambiguous | 0.34 | Likely Benign | 0.246 | Likely Benign | -0.95 | Neutral | 0.270 | Benign | 0.136 | Benign | 1.96 | Pathogenic | 0.32 | Tolerated | 0.1646 | 0.1660 | 0 | -2 | -5.3 | 44.01 | ||||||||||||||||||||||||||||||
| c.1330A>C | K444Q 2D  3DClick to see structure in 3D Viewer AISynGAP1 K444Q is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions from REVEL, SIFT, and FATHMM; pathogenic predictions from premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also indicates a likely pathogenic outcome. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, and the SGM‑Consensus (majority vote) is pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is inconclusive. Overall, the preponderance of evidence points to a pathogenic effect for K444Q, and this conclusion does not conflict with any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.203355 | Structured | 0.262172 | Uncertain | 0.955 | 0.213 | 0.000 | -12.876 | Likely Pathogenic | 0.992 | Likely Pathogenic | Likely Pathogenic | 1.34 | Ambiguous | 0.0 | 1.36 | Ambiguous | 1.35 | Ambiguous | 1.04 | Destabilizing | 0.382 | Likely Benign | -3.82 | Deleterious | 0.998 | Probably Damaging | 0.997 | Probably Damaging | 3.43 | Benign | 0.07 | Tolerated | 0.4112 | 0.1057 | 1 | 1 | 0.4 | -0.04 | |||||||||||||||||||||||||||||
| c.1360A>C | I454L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I454L is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, and FATHMM, while polyPhen‑2 (HumDiv and HumVar) and AlphaMissense‑Default predict a pathogenic impact. The remaining tools—FoldX, Rosetta, Foldetta, premPS, ESM1b, and AlphaMissense‑Optimized—yield uncertain or inconclusive results and are treated as unavailable evidence. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) leaning toward benign, and Foldetta also uncertain. Overall, the majority of reliable predictors and the SGM Consensus favor a benign classification. Thus, the variant is most likely benign, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.254060 | Structured | 0.312811 | Uncertain | 0.965 | 0.182 | 0.000 | -7.852 | In-Between | 0.786 | Likely Pathogenic | Ambiguous | 0.57 | Ambiguous | 0.1 | 1.36 | Ambiguous | 0.97 | Ambiguous | 0.80 | Ambiguous | 0.246 | Likely Benign | -1.98 | Neutral | 0.908 | Possibly Damaging | 0.943 | Probably Damaging | 3.51 | Benign | 0.26 | Tolerated | 0.0653 | 0.2857 | 2 | 2 | -0.7 | 0.00 | ||||||||||||||||||||||||||||||
| c.1360A>G | I454V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I454V is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and AlphaMissense‑Default. Predictions that are inconclusive or unavailable are FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus as likely benign, and Foldetta as uncertain. Based on the collective evidence, the variant is most likely benign, and this conclusion does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.254060 | Structured | 0.312811 | Uncertain | 0.965 | 0.182 | 0.000 | -4.719 | Likely Benign | 0.657 | Likely Pathogenic | Likely Benign | 1.47 | Ambiguous | 0.0 | 1.36 | Ambiguous | 1.42 | Ambiguous | 0.69 | Ambiguous | 0.132 | Likely Benign | -0.79 | Neutral | 0.935 | Possibly Damaging | 0.858 | Possibly Damaging | 3.40 | Benign | 0.18 | Tolerated | 0.0833 | 0.2959 | 4 | 3 | -0.3 | -14.03 | |||||||||||||||||||||||||||||
| c.1819C>G | L607V 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L607V is listed in ClinVar with an uncertain significance (ClinVar ID 1450275.0) and is present in gnomAD (ID 6‑33440871‑C‑G). Prediction tools that agree on a benign effect include only AlphaMissense‑Optimized. All other evaluated algorithms—REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN)—predict a pathogenic impact. High‑accuracy assessments further support this: AlphaMissense‑Optimized reports benign, whereas the SGM‑Consensus, derived from the majority of pathogenic predictions, indicates pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta outputs, is inconclusive and therefore not considered evidence. Overall, the preponderance of computational evidence points to a pathogenic effect for L607V, a conclusion that contrasts with the current ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.048328 | Structured | 0.194229 | Uncertain | 0.869 | 0.250 | 0.000 | Uncertain | 2 | 6-33440871-C-G | 2 | 1.24e-6 | -11.190 | Likely Pathogenic | 0.637 | Likely Pathogenic | Likely Benign | 1.04 | Ambiguous | 0.2 | 1.36 | Ambiguous | 1.20 | Ambiguous | 0.90 | Ambiguous | 0.715 | Likely Pathogenic | -2.99 | Deleterious | 0.985 | Probably Damaging | 0.992 | Probably Damaging | -1.50 | Pathogenic | 0.01 | Affected | 3.37 | 35 | 0.1634 | 0.3577 | 2 | 1 | 0.4 | -14.03 | 216.3 | 28.1 | 0.1 | 0.0 | 0.9 | 0.2 | X | Potentially Benign | Leu607 is located in a short helical region (res. Ser606-Phe608) within an α-α loop connecting two α helices (res. Glu582-Met603 and res. Glu617-Asn635). In the WT simulations, the iso-butyl side chain of Leu607 does not interact with any other residues, but it could potentially interact directly with Ras due to its location at the GAP domain.In the variant simulations, Val607, which has similar size and physicochemical properties to leucine, does not cause any negative effects on the protein structure. However, due to its location at the GAP-Ras interface, the residue swap could affect the complex formation with the GTPase, but this cannot be investigated using solvent-only simulations. | |||||||||||||
| c.1833G>A | M611I 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant M611I is reported in gnomAD (ID 6‑33440885‑G‑A) but has no ClinVar entry. Functional prediction tools cluster into two groups: benign predictions come from REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and AlphaMissense‑Optimized; pathogenic predictions arise from SGM‑Consensus, ESM1b, FATHMM, and AlphaMissense‑Default. High‑accuracy assessments further clarify the variant’s likely effect: AlphaMissense‑Optimized classifies it as benign, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates pathogenicity, and Foldetta, which integrates FoldX‑MD and Rosetta outputs, yields an uncertain stability change. No folding‑stability method provides definitive evidence. Overall, the majority of predictions lean toward pathogenicity, and this conclusion does not conflict with ClinVar status, which lacks an entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.236433 | Structured | 0.210791 | Uncertain | 0.870 | 0.253 | 0.000 | 6-33440885-G-A | 1 | 6.19e-7 | -8.552 | Likely Pathogenic | 0.736 | Likely Pathogenic | Likely Benign | 1.45 | Ambiguous | 0.4 | 1.36 | Ambiguous | 1.41 | Ambiguous | 0.72 | Ambiguous | 0.292 | Likely Benign | -2.10 | Neutral | 0.250 | Benign | 0.091 | Benign | -1.14 | Pathogenic | 0.38 | Tolerated | 3.37 | 35 | 0.1009 | 0.2302 | 1 | 2 | 2.6 | -18.03 | ||||||||||||||||||||||||
| c.1833G>C | M611I 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant M611I is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and AlphaMissense‑Optimized. Tools that agree on a pathogenic effect are SGM‑Consensus, ESM1b, FATHMM, and AlphaMissense‑Default. Predictions that are inconclusive are FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized predicts a benign outcome, while the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta outputs, is uncertain. Overall, the majority of evidence points toward a pathogenic classification, and this assessment does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.236433 | Structured | 0.210791 | Uncertain | 0.870 | 0.253 | 0.000 | -8.552 | Likely Pathogenic | 0.736 | Likely Pathogenic | Likely Benign | 1.45 | Ambiguous | 0.4 | 1.36 | Ambiguous | 1.41 | Ambiguous | 0.72 | Ambiguous | 0.292 | Likely Benign | -2.10 | Neutral | 0.250 | Benign | 0.091 | Benign | -1.14 | Pathogenic | 0.38 | Tolerated | 3.37 | 35 | 0.1009 | 0.2302 | 1 | 2 | 2.6 | -18.03 | |||||||||||||||||||||||||||
| c.1833G>T | M611I 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant M611I is not reported in ClinVar and is absent from gnomAD. In silico predictors that classify the variant as benign include REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and AlphaMissense‑Optimized. Predictors that classify it as pathogenic are SGM‑Consensus, ESM1b, FATHMM, and AlphaMissense‑Default. Four tools (FoldX, Rosetta, premPS, and Foldetta) provide uncertain or unavailable results. High‑accuracy assessment shows AlphaMissense‑Optimized predicts benign, while the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts pathogenic; Foldetta’s stability output is unavailable. Overall, the majority of predictions lean toward a benign effect, and this conclusion does not contradict the ClinVar status, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.236433 | Structured | 0.210791 | Uncertain | 0.870 | 0.253 | 0.000 | -8.552 | Likely Pathogenic | 0.736 | Likely Pathogenic | Likely Benign | 1.45 | Ambiguous | 0.4 | 1.36 | Ambiguous | 1.41 | Ambiguous | 0.72 | Ambiguous | 0.292 | Likely Benign | -2.10 | Neutral | 0.250 | Benign | 0.091 | Benign | -1.14 | Pathogenic | 0.38 | Tolerated | 3.37 | 35 | 0.1009 | 0.2302 | 1 | 2 | 2.6 | -18.03 | |||||||||||||||||||||||||||
| c.1864A>T | T622S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant T622S is not reported in ClinVar and is absent from gnomAD. Prediction tools cluster into two groups: benign predictions come from SIFT and AlphaMissense‑Optimized, while pathogenic predictions arise from REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). High‑accuracy assessments show AlphaMissense‑Optimized labeling the variant as benign, whereas the SGM‑Consensus predicts it to be likely pathogenic; Foldetta, which integrates FoldX‑MD and Rosetta outputs, yields an uncertain result. No other folding‑stability tools provide conclusive evidence. Overall, the preponderance of predictions, including the SGM‑Consensus, indicates that T622S is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.268042 | Structured | 0.071403 | Uncertain | 0.957 | 0.198 | 0.000 | -9.840 | Likely Pathogenic | 0.669 | Likely Pathogenic | Likely Benign | 0.78 | Ambiguous | 0.1 | 1.36 | Ambiguous | 1.07 | Ambiguous | 0.80 | Ambiguous | 0.705 | Likely Pathogenic | -3.52 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | -1.50 | Pathogenic | 0.09 | Tolerated | 0.2523 | 0.3183 | 1 | 1 | -0.1 | -14.03 | |||||||||||||||||||||||||||||
| c.1865C>G | T622S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant T622S is not reported in ClinVar and is absent from gnomAD. Prediction tools cluster into two groups: benign predictions come from SIFT and AlphaMissense‑Optimized, while pathogenic predictions arise from REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). High‑accuracy assessments show AlphaMissense‑Optimized labeling the variant as benign, whereas the SGM‑Consensus predicts it to be likely pathogenic; Foldetta, which integrates FoldX‑MD and Rosetta outputs, yields an uncertain result. No other folding‑stability tools provide conclusive evidence. Overall, the preponderance of predictions, including the SGM‑Consensus, indicates that T622S is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.268042 | Structured | 0.071403 | Uncertain | 0.957 | 0.198 | 0.000 | -9.840 | Likely Pathogenic | 0.669 | Likely Pathogenic | Likely Benign | 0.78 | Ambiguous | 0.1 | 1.36 | Ambiguous | 1.07 | Ambiguous | 0.80 | Ambiguous | 0.595 | Likely Pathogenic | -3.52 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | -1.50 | Pathogenic | 0.09 | Tolerated | 0.2523 | 0.3183 | 1 | 1 | -0.1 | -14.03 | |||||||||||||||||||||||||||||
| c.2003C>G | S668C 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant S668C is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split opinion: benign predictions come from premPS, FATHMM, and AlphaMissense‑Optimized, whereas pathogenic predictions are returned by REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and the SGM‑Consensus. Uncertain results are reported by FoldX, Rosetta, and Foldetta. High‑accuracy assessments indicate that AlphaMissense‑Optimized predicts a benign effect, while the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) supports a pathogenic outcome; Foldetta remains inconclusive. Overall, the majority of evidence points toward a pathogenic impact, and this conclusion does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.247041 | Structured | 0.084935 | Uncertain | 0.922 | 0.370 | 0.000 | -12.815 | Likely Pathogenic | 0.758 | Likely Pathogenic | Likely Benign | 1.31 | Ambiguous | 0.6 | 1.36 | Ambiguous | 1.34 | Ambiguous | 0.18 | Likely Benign | 0.503 | Likely Pathogenic | -4.99 | Deleterious | 0.999 | Probably Damaging | 0.944 | Probably Damaging | 3.27 | Benign | 0.02 | Affected | 0.0962 | 0.5528 | 0 | -1 | 3.3 | 16.06 | |||||||||||||||||||||||||||||
| c.701G>C | R234P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R234P is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include premPS and FATHMM, whereas the majority of evaluated algorithms (REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus) predict a pathogenic impact. Uncertain results are reported by FoldX, Rosetta, and Foldetta. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as likely pathogenic, and Foldetta as inconclusive. Based on the collective evidence, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.239899 | Structured | 0.311558 | Uncertain | 0.804 | 0.322 | 0.000 | -10.126 | Likely Pathogenic | 0.972 | Likely Pathogenic | Likely Pathogenic | 1.33 | Ambiguous | 0.6 | 1.36 | Ambiguous | 1.35 | Ambiguous | 0.13 | Likely Benign | 0.826 | Likely Pathogenic | -4.43 | Deleterious | 0.929 | Possibly Damaging | 0.519 | Possibly Damaging | 5.93 | Benign | 0.04 | Affected | 0.1972 | 0.4336 | 0 | -2 | 2.9 | -59.07 | |||||||||||||||||||||||||||||
| c.721A>G | K241E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 K241E missense variant has no ClinVar entry and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include SIFT and FATHMM, whereas a majority of tools predict a pathogenic impact: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Predictions that are inconclusive or unavailable are FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments further support a deleterious interpretation: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is also pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, remains uncertain. Overall, the preponderance of evidence from multiple independent predictors indicates that the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status because no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.196879 | Structured | 0.349250 | Uncertain | 0.797 | 0.347 | 0.000 | -12.582 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 0.75 | Ambiguous | 0.1 | 1.36 | Ambiguous | 1.06 | Ambiguous | 0.68 | Ambiguous | 0.878 | Likely Pathogenic | -3.43 | Deleterious | 0.982 | Probably Damaging | 0.679 | Possibly Damaging | 6.00 | Benign | 0.10 | Tolerated | 0.3610 | 0.1179 | 0 | 1 | 0.4 | 0.94 | |||||||||||||||||||||||||||||
| c.1549C>G | L517V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L517V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree that the change is deleterious: pathogenic predictions come from SGM‑Consensus, REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and FATHMM, while benign predictions are made by SIFT and AlphaMissense‑Optimized; the remaining tools (Rosetta, Foldetta, AlphaMissense‑Default) are uncertain. High‑accuracy assessments further support a pathogenic bias: AlphaMissense‑Optimized predicts benign, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is uncertain. Overall, the preponderance of evidence points to a likely pathogenic effect of the variant, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.118441 | Structured | 0.147645 | Uncertain | 0.938 | 0.296 | 0.000 | -10.691 | Likely Pathogenic | 0.498 | Ambiguous | Likely Benign | 2.04 | Destabilizing | 0.3 | 1.37 | Ambiguous | 1.71 | Ambiguous | 1.14 | Destabilizing | 0.577 | Likely Pathogenic | -2.68 | Deleterious | 0.998 | Probably Damaging | 0.992 | Probably Damaging | -1.26 | Pathogenic | 0.17 | Tolerated | 0.1405 | 0.2398 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.1693C>G | L565V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L565V has no ClinVar entry and is not reported in gnomAD. Prediction tools cluster into two consensus groups: benign predictions come from REVEL and FATHMM, while pathogenic predictions are made by SGM‑Consensus, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and Foldetta. Two tools (AlphaMissense‑Optimized and Rosetta) give uncertain results. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is inconclusive, SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) reports “Likely Pathogenic,” and Foldetta, which integrates FoldX‑MD and Rosetta outputs, predicts a pathogenic impact on protein stability. Overall, the preponderance of evidence indicates that the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status because none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.034884 | Structured | 0.045819 | Uncertain | 0.922 | 0.205 | 0.000 | -11.118 | Likely Pathogenic | 0.833 | Likely Pathogenic | Ambiguous | 2.69 | Destabilizing | 0.0 | 1.37 | Ambiguous | 2.03 | Destabilizing | 1.40 | Destabilizing | 0.303 | Likely Benign | -2.99 | Deleterious | 0.996 | Probably Damaging | 0.992 | Probably Damaging | 2.87 | Benign | 0.03 | Affected | 0.1558 | 0.2488 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.2152C>A | L718I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L718I is not reported in ClinVar and is absent from gnomAD. Prediction tools cluster into two groups: benign predictions come from REVEL, PROVEAN, and AlphaMissense‑Optimized; pathogenic predictions come from SGM‑Consensus (Likely Pathogenic), FoldX, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. Three tools (Foldetta, premPS, Rosetta) give uncertain results and are not considered evidence. High‑accuracy methods specifically show AlphaMissense‑Optimized as benign, SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. Because the majority of reliable predictors (eight out of eleven) indicate pathogenicity, the variant is most likely pathogenic. This assessment does not contradict ClinVar, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.298791 | Structured | 0.438417 | Uncertain | 0.966 | 0.385 | 0.000 | -10.560 | Likely Pathogenic | 0.615 | Likely Pathogenic | Likely Benign | 2.21 | Destabilizing | 0.2 | 1.37 | Ambiguous | 1.79 | Ambiguous | 0.89 | Ambiguous | 0.296 | Likely Benign | -1.90 | Neutral | 0.999 | Probably Damaging | 0.997 | Probably Damaging | 1.37 | Pathogenic | 0.00 | Affected | 0.0876 | 0.3206 | 2 | 2 | 0.7 | 0.00 | |||||||||||||||||||||||||||||
| c.1078G>C | E360Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E360Q is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL and premPS, while a majority of tools predict a pathogenic outcome: SIFT, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, and AlphaMissense‑Default. Uncertain or inconclusive results come from FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (derived from the unanimous pathogenic vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the preponderance of evidence points to a pathogenic effect for E360Q, and this conclusion does not contradict any ClinVar annotation, as none exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.250310 | Structured | 0.421183 | Uncertain | 0.955 | 0.498 | 0.250 | -11.012 | Likely Pathogenic | 0.925 | Likely Pathogenic | Ambiguous | 0.55 | Ambiguous | 0.1 | 1.38 | Ambiguous | 0.97 | Ambiguous | -0.02 | Likely Benign | 0.343 | Likely Benign | -2.76 | Deleterious | 0.997 | Probably Damaging | 0.986 | Probably Damaging | 1.61 | Pathogenic | 0.03 | Affected | 0.1824 | 0.8282 | 2 | 2 | 0.0 | -0.98 | |||||||||||||||||||||||||||||
| c.1528A>T | I510F 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I510F is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include premPS and AlphaMissense‑Optimized, whereas the majority of tools (SGM‑Consensus, REVEL, FoldX, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and Foldetta) predict a pathogenic impact; Rosetta remains uncertain. High‑accuracy assessments further support this: AlphaMissense‑Optimized classifies the variant as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts pathogenicity, and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenicity. Overall, the preponderance of evidence indicates that the variant is most likely pathogenic, and this conclusion does not contradict any existing ClinVar annotation, as none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.025762 | Structured | 0.250630 | Uncertain | 0.945 | 0.273 | 0.000 | -8.185 | Likely Pathogenic | 0.713 | Likely Pathogenic | Likely Benign | 4.66 | Destabilizing | 0.7 | 1.38 | Ambiguous | 3.02 | Destabilizing | 0.50 | Likely Benign | 0.692 | Likely Pathogenic | -2.64 | Deleterious | 0.991 | Probably Damaging | 0.854 | Possibly Damaging | -1.14 | Pathogenic | 0.01 | Affected | 0.0552 | 0.1794 | 1 | 0 | -1.7 | 34.02 | |||||||||||||||||||||||||||||
| c.869T>A | L290Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L290Q is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that agree on a benign effect include only SIFT, whereas the majority of tools predict a pathogenic impact: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (which itself is “Likely Pathogenic”). Uncertain or inconclusive results come from FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus is likely pathogenic, and Foldetta’s stability prediction is unavailable. Overall, the preponderance of evidence indicates that the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.127496 | Structured | 0.399723 | Uncertain | 0.904 | 0.255 | 0.000 | -12.776 | Likely Pathogenic | 0.993 | Likely Pathogenic | Likely Pathogenic | 1.12 | Ambiguous | 0.1 | 1.38 | Ambiguous | 1.25 | Ambiguous | 0.68 | Ambiguous | 0.697 | Likely Pathogenic | -5.52 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.97 | Pathogenic | 0.06 | Tolerated | 0.1079 | 0.1014 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.1440G>C | E480D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E480D missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, and ESM1b, whereas polyPhen‑2 (HumDiv and HumVar) and FATHMM predict a pathogenic impact. The remaining tools—FoldX, Rosetta, premPS, AlphaMissense‑Default, and Foldetta—return uncertain or inconclusive results and are treated as unavailable evidence. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized predicts benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) yields a benign majority, and Foldetta remains uncertain. Overall, the balance of evidence favors a benign classification for E480D, and this conclusion does not contradict the lack of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.216401 | Structured | 0.426867 | Uncertain | 0.798 | 0.250 | 0.000 | -3.001 | Likely Benign | 0.475 | Ambiguous | Likely Benign | 0.62 | Ambiguous | 0.2 | 1.39 | Ambiguous | 1.01 | Ambiguous | 0.61 | Ambiguous | 0.405 | Likely Benign | -0.77 | Neutral | 0.989 | Probably Damaging | 0.979 | Probably Damaging | -1.27 | Pathogenic | 0.28 | Tolerated | 0.1526 | 0.4130 | 3 | 2 | 0.0 | -14.03 | ||||||||||||||||||||||||||||||
| c.1440G>T | E480D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E480D missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, and ESM1b, while polyPhen‑2 (HumDiv and HumVar) and FATHMM predict a pathogenic outcome. The remaining tools—FoldX, Rosetta, premPS, AlphaMissense‑Default, and Foldetta—return uncertain or inconclusive results and are treated as unavailable evidence. High‑accuracy assessments show AlphaMissense‑Optimized classifying the variant as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) leaning toward benign, and Foldetta providing no definitive stability change. Overall, the balance of evidence favors a benign interpretation, and this conclusion does not contradict the lack of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.216401 | Structured | 0.426867 | Uncertain | 0.798 | 0.250 | 0.000 | -3.001 | Likely Benign | 0.475 | Ambiguous | Likely Benign | 0.62 | Ambiguous | 0.2 | 1.39 | Ambiguous | 1.01 | Ambiguous | 0.61 | Ambiguous | 0.405 | Likely Benign | -0.77 | Neutral | 0.989 | Probably Damaging | 0.979 | Probably Damaging | -1.27 | Pathogenic | 0.28 | Tolerated | 0.1526 | 0.4130 | 3 | 2 | 0.0 | -14.03 | ||||||||||||||||||||||||||||||
| c.1636T>A | C546S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C546S is reported in gnomAD (ID 6‑33438879‑T‑A) but has no ClinVar entry. Functional prediction tools cluster into two groups: benign predictions from FoldX and SIFT; pathogenic predictions from REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Two tools give uncertain results: Rosetta and Foldetta. High‑accuracy assessments reinforce a pathogenic signal: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—labels the variant as Likely Pathogenic; Foldetta, which integrates FoldX‑MD and Rosetta outputs, remains inconclusive. Overall, the preponderance of evidence, including the high‑accuracy tools, indicates that C546S is most likely pathogenic, and this assessment does not conflict with any ClinVar classification because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.027463 | Structured | 0.009041 | Uncertain | 0.960 | 0.288 | 0.000 | 6-33438879-T-A | 1 | 6.20e-7 | -8.079 | Likely Pathogenic | 0.988 | Likely Pathogenic | Likely Pathogenic | 0.44 | Likely Benign | 0.1 | 1.39 | Ambiguous | 0.92 | Ambiguous | 1.65 | Destabilizing | 0.836 | Likely Pathogenic | -8.04 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.21 | Pathogenic | 0.17 | Tolerated | 3.37 | 35 | 0.4343 | 0.1950 | -1 | 0 | -3.3 | -16.06 | ||||||||||||||||||||||||
| c.1637G>C | C546S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C546S is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include FoldX and SIFT, whereas the majority of tools predict a pathogenic impact: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Rosetta and Foldetta are uncertain, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates a likely pathogenic outcome. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, SGM Consensus as likely pathogenic, and Foldetta as uncertain. Overall, the preponderance of evidence points to a pathogenic effect for C546S, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.027463 | Structured | 0.009041 | Uncertain | 0.960 | 0.288 | 0.000 | -8.079 | Likely Pathogenic | 0.988 | Likely Pathogenic | Likely Pathogenic | 0.44 | Likely Benign | 0.1 | 1.39 | Ambiguous | 0.92 | Ambiguous | 1.65 | Destabilizing | 0.788 | Likely Pathogenic | -8.04 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.21 | Pathogenic | 0.17 | Tolerated | 3.37 | 35 | 0.4343 | 0.1950 | -1 | 0 | -3.3 | -16.06 | |||||||||||||||||||||||||||
| c.1739G>A | G580D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G580D is not reported in ClinVar and has no entries in gnomAD. Consensus from multiple in silico predictors indicates a pathogenic effect: SGM‑Consensus, REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify it as pathogenic, while premPS and Rosetta are uncertain. High‑accuracy tools reinforce this assessment: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenic. No benign predictions are present. Consequently, the variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.104810 | Structured | 0.025952 | Uncertain | 0.853 | 0.236 | 0.000 | -10.086 | Likely Pathogenic | 0.965 | Likely Pathogenic | Likely Pathogenic | 2.85 | Destabilizing | 0.1 | 1.39 | Ambiguous | 2.12 | Destabilizing | 0.83 | Ambiguous | 0.712 | Likely Pathogenic | -6.73 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | -1.25 | Pathogenic | 0.04 | Affected | 0.1793 | 0.1971 | 1 | -1 | -3.1 | 58.04 | |||||||||||||||||||||||||||||
| c.1822T>G | F608V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F608V is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that agree on a pathogenic effect include REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized; all of these return a pathogenic or likely pathogenic label. No tool in the dataset predicts a benign outcome. Uncertain results are reported only for Rosetta and Foldetta, which are treated as unavailable evidence. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta remains uncertain. Taken together, the overwhelming majority of predictions indicate that F608V is most likely pathogenic, and this conclusion does not contradict any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.106997 | Structured | 0.197190 | Uncertain | 0.891 | 0.247 | 0.000 | -16.084 | Likely Pathogenic | 0.982 | Likely Pathogenic | Likely Pathogenic | 2.21 | Destabilizing | 0.1 | 1.39 | Ambiguous | 1.80 | Ambiguous | 1.47 | Destabilizing | 0.917 | Likely Pathogenic | -6.97 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | -1.59 | Pathogenic | 0.01 | Affected | 0.2193 | 0.2199 | -1 | -1 | 1.4 | -48.04 | |||||||||||||||||||||||||||||
| c.623C>G | P208R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P208R missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are REVEL and FATHMM. The majority of other in silico predictors (FoldX, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) and the SGM‑Consensus score (Likely Pathogenic) all indicate a pathogenic impact, while Rosetta remains uncertain. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts pathogenic; the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. Based on the preponderance of pathogenic predictions and the absence of any benign consensus, the variant is most likely pathogenic, with no contradiction to ClinVar status because no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.271506 | Structured | 0.399506 | Uncertain | 0.864 | 0.345 | 0.125 | -11.929 | Likely Pathogenic | 0.966 | Likely Pathogenic | Likely Pathogenic | 5.25 | Destabilizing | 1.6 | 1.39 | Ambiguous | 3.32 | Destabilizing | 1.16 | Destabilizing | 0.472 | Likely Benign | -7.50 | Deleterious | 1.000 | Probably Damaging | 0.996 | Probably Damaging | 3.75 | Benign | 0.00 | Affected | 0.1676 | 0.3118 | 0 | -2 | -2.9 | 59.07 | |||||||||||||||||||||||||||||
| c.632G>A | S211N 2D  AIThe SynGAP1 S211N missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, and FATHMM, whereas those that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, and premPS. Four tools (FoldX, Rosetta, Foldetta, AlphaMissense‑Optimized) give uncertain or inconclusive results. High‑accuracy assessments are likewise inconclusive: AlphaMissense‑Optimized is uncertain; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is a 2‑vs‑2 tie, and Foldetta is uncertain. Consequently, the overall prediction leans toward pathogenicity, with no ClinVar entry to contradict this assessment. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.209395 | Structured | 0.389893 | Uncertain | 0.846 | 0.300 | 0.125 | -9.995 | Likely Pathogenic | 0.794 | Likely Pathogenic | Ambiguous | 0.60 | Ambiguous | 1.2 | 1.39 | Ambiguous | 1.00 | Ambiguous | 1.21 | Destabilizing | 0.174 | Likely Benign | -2.22 | Neutral | 0.982 | Probably Damaging | 0.747 | Possibly Damaging | 3.95 | Benign | 0.09 | Tolerated | 0.1534 | 0.4863 | 1 | 1 | -2.7 | 27.03 | ||||||||||||||||||||||||||||||
| c.745G>A | A249T 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant A249T is listed in ClinVar (ID 1031675.0) with an uncertain significance annotation and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, SIFT, ESM1b, and FATHMM, whereas polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and AlphaMissense‑Default predict a pathogenic outcome. Predictions that are inconclusive are FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely benign, and Foldetta (combining FoldX‑MD and Rosetta) as uncertain. Overall, the balance of evidence favors a benign interpretation, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | PH | 0.505461 | Disordered | 0.255452 | Uncertain | 0.810 | 0.336 | 0.125 | Uncertain | 1 | -3.564 | Likely Benign | 0.805 | Likely Pathogenic | Ambiguous | 1.50 | Ambiguous | 0.6 | 1.39 | Ambiguous | 1.45 | Ambiguous | 0.30 | Likely Benign | 0.487 | Likely Benign | -0.96 | Neutral | 0.990 | Probably Damaging | 0.815 | Possibly Damaging | 5.65 | Benign | 0.40 | Tolerated | 3.39 | 15 | 0.0909 | 0.4972 | 1 | 0 | -2.5 | 30.03 | 214.5 | -43.3 | 0.0 | 0.0 | 0.5 | 0.2 | X | Potentially Benign | The methyl group of Ala249, located on the surface of an α helix (res. Ala236-Val250) facing an anti-parallel β sheet strand (res. Ile205-Val209), packs against nearby hydrophobic residues such as Leu200, Leu246, and Val250. In the variant simulations, the hydroxyl group of Thr249, which is not suitable for hydrophobic packing, forms a stable hydrogen bond with the backbone carbonyl of Asn245 in the same helix. Although this interaction could theoretically weaken the structural integrity of the α helix, this destabilizing effect is not observed in the variant simulations. | ||||||||||||||||
| c.1784T>A | L595Q 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L595Q is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Prediction tools that classify the variant as benign include only FATHMM. All other evaluated algorithms—REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized—predict a pathogenic effect, and the SGM‑Consensus score indicates a likely pathogenic outcome. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized returns a pathogenic prediction, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also yields a likely pathogenic result, while Foldetta’s stability analysis is inconclusive. Overall, the majority of computational evidence points to a pathogenic effect, which does not contradict the ClinVar designation of uncertain significance but suggests a higher likelihood of pathogenicity. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.015344 | Structured | 0.128444 | Uncertain | 0.920 | 0.150 | 0.000 | Uncertain | 1 | -15.101 | Likely Pathogenic | 0.984 | Likely Pathogenic | Likely Pathogenic | 0.79 | Ambiguous | 0.1 | 1.40 | Ambiguous | 1.10 | Ambiguous | 1.99 | Destabilizing | 0.733 | Likely Pathogenic | -5.97 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 2.75 | Benign | 0.00 | Affected | 3.37 | 35 | 0.1074 | 0.1563 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||
| c.1994A>C | Y665S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant Y665S is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, SIFT, FATHMM, and AlphaMissense‑Optimized. Those that agree on a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and ESM1b. The remaining tools (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Default) give uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized predicts benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) leans pathogenic, and Foldetta is uncertain (treated as unavailable). Overall, the balance of evidence, particularly the pathogenic signal from the SGM Consensus and the equal split among standard predictors, indicates that the variant is most likely pathogenic. This conclusion does not contradict ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.098513 | Structured | 0.086641 | Uncertain | 0.922 | 0.361 | 0.000 | -9.110 | Likely Pathogenic | 0.453 | Ambiguous | Likely Benign | 1.24 | Ambiguous | 0.2 | 1.40 | Ambiguous | 1.32 | Ambiguous | 0.87 | Ambiguous | 0.202 | Likely Benign | -2.50 | Deleterious | 1.000 | Probably Damaging | 0.994 | Probably Damaging | 3.55 | Benign | 0.62 | Tolerated | 0.3995 | 0.1793 | -3 | -2 | 0.5 | -76.10 | ||||||||||||||||||||||||||||||
| c.937G>A | E313K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E313K is listed in ClinVar as Benign (ClinVar ID 3695040.0) and is not reported in gnomAD. Prediction tools that report a benign effect are absent; all available predictors that provide a definitive call classify the variant as pathogenic. These include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized predicts Pathogenic, the SGM‑Consensus indicates Likely Pathogenic, while Foldetta (combining FoldX‑MD and Rosetta outputs) is Uncertain. Based on the overwhelming pathogenic predictions, the variant is most likely pathogenic, which contradicts its ClinVar benign classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.170161 | Structured | 0.366526 | Uncertain | 0.898 | 0.304 | 0.125 | Likely Benign | 1 | -12.902 | Likely Pathogenic | 0.959 | Likely Pathogenic | Likely Pathogenic | 0.64 | Ambiguous | 0.6 | 1.40 | Ambiguous | 1.02 | Ambiguous | 0.75 | Ambiguous | 0.575 | Likely Pathogenic | -3.31 | Deleterious | 1.000 | Probably Damaging | 0.995 | Probably Damaging | 1.90 | Pathogenic | 0.02 | Affected | 0.2540 | 0.7708 | 0 | 1 | -0.4 | -0.94 | |||||||||||||||||||||||||||
| c.977A>G | H326R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 H326R missense variant is not listed in ClinVar and has no entry in gnomAD. Prediction tools that agree on a benign effect include SIFT and Foldetta, whereas the remaining tools (SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized) all predict a pathogenic impact. High‑accuracy methods give the following results: AlphaMissense‑Optimized classifies the variant as pathogenic; the SGM‑Consensus, which is a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports it as likely pathogenic; Foldetta, a protein‑folding stability approach that integrates FoldX‑MD and Rosetta outputs, predicts a benign effect. Because the majority of evidence points to a deleterious change, the variant is most likely pathogenic, and this assessment is not contradicted by the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.342579 | Structured | 0.418150 | Uncertain | 0.944 | 0.455 | 0.000 | -12.369 | Likely Pathogenic | 0.980 | Likely Pathogenic | Likely Pathogenic | -0.65 | Ambiguous | 0.1 | 1.40 | Ambiguous | 0.38 | Likely Benign | 0.83 | Ambiguous | 0.601 | Likely Pathogenic | -6.89 | Deleterious | 0.997 | Probably Damaging | 0.994 | Probably Damaging | 1.97 | Pathogenic | 0.10 | Tolerated | 0.1832 | 0.2413 | 2 | 0 | -1.3 | 19.05 | |||||||||||||||||||||||||||||
| c.1058T>A | L353Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 L353Q missense variant has no ClinVar entry and is absent from gnomAD. Among in‑silico predictors, only REVEL classifies it as benign, whereas the majority—FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus—label it pathogenic. Predictions marked uncertain include Rosetta, Foldetta, ESM1b, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Overall, the preponderance of pathogenic predictions outweighs the single benign call, and no ClinVar record contradicts this assessment. Thus, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.137348 | Structured | 0.373584 | Uncertain | 0.926 | 0.315 | 0.000 | -7.074 | In-Between | 0.884 | Likely Pathogenic | Ambiguous | 2.38 | Destabilizing | 0.2 | 1.41 | Ambiguous | 1.90 | Ambiguous | 2.00 | Destabilizing | 0.388 | Likely Benign | -3.56 | Deleterious | 0.947 | Possibly Damaging | 0.556 | Possibly Damaging | 1.30 | Pathogenic | 0.01 | Affected | 0.1098 | 0.1303 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.1106C>T | T369I 2D  AIThe SynGAP1 missense variant T369I is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote). Only FATHMM predicts a pathogenic outcome. Stability‑based methods (FoldX, Rosetta, Foldetta) are inconclusive, providing no definitive evidence. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus as likely benign, and Foldetta as uncertain. Overall, the majority of evidence points to a benign impact; there is no conflict with ClinVar status, which contains no entry for this variant. Thus, the variant is most likely benign based on current predictions. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.468512 | Structured | 0.437011 | Uncertain | 0.417 | 0.707 | 0.500 | -6.759 | Likely Benign | 0.289 | Likely Benign | Likely Benign | 0.60 | Ambiguous | 0.8 | 1.41 | Ambiguous | 1.01 | Ambiguous | -0.08 | Likely Benign | 0.078 | Likely Benign | -2.37 | Neutral | 0.396 | Benign | 0.142 | Benign | 1.72 | Pathogenic | 0.13 | Tolerated | 0.1106 | 0.7207 | 0 | -1 | 5.2 | 12.05 | |||||||||||||||||||||||||||||
| c.1256A>T | E419V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E419V missense variant is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign predictions come from REVEL, FoldX, FATHMM, and premPS, whereas pathogenic predictions arise from PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as Likely Pathogenic. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts Pathogenic, the SGM Consensus also indicates Likely Pathogenic, while Foldetta, which integrates FoldX‑MD and Rosetta outputs, yields an uncertain result. No evidence from the high‑accuracy tools contradicts the pathogenic prediction. Overall, the majority of computational evidence points to a pathogenic effect, which is consistent with the lack of ClinVar reporting and gnomAD absence, suggesting the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.102787 | Structured | 0.371949 | Uncertain | 0.961 | 0.261 | 0.000 | -12.290 | Likely Pathogenic | 0.980 | Likely Pathogenic | Likely Pathogenic | 0.45 | Likely Benign | 0.0 | 1.41 | Ambiguous | 0.93 | Ambiguous | 0.27 | Likely Benign | 0.494 | Likely Benign | -6.55 | Deleterious | 0.998 | Probably Damaging | 0.983 | Probably Damaging | 3.35 | Benign | 0.01 | Affected | 0.0953 | 0.6834 | -2 | -2 | 7.7 | -29.98 | |||||||||||||||||||||||||||||
| c.1258T>C | F420L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F420L is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and FATHMM. Tools that predict a pathogenic effect comprise the SGM‑Consensus (Likely Pathogenic), premPS, PROVEAN, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. FoldX, Rosetta, and Foldetta provide uncertain or inconclusive stability results and are treated as unavailable evidence. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. Overall, the majority of evaluated predictors (six pathogenic vs. five benign) indicate that F420L is most likely pathogenic, and this conclusion does not contradict any existing ClinVar annotation, as none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.104810 | Structured | 0.384475 | Uncertain | 0.974 | 0.255 | 0.000 | -8.432 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 1.76 | Ambiguous | 0.0 | 1.41 | Ambiguous | 1.59 | Ambiguous | 1.04 | Destabilizing | 0.262 | Likely Benign | -5.39 | Deleterious | 0.009 | Benign | 0.005 | Benign | 4.22 | Benign | 0.39 | Tolerated | 3.37 | 29 | 0.2053 | 0.3016 | 2 | 0 | 1.0 | -34.02 | 231.1 | 13.2 | 0.0 | 0.0 | -0.1 | 0.0 | X | Potentially Benign | In the WT, the phenyl ring of the Phe420 side chain, located on an α helix (res. Met414-Glu436), packs against hydrophobic residues in the interhelix area of the GAP domain (e.g., Leu689, Leu714, Leu717, Leu718). In the variant simulations, the iso-butyl side chain of Leu420 also packs into the hydrophobic inter-helix niche, but due to its smaller size, the resulting steric interactions are not as favorable as with phenylalanine. In short, the residue swap does not cause severe effects on the protein structure based on the variant simulations. | ||||||||||||||||||
| c.1260T>A | F420L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F420L is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and FATHMM. Tools that predict a pathogenic effect comprise the SGM‑Consensus (Likely Pathogenic), premPS, PROVEAN, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. FoldX, Rosetta, and Foldetta provide uncertain or inconclusive stability results and are treated as unavailable evidence. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. Overall, the majority of evaluated predictors (six pathogenic vs. five benign) indicate that F420L is most likely pathogenic, and this conclusion does not contradict any existing ClinVar annotation, as none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.104810 | Structured | 0.384475 | Uncertain | 0.974 | 0.255 | 0.000 | -8.432 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 1.76 | Ambiguous | 0.0 | 1.41 | Ambiguous | 1.59 | Ambiguous | 1.04 | Destabilizing | 0.146 | Likely Benign | -5.39 | Deleterious | 0.009 | Benign | 0.005 | Benign | 4.22 | Benign | 0.39 | Tolerated | 3.37 | 29 | 0.2053 | 0.3016 | 2 | 0 | 1.0 | -34.02 | 231.1 | 13.2 | 0.0 | 0.0 | -0.1 | 0.0 | X | Potentially Benign | In the WT, the phenyl ring of the Phe420 side chain, located on an α helix (res. Met414-Glu436), packs against hydrophobic residues in the interhelix area of the GAP domain (e.g., Leu689, Leu714, Leu717, Leu718). In the variant simulations, the iso-butyl side chain of Leu420 also packs into the hydrophobic inter-helix niche, but due to its smaller size, the resulting steric interactions are not as favorable as with phenylalanine. In short, the residue swap does not cause severe effects on the protein structure based on the variant simulations. | ||||||||||||||||||
| c.1260T>G | F420L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F420L is listed in ClinVar (ID 1397885.0) with an “Uncertain” clinical significance and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM. Those that predict a pathogenic effect comprise premPS, PROVEAN, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Stability‑based methods (FoldX, Rosetta, Foldetta) yield inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus also as pathogenic, while Foldetta remains uncertain. Overall, the majority of evidence points toward a pathogenic impact, which does not contradict the ClinVar “Uncertain” status but suggests the variant is more likely pathogenic rather than benign. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.104810 | Structured | 0.384475 | Uncertain | 0.974 | 0.255 | 0.000 | Uncertain | 1 | -8.432 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 1.76 | Ambiguous | 0.0 | 1.41 | Ambiguous | 1.59 | Ambiguous | 1.04 | Destabilizing | 0.146 | Likely Benign | -5.39 | Deleterious | 0.009 | Benign | 0.005 | Benign | 4.22 | Benign | 0.39 | Tolerated | 3.37 | 29 | 0.2053 | 0.3016 | 2 | 0 | 1.0 | -34.02 | 231.1 | 13.2 | 0.0 | 0.0 | -0.1 | 0.0 | X | Potentially Benign | In the WT, the phenyl ring of the Phe420 side chain, located on an α helix (res. Met414-Glu436), packs against hydrophobic residues in the interhelix area of the GAP domain (e.g., Leu689, Leu714, Leu717, Leu718). In the variant simulations, the iso-butyl side chain of Leu420 also packs into the hydrophobic inter-helix niche, but due to its smaller size, the resulting steric interactions are not as favorable as with phenylalanine. In short, the residue swap does not cause severe effects on the protein structure based on the variant simulations. | ||||||||||||||||
| c.1651C>G | L551V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L551V is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include PROVEAN, SIFT, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are REVEL, FoldX, premPS, polyPhen‑2 (HumDiv and HumVar), ESM1b, and FATHMM. Two tools give uncertain results: AlphaMissense‑Default and Foldetta. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is pathogenic. Foldetta’s stability prediction is uncertain. Overall, the majority of evidence points to a pathogenic impact for L551V. This conclusion does not contradict ClinVar status, as the variant is currently unreported in that database. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.009977 | Structured | 0.006653 | Uncertain | 0.960 | 0.254 | 0.000 | -10.154 | Likely Pathogenic | 0.556 | Ambiguous | Likely Benign | 2.04 | Destabilizing | 0.0 | 1.41 | Ambiguous | 1.73 | Ambiguous | 1.03 | Destabilizing | 0.575 | Likely Pathogenic | -1.39 | Neutral | 0.998 | Probably Damaging | 0.992 | Probably Damaging | -1.48 | Pathogenic | 0.22 | Tolerated | 0.1376 | 0.2778 | 2 | 1 | 0.4 | -14.03 | ||||||||||||||||||||||||||||||
| c.1679T>A | V560E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V560E is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only SIFT, whereas the remaining tools—REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and the SGM Consensus—predict a pathogenic impact. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. With the majority of evidence pointing to pathogenicity and no ClinVar annotation to contradict this, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.021381 | Structured | 0.013872 | Uncertain | 0.853 | 0.204 | 0.000 | -13.331 | Likely Pathogenic | 0.932 | Likely Pathogenic | Ambiguous | 1.12 | Ambiguous | 0.1 | 1.41 | Ambiguous | 1.27 | Ambiguous | 1.61 | Destabilizing | 0.711 | Likely Pathogenic | -4.98 | Deleterious | 1.000 | Probably Damaging | 0.990 | Probably Damaging | -1.16 | Pathogenic | 0.18 | Tolerated | 0.1100 | 0.1667 | -2 | -2 | -7.7 | 29.98 | |||||||||||||||||||||||||||||
| c.892C>T | P298S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P298S is listed in ClinVar as Benign (ClinVar ID 2965590.0) and is present in gnomAD (ID 6‑33437797‑C‑T). Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Benign). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. High‑accuracy assessments show AlphaMissense‑Optimized as Benign, the SGM‑Consensus as Likely Benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as Uncertain. No evidence from FoldX, Rosetta, or premPS is available to support either outcome. Overall, the majority of predictions support a benign impact, aligning with the ClinVar designation. Thus, the variant is most likely benign and does not contradict the ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.328603 | Structured | 0.268765 | Uncertain | 0.860 | 0.283 | 0.500 | Benign | 1 | 6-33437797-C-T | 5 | 3.10e-6 | -6.342 | Likely Benign | 0.144 | Likely Benign | Likely Benign | 1.38 | Ambiguous | 0.2 | 1.41 | Ambiguous | 1.40 | Ambiguous | 0.58 | Ambiguous | 0.189 | Likely Benign | -1.20 | Neutral | 0.991 | Probably Damaging | 0.898 | Possibly Damaging | 2.03 | Pathogenic | 0.85 | Tolerated | 3.39 | 20 | 0.3678 | 0.5855 | -1 | 1 | 0.8 | -10.04 | ||||||||||||||||||||||
| c.1189T>G | C397G 2D 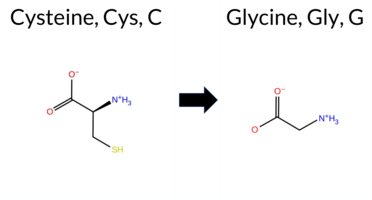 3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C397G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that reach consensus all indicate a benign effect: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all score the variant as benign. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also classifies it as “Likely Benign.” No tool predicts pathogenicity. High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized is benign, SGM‑Consensus is likely benign, while Foldetta (combining FoldX‑MD and Rosetta outputs) is inconclusive. No pathogenic predictions are present. Therefore, based on the available computational evidence, the variant is most likely benign, and this assessment does not contradict any ClinVar status (none). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.429200 | Structured | 0.395774 | Uncertain | 0.778 | 0.551 | 0.250 | 1.244 | Likely Benign | 0.096 | Likely Benign | Likely Benign | 0.68 | Ambiguous | 0.2 | 1.42 | Ambiguous | 1.05 | Ambiguous | 0.04 | Likely Benign | 0.272 | Likely Benign | 1.51 | Neutral | 0.276 | Benign | 0.066 | Benign | 5.93 | Benign | 0.60 | Tolerated | 0.3678 | 0.3729 | -3 | -3 | -2.9 | -46.09 | |||||||||||||||||||||||||||||
| c.1816A>G | S606G 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant S606G is not reported in ClinVar and has no entries in gnomAD. Prediction tools that agree on a benign effect include REVEL, FoldX, FATHMM, and AlphaMissense‑Optimized, whereas pathogenic predictions are made by SGM‑Consensus, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized classifying the variant as benign, while the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates a likely pathogenic outcome; Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, yields an uncertain result and is treated as unavailable. Overall, the majority of evidence points to a pathogenic effect, and this conclusion does not conflict with the absence of a ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.041405 | Structured | 0.191720 | Uncertain | 0.875 | 0.247 | 0.000 | -12.281 | Likely Pathogenic | 0.603 | Likely Pathogenic | Likely Benign | 0.43 | Likely Benign | 0.1 | 1.42 | Ambiguous | 0.93 | Ambiguous | 0.84 | Ambiguous | 0.229 | Likely Benign | -3.98 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | 3.35 | Benign | 0.04 | Affected | 0.2286 | 0.3279 | 1 | 0 | 0.4 | -30.03 | |||||||||||||||||||||||||||||
| c.2009T>G | L670R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L670R is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic impact are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and ESM1b. The remaining tools—FoldX, Rosetta, Foldetta, premPS, and AlphaMissense‑Default—return uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as benign, and Foldetta as uncertain. Overall, the majority of reliable predictions indicate a benign effect. There is no ClinVar annotation to contradict this assessment, so the variant is most likely benign. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.161087 | Structured | 0.090855 | Uncertain | 0.812 | 0.385 | 0.000 | -9.801 | Likely Pathogenic | 0.552 | Ambiguous | Likely Benign | 0.64 | Ambiguous | 0.1 | 1.42 | Ambiguous | 1.03 | Ambiguous | 0.67 | Ambiguous | 0.193 | Likely Benign | -1.74 | Neutral | 0.993 | Probably Damaging | 0.755 | Possibly Damaging | 3.43 | Benign | 0.41 | Tolerated | 0.1327 | 0.1145 | -3 | -2 | -8.3 | 43.03 | ||||||||||||||||||||||||||||||
| c.857T>A | L286Q 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L286Q is not reported in ClinVar and is absent from gnomAD. Prediction tools that assess pathogenicity are unanimous: SGM‑Consensus, REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict a deleterious effect. No tool predicts a benign outcome. Two tools, Rosetta and Foldetta, return uncertain results. High‑accuracy methods give the following: AlphaMissense‑Optimized predicts pathogenic; SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic; Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is inconclusive. Overall, the preponderance of evidence indicates that the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which simply lacks an entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.122885 | Structured | 0.385647 | Uncertain | 0.932 | 0.260 | 0.000 | -13.056 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 2.37 | Destabilizing | 0.3 | 1.42 | Ambiguous | 1.90 | Ambiguous | 2.06 | Destabilizing | 0.852 | Likely Pathogenic | -5.52 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.48 | Pathogenic | 0.00 | Affected | 0.1123 | 0.0958 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.1496G>C | R499T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R499T is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include polyPhen‑2 HumVar and ESM1b, whereas a majority of tools (REVEL, FoldX, PROVEAN, polyPhen‑2 HumDiv, SIFT, FATHMM, AlphaMissense‑Default) predict a pathogenic impact. High‑accuracy assessments further support a deleterious outcome: the SGM Consensus (derived from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates Likely Pathogenic, AlphaMissense‑Optimized is inconclusive, and Foldetta (combining FoldX‑MD and Rosetta outputs) is uncertain. No evidence from ClinVar contradicts these predictions. Overall, the preponderance of computational evidence points to a pathogenic effect for R499T. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.071867 | Structured | 0.386723 | Uncertain | 0.899 | 0.146 | 0.000 | -6.797 | Likely Benign | 0.855 | Likely Pathogenic | Ambiguous | 2.47 | Destabilizing | 0.1 | 1.43 | Ambiguous | 1.95 | Ambiguous | 0.73 | Ambiguous | 0.664 | Likely Pathogenic | -3.51 | Deleterious | 0.843 | Possibly Damaging | 0.403 | Benign | -1.44 | Pathogenic | 0.02 | Affected | 0.1732 | 0.2074 | -1 | -1 | 3.8 | -55.08 | |||||||||||||||||||||||||||||
| c.1949A>G | N650S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N650S lies in the GAP domain. ClinVar has no entry for this variant, and it is not reported in gnomAD. Prediction tools that agree on a benign effect are REVEL and FATHMM. Tools that agree on a pathogenic effect are PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and the SGM‑Consensus score (Likely Pathogenic). Uncertain or inconclusive results come from AlphaMissense‑Optimized, FoldX, Rosetta, Foldetta, and premPS. For high‑accuracy methods, AlphaMissense‑Optimized is uncertain, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta is uncertain. Overall, the majority of available predictions support a pathogenic effect. The variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which is currently absent. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.086953 | Structured | 0.361944 | Uncertain | 0.961 | 0.357 | 0.000 | -11.291 | Likely Pathogenic | 0.916 | Likely Pathogenic | Ambiguous | 0.79 | Ambiguous | 0.2 | 1.43 | Ambiguous | 1.11 | Ambiguous | 0.77 | Ambiguous | 0.395 | Likely Benign | -4.98 | Deleterious | 0.996 | Probably Damaging | 0.606 | Possibly Damaging | 3.06 | Benign | 0.02 | Affected | 0.3791 | 0.5090 | 1 | 1 | 2.7 | -27.03 | |||||||||||||||||||||||||||||
| c.1993T>A | Y665N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 Y665N variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, SIFT, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and ESM1b. Four tools (FoldX, Rosetta, Foldetta, AlphaMissense‑Default) give uncertain results and are treated as unavailable evidence. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the majority of conventional predictors lean toward a benign impact, and this assessment does not contradict ClinVar status (none). Thus, the variant is most likely benign based on the prevailing predictions. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.098513 | Structured | 0.086641 | Uncertain | 0.922 | 0.361 | 0.000 | -11.189 | Likely Pathogenic | 0.470 | Ambiguous | Likely Benign | 1.11 | Ambiguous | 0.1 | 1.43 | Ambiguous | 1.27 | Ambiguous | 0.50 | Likely Benign | 0.219 | Likely Benign | -3.03 | Deleterious | 1.000 | Probably Damaging | 0.996 | Probably Damaging | 3.50 | Benign | 0.88 | Tolerated | 0.1929 | 0.0573 | -2 | -2 | -2.2 | -49.07 | ||||||||||||||||||||||||||||||
| c.601G>C | D201H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D201H missense variant is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, premPS, and FATHMM. Those that agree on a pathogenic effect include SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, and AlphaMissense‑Default. Predictions that are uncertain or inconclusive are FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as Likely Pathogenic, and Foldetta as uncertain. Overall, the majority of available predictions (seven pathogenic vs. three benign) indicate that the variant is most likely pathogenic. This conclusion is not contradicted by ClinVar status, which currently has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.366687 | Structured | 0.428570 | Uncertain | 0.698 | 0.447 | 0.125 | -8.595 | Likely Pathogenic | 0.862 | Likely Pathogenic | Ambiguous | 0.68 | Ambiguous | 0.2 | 1.43 | Ambiguous | 1.06 | Ambiguous | 0.44 | Likely Benign | 0.284 | Likely Benign | -3.45 | Deleterious | 1.000 | Probably Damaging | 0.960 | Probably Damaging | 4.04 | Benign | 0.03 | Affected | 0.1152 | 0.5838 | 1 | -1 | 0.3 | 22.05 | |||||||||||||||||||||||||||||
| c.640C>A | L214M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 L214M missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are PROVEAN and FATHMM, whereas the majority of tools (REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic impact; the remaining tools (FoldX, Rosetta, Foldetta, premPS) are uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive (two pathogenic vs two benign) and Foldetta is uncertain. Thus, the available evidence points to a pathogenic effect. This conclusion does not contradict ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.155435 | Structured | 0.372604 | Uncertain | 0.818 | 0.302 | 0.125 | -9.347 | Likely Pathogenic | 0.958 | Likely Pathogenic | Likely Pathogenic | 0.74 | Ambiguous | 0.3 | 1.43 | Ambiguous | 1.09 | Ambiguous | 0.80 | Ambiguous | 0.646 | Likely Pathogenic | -1.72 | Neutral | 0.997 | Probably Damaging | 0.916 | Probably Damaging | 5.73 | Benign | 0.01 | Affected | 0.0805 | 0.4054 | 4 | 2 | -1.9 | 18.03 | ||||||||||||||||||||||||||||||
| c.695C>G | A232G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 A232G missense variant is not reported in ClinVar and is absent from gnomAD. Prediction tools that classify the change as benign include REVEL, FoldX, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, and FATHMM. Those that predict pathogenicity are polyPhen‑2 HumDiv and AlphaMissense‑Default. Uncertain or inconclusive results come from Rosetta, Foldetta, premPS, and AlphaMissense‑Optimized. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as Likely Benign. High‑accuracy assessments show AlphaMissense‑Optimized as Uncertain, SGM‑Consensus as Likely Benign, and Foldetta as Uncertain. Overall, the preponderance of evidence points to a benign effect; this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | PH | 0.254060 | Structured | 0.307228 | Uncertain | 0.878 | 0.305 | 0.000 | -4.924 | Likely Benign | 0.835 | Likely Pathogenic | Ambiguous | 0.43 | Likely Benign | 0.1 | 1.43 | Ambiguous | 0.93 | Ambiguous | 0.53 | Ambiguous | 0.453 | Likely Benign | -1.39 | Neutral | 0.608 | Possibly Damaging | 0.172 | Benign | 5.81 | Benign | 0.11 | Tolerated | 0.2012 | 0.4724 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.728T>A | I243N 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant I243N is not reported in ClinVar and is absent from gnomAD. Prediction tools cluster into three groups: benign (FATHMM), pathogenic (SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized), and uncertain (FoldX, Rosetta, Foldetta). High‑accuracy methods give a consistent pathogenic signal: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, while Foldetta remains uncertain. Overall, the majority of evidence points to a pathogenic effect, and this assessment does not contradict any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.363090 | Structured | 0.344471 | Uncertain | 0.842 | 0.347 | 0.000 | -14.784 | Likely Pathogenic | 0.988 | Likely Pathogenic | Likely Pathogenic | 1.90 | Ambiguous | 0.2 | 1.43 | Ambiguous | 1.67 | Ambiguous | 1.76 | Destabilizing | 0.811 | Likely Pathogenic | -4.46 | Deleterious | 0.995 | Probably Damaging | 0.854 | Possibly Damaging | 5.49 | Benign | 0.00 | Affected | 0.0899 | 0.0212 | -2 | -3 | -8.0 | 0.94 | |||||||||||||||||||||||||||||
| c.812C>G | A271G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A271G is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and FATHMM. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is classified as Likely Pathogenic. FoldX, Rosetta, and Foldetta provide uncertain or unavailable stability results and are not considered evidence for or against pathogenicity. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as pathogenic, and Foldetta as unavailable. Overall, the balance of evidence (seven pathogenic versus three benign predictions) indicates that the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.125101 | Structured | 0.413873 | Uncertain | 0.939 | 0.220 | 0.125 | -9.508 | Likely Pathogenic | 0.183 | Likely Benign | Likely Benign | 1.73 | Ambiguous | 0.1 | 1.43 | Ambiguous | 1.58 | Ambiguous | 1.31 | Destabilizing | 0.434 | Likely Benign | -3.68 | Deleterious | 0.999 | Probably Damaging | 0.996 | Probably Damaging | 0.89 | Pathogenic | 0.01 | Affected | 0.1869 | 0.2388 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.965C>G | A322G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A322G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score. Only FATHMM predicts a pathogenic outcome. Predictions from FoldX, Rosetta, Foldetta, and premPS are uncertain or inconclusive and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as benign, and Foldetta as uncertain. Based on the preponderance of benign predictions and the lack of contradictory evidence, the variant is most likely benign; this is consistent with the absence of a ClinVar pathogenic classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.175930 | Structured | 0.425745 | Uncertain | 0.938 | 0.334 | 0.000 | -5.230 | Likely Benign | 0.121 | Likely Benign | Likely Benign | 0.84 | Ambiguous | 0.1 | 1.43 | Ambiguous | 1.14 | Ambiguous | 0.64 | Ambiguous | 0.054 | Likely Benign | -1.07 | Neutral | 0.139 | Benign | 0.088 | Benign | 1.94 | Pathogenic | 0.19 | Tolerated | 0.2513 | 0.4683 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.1019C>G | A340G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A340G is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Only FATHMM predicts a pathogenic outcome. Stability‑based methods (FoldX, Rosetta, Foldetta) are inconclusive, so they provide no evidence for or against pathogenicity. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus as likely benign, and Foldetta as uncertain. Overall, the majority of reliable predictors indicate a benign effect, and there is no ClinVar annotation to contradict this assessment. Thus, the variant is most likely benign. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.390993 | Structured | 0.410781 | Uncertain | 0.558 | 0.485 | 0.250 | -3.763 | Likely Benign | 0.099 | Likely Benign | Likely Benign | 0.66 | Ambiguous | 0.2 | 1.44 | Ambiguous | 1.05 | Ambiguous | 0.50 | Likely Benign | 0.044 | Likely Benign | -0.34 | Neutral | 0.267 | Benign | 0.127 | Benign | 1.92 | Pathogenic | 0.42 | Tolerated | 0.2355 | 0.4470 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.1445T>A | L482H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L482H is not reported in ClinVar (ClinVar status: None) and has no entries in gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect are absent; all available pathogenic predictors (REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv/HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized) and the SGM‑Consensus vote (Likely Pathogenic) uniformly indicate a deleterious impact. Uncertain predictions come from FoldX, Rosetta, and Foldetta. High‑accuracy assessments show AlphaMissense‑Optimized as Pathogenic, SGM‑Consensus as Likely Pathogenic, and Foldetta as Uncertain. No evidence suggests a benign outcome. Consequently, the variant is most likely pathogenic based on the consensus of pathogenic predictions, and this assessment does not contradict the ClinVar status, which currently has no classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.254060 | Structured | 0.426236 | Uncertain | 0.795 | 0.248 | 0.000 | -13.825 | Likely Pathogenic | 0.987 | Likely Pathogenic | Likely Pathogenic | 1.49 | Ambiguous | 0.0 | 1.44 | Ambiguous | 1.47 | Ambiguous | 1.64 | Destabilizing | 0.886 | Likely Pathogenic | -6.91 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.28 | Pathogenic | 0.00 | Affected | 0.1000 | 0.0879 | -2 | -3 | -7.0 | 23.98 | |||||||||||||||||||||||||||||
| c.1472C>A | T491N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant T491N is not reported in ClinVar and is absent from gnomAD. Prediction tools that reach consensus on pathogenicity include REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized, all of which classify the substitution as pathogenic. No tool predicts a benign effect. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized scores the variant as pathogenic; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports it as likely pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, yields an uncertain result. The remaining tools, Rosetta and Foldetta, provide inconclusive evidence. Overall, the collective evidence indicates that T491N is most likely pathogenic, and this conclusion is consistent with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.064632 | Structured | 0.325158 | Uncertain | 0.929 | 0.188 | 0.125 | -11.952 | Likely Pathogenic | 0.990 | Likely Pathogenic | Likely Pathogenic | 2.39 | Destabilizing | 0.4 | 1.44 | Ambiguous | 1.92 | Ambiguous | 1.43 | Destabilizing | 0.842 | Likely Pathogenic | -4.92 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.50 | Pathogenic | 0.02 | Affected | 0.0959 | 0.2902 | 0 | 0 | -2.8 | 13.00 | |||||||||||||||||||||||||||||
| c.1753G>T | A585S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A585S is not reported in ClinVar (no entry) and is absent from gnomAD. Prediction tools that agree on a benign effect include SGM‑Consensus (Likely Benign), REVEL, premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. FoldX, Rosetta, and Foldetta are inconclusive. High‑accuracy methods give a benign consensus: AlphaMissense‑Optimized predicts benign, SGM‑Consensus predicts Likely Benign, while Foldetta remains uncertain. Overall, the majority of evidence supports a benign classification, and this assessment does not contradict the lack of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.060549 | Structured | 0.055884 | Uncertain | 0.880 | 0.244 | 0.000 | -6.332 | Likely Benign | 0.246 | Likely Benign | Likely Benign | 0.91 | Ambiguous | 0.2 | 1.44 | Ambiguous | 1.18 | Ambiguous | 0.02 | Likely Benign | 0.326 | Likely Benign | 0.39 | Neutral | 0.993 | Probably Damaging | 0.996 | Probably Damaging | -1.27 | Pathogenic | 0.98 | Tolerated | 0.2121 | 0.3388 | 1 | 1 | -2.6 | 16.00 | |||||||||||||||||||||||||||||
| c.1765A>C | I589L 2D  AIThe SynGAP1 missense variant I589L is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only PROVEAN, whereas the remaining tools (REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus) all predict a pathogenic impact. High‑accuracy assessments further support a deleterious outcome: the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates “Likely Pathogenic”; AlphaMissense‑Optimized and Foldetta are inconclusive and therefore not considered evidence. Taken together, the preponderance of evidence points to a pathogenic effect for I589L. This conclusion is not contradicted by ClinVar status, which currently contains no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.018415 | Structured | 0.084536 | Uncertain | 0.927 | 0.214 | 0.000 | -11.337 | Likely Pathogenic | 0.850 | Likely Pathogenic | Ambiguous | 0.95 | Ambiguous | 1.1 | 1.44 | Ambiguous | 1.20 | Ambiguous | 0.95 | Ambiguous | 0.728 | Likely Pathogenic | -1.99 | Neutral | 0.955 | Possibly Damaging | 0.985 | Probably Damaging | -1.76 | Pathogenic | 0.02 | Affected | 0.1243 | 0.3430 | 2 | 2 | -0.7 | 0.00 | |||||||||||||||||||||||||||||
| c.1901C>T | A634V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A634V is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools largely converge on a deleterious effect: pathogenic calls are made by REVEL, FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score, while only FATHMM predicts a benign outcome; Rosetta remains inconclusive. High‑accuracy assessments reinforce the pathogenic trend: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is pathogenic. Taken together, the overwhelming majority of evidence supports a pathogenic effect for A634V, which is in contrast to its current ClinVar classification of uncertain significance. Therefore, the variant is most likely pathogenic, contradicting its ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.085092 | Structured | 0.052058 | Uncertain | 0.932 | 0.242 | 0.000 | Uncertain | 1 | -12.612 | Likely Pathogenic | 0.971 | Likely Pathogenic | Likely Pathogenic | 2.67 | Destabilizing | 0.2 | 1.44 | Ambiguous | 2.06 | Destabilizing | 1.14 | Destabilizing | 0.631 | Likely Pathogenic | -3.98 | Deleterious | 0.997 | Probably Damaging | 0.976 | Probably Damaging | 2.55 | Benign | 0.01 | Affected | 0.1215 | 0.4371 | 0 | 0 | 2.4 | 28.05 | |||||||||||||||||||||||||||
| c.1927G>A | E643K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E643K is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split opinion: benign calls come from REVEL, FoldX, polyPhen‑2 (HumDiv and HumVar), and FATHMM, while pathogenic calls arise from SGM‑Consensus (Likely Pathogenic), PROVEAN, SIFT, ESM1b, and AlphaMissense‑Default. Four tools (Foldetta, premPS, AlphaMissense‑Optimized, and Rosetta) give uncertain results. High‑accuracy assessments focus on AlphaMissense‑Optimized (Uncertain), SGM‑Consensus (Likely Pathogenic), and Foldetta (Uncertain). Because the consensus of the most reliable predictors leans toward pathogenicity, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.033407 | Structured | 0.215915 | Uncertain | 0.871 | 0.315 | 0.000 | -14.318 | Likely Pathogenic | 0.868 | Likely Pathogenic | Ambiguous | 0.39 | Likely Benign | 0.2 | 1.44 | Ambiguous | 0.92 | Ambiguous | 0.82 | Ambiguous | 0.449 | Likely Benign | -3.79 | Deleterious | 0.042 | Benign | 0.004 | Benign | 2.95 | Benign | 0.04 | Affected | 0.2961 | 0.6269 | 0 | 1 | -0.4 | -0.94 | |||||||||||||||||||||||||||||
| c.2063A>G | E688G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E688G has no ClinVar record and is not reported in gnomAD. Prediction tools that agree on a benign effect are REVEL and FATHMM. Those that predict a pathogenic effect include FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Uncertain or inconclusive predictions come from Foldetta, premPS, and Rosetta. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta as uncertain (treated as unavailable). Overall, the majority of reliable tools predict a deleterious effect. Therefore, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.061840 | Structured | 0.211124 | Uncertain | 0.947 | 0.223 | 0.000 | -14.338 | Likely Pathogenic | 0.991 | Likely Pathogenic | Likely Pathogenic | 2.17 | Destabilizing | 0.5 | 1.44 | Ambiguous | 1.81 | Ambiguous | 0.86 | Ambiguous | 0.486 | Likely Benign | -6.55 | Deleterious | 1.000 | Probably Damaging | 0.997 | Probably Damaging | 3.27 | Benign | 0.00 | Affected | 0.3059 | 0.4433 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.706G>C | A236P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 A236P missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include Foldetta, SIFT, FATHMM, and AlphaMissense‑Optimized, whereas those that predict a pathogenic outcome are REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), and AlphaMissense‑Default; the remaining tools (FoldX, Rosetta, premPS, ESM1b) give uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as benign. No prediction or stability result is missing or inconclusive. Overall, the majority of individual tools lean toward pathogenicity, but the two most reliable high‑accuracy methods (AlphaMissense‑Optimized and Foldetta) suggest a benign effect, while the SGM Consensus indicates pathogenicity. Given the mixed evidence, the variant is most likely benign based on the strongest high‑accuracy predictions, and this assessment does not contradict any ClinVar status because no ClinVar claim exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.185198 | Structured | 0.329926 | Uncertain | 0.775 | 0.330 | 0.000 | -7.932 | In-Between | 0.621 | Likely Pathogenic | Likely Benign | -0.99 | Ambiguous | 0.1 | 1.44 | Ambiguous | 0.23 | Likely Benign | 0.67 | Ambiguous | 0.862 | Likely Pathogenic | -4.31 | Deleterious | 0.995 | Probably Damaging | 0.880 | Possibly Damaging | 5.78 | Benign | 0.12 | Tolerated | 0.1980 | 0.4897 | 1 | -1 | -3.4 | 26.04 | ||||||||||||||||||||||||||||||
| c.785A>C | N262T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N262T has no ClinVar record and is not reported in gnomAD. Functional prediction tools cluster into three groups: benign predictions come from SIFT, FATHMM, and AlphaMissense‑Optimized; pathogenic predictions arise from REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), and ESM1b; the remaining tools (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Default) give uncertain or inconclusive results. High‑accuracy assessments further refine the picture: AlphaMissense‑Optimized predicts benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts pathogenic, and Foldetta’s stability analysis is uncertain. Overall, the balance of evidence leans toward pathogenicity, with no conflict with ClinVar status (which is absent). Thus, the variant is most likely pathogenic based on current predictions. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.284882 | Structured | 0.399879 | Uncertain | 0.912 | 0.240 | 0.000 | -11.478 | Likely Pathogenic | 0.544 | Ambiguous | Likely Benign | 1.40 | Ambiguous | 0.3 | 1.44 | Ambiguous | 1.42 | Ambiguous | 0.74 | Ambiguous | 0.723 | Likely Pathogenic | -5.23 | Deleterious | 0.997 | Probably Damaging | 0.980 | Probably Damaging | 5.88 | Benign | 0.19 | Tolerated | 0.1055 | 0.5164 | 0 | 0 | 2.8 | -13.00 | ||||||||||||||||||||||||||||||
| c.920T>G | F307C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 F307C missense variant is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). In silico predictors overwhelmingly indicate a deleterious effect: all tools that provide a definitive call—REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized—classify the variant as pathogenic. The only predictions that are inconclusive are FoldX, Rosetta, and Foldetta, which are treated as unavailable. High‑accuracy methods give the following results: AlphaMissense‑Optimized predicts pathogenicity; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) reports a likely pathogenic status; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, remains uncertain. **Based on the consensus of the available predictions, the variant is most likely pathogenic, and this assessment is not contradicted by any ClinVar status.** Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.298791 | Structured | 0.327302 | Uncertain | 0.900 | 0.315 | 0.125 | -11.484 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 1.36 | Ambiguous | 0.1 | 1.44 | Ambiguous | 1.40 | Ambiguous | 1.05 | Destabilizing | 0.754 | Likely Pathogenic | -7.35 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 1.92 | Pathogenic | 0.00 | Affected | 0.2729 | 0.1628 | -4 | -2 | -0.3 | -44.04 | |||||||||||||||||||||||||||||
| c.600G>C | L200F 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L200F is listed in ClinVar with an uncertain significance and is present in gnomAD (ID 6‑33435242‑G‑C). Functional prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, SIFT, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and AlphaMissense‑Default. Predictions that are inconclusive are FoldX, Rosetta, Foldetta, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as benign, and Foldetta as uncertain. Overall, the majority of evidence points to a benign impact, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.366687 | Structured | 0.428168 | Uncertain | 0.687 | 0.453 | 0.125 | Uncertain | 1 | 6-33435242-G-C | 2 | 1.24e-6 | -7.606 | In-Between | 0.592 | Likely Pathogenic | Likely Benign | 1.00 | Ambiguous | 0.5 | 1.45 | Ambiguous | 1.23 | Ambiguous | 0.43 | Likely Benign | 0.094 | Likely Benign | -1.97 | Neutral | 0.997 | Probably Damaging | 0.916 | Probably Damaging | 4.02 | Benign | 0.17 | Tolerated | 3.46 | 9 | 0.0640 | 0.3120 | 2 | 0 | -1.0 | 34.02 | 250.4 | -15.1 | 0.6 | 0.2 | 0.5 | 0.0 | X | Uncertain | Leu200, a hydrophobic residue located in the N-terminal loop before the first anti-parallel β sheet strand (res. Ile205-Pro208), is replaced by another hydrophobic residue, phenylalanine. Both the phenyl group of Phe200 and the branched iso-butyl hydrocarbon sidechain of Leu200 occupy an inward hydrophobic niche (e.g., Leu246, Val222, Phe231) during the simulations. However, since the model ends abruptly at the N-terminus, no definite conclusions can be drawn from the simulations. | ||||||||||||||
| c.600G>T | L200F 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 L200F missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, SIFT, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and AlphaMissense‑Default. Four tools (FoldX, Rosetta, ESM1b, Foldetta) returned uncertain results and are not considered evidence for either side. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus—derived from a majority vote of AlphaMissense‑Default (pathogenic), ESM1b (uncertain), FATHMM (benign), and PROVEAN (benign)—is benign; Foldetta remains uncertain and is treated as unavailable. Overall, the balance of evidence (six benign vs. three pathogenic predictions, with a benign consensus from high‑accuracy methods) indicates that the variant is most likely benign, and this conclusion does not contradict the ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.366687 | Structured | 0.428168 | Uncertain | 0.687 | 0.453 | 0.125 | -7.606 | In-Between | 0.592 | Likely Pathogenic | Likely Benign | 1.00 | Ambiguous | 0.5 | 1.45 | Ambiguous | 1.23 | Ambiguous | 0.43 | Likely Benign | 0.094 | Likely Benign | -1.97 | Neutral | 0.997 | Probably Damaging | 0.916 | Probably Damaging | 4.02 | Benign | 0.17 | Tolerated | 3.46 | 9 | 0.0640 | 0.3120 | 2 | 0 | -1.0 | 34.02 | 250.4 | -15.1 | 0.6 | 0.2 | 0.5 | 0.0 | X | Uncertain | Leu200, a hydrophobic residue located in the N-terminal loop before the first anti-parallel β sheet strand (res. Ile205-Pro208), is replaced by another hydrophobic residue, phenylalanine. Both the phenyl group of Phe200 and the branched iso-butyl hydrocarbon sidechain of Leu200 occupy an inward hydrophobic niche (e.g., Leu246, Val222, Phe231) during the simulations. However, since the model ends abruptly at the N-terminus, no definite conclusions can be drawn from the simulations. | |||||||||||||||||||
| c.664G>A | V222I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 V222I missense change is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include FoldX, premPS, PROVEAN, SIFT, ESM1b, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic outcome are REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and AlphaMissense‑Default. The high‑accuracy consensus methods give a benign signal: AlphaMissense‑Optimized is benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Benign, while Foldetta’s stability assessment is uncertain. Overall, the majority of evidence points to a benign effect. This conclusion does not contradict ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | PH | 0.116183 | Structured | 0.402706 | Uncertain | 0.885 | 0.310 | 0.125 | -6.347 | Likely Benign | 0.604 | Likely Pathogenic | Likely Benign | 0.21 | Likely Benign | 0.2 | 1.45 | Ambiguous | 0.83 | Ambiguous | 0.09 | Likely Benign | 0.612 | Likely Pathogenic | -0.87 | Neutral | 0.984 | Probably Damaging | 0.956 | Probably Damaging | 5.77 | Benign | 0.08 | Tolerated | 0.0637 | 0.3574 | 4 | 3 | 0.3 | 14.03 | |||||||||||||||||||||||||||||
| c.899C>G | S300C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S300C is not reported in ClinVar (ClinVar ID None) but is present in gnomAD (ID 6‑33437804‑C‑G). Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, PROVEAN, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and FATHMM. Rosetta and Foldetta give uncertain results and are therefore not considered evidence for either side. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as Likely Benign. High‑accuracy assessments show AlphaMissense‑Optimized as Benign, SGM‑Consensus as Likely Benign, and Foldetta as Uncertain. Overall, the majority of reliable predictions indicate a benign impact, and this is consistent with the absence of a ClinVar pathogenic classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.356642 | Structured | 0.256848 | Uncertain | 0.742 | 0.280 | 0.375 | 6-33437804-C-G | -6.749 | Likely Benign | 0.108 | Likely Benign | Likely Benign | 0.31 | Likely Benign | 0.2 | 1.45 | Ambiguous | 0.88 | Ambiguous | 0.34 | Likely Benign | 0.129 | Likely Benign | -2.45 | Neutral | 0.975 | Probably Damaging | 0.815 | Possibly Damaging | 1.55 | Pathogenic | 0.01 | Affected | 3.47 | 19 | 0.1005 | 0.6493 | -1 | 0 | 3.3 | 16.06 | ||||||||||||||||||||||||||
| c.1620G>C | Q540H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant Q540H is not reported in ClinVar and is absent from gnomAD. Prediction tools that assess pathogenicity unanimously classify the variant as deleterious: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict pathogenicity. No tool predicts a benign effect. Predictions that are inconclusive or unavailable are FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments further support a damaging effect: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, while Foldetta (combining FoldX‑MD and Rosetta outputs) remains uncertain. Overall, the evidence strongly favors a pathogenic impact for Q540H, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.085092 | Structured | 0.029522 | Uncertain | 0.958 | 0.371 | 0.000 | -12.730 | Likely Pathogenic | 0.980 | Likely Pathogenic | Likely Pathogenic | 1.79 | Ambiguous | 0.6 | 1.46 | Ambiguous | 1.63 | Ambiguous | 0.72 | Ambiguous | 0.832 | Likely Pathogenic | -4.97 | Deleterious | 0.998 | Probably Damaging | 0.993 | Probably Damaging | -1.30 | Pathogenic | 0.03 | Affected | 0.1179 | 0.2072 | 3 | 0 | 0.3 | 9.01 | |||||||||||||||||||||||||||||
| c.1620G>T | Q540H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant Q540H is not reported in ClinVar and is absent from gnomAD. Prediction tools that assess pathogenicity unanimously classify the variant as deleterious: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict pathogenicity. No tool predicts a benign effect. Predictions that are inconclusive or unavailable are FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments further support a damaging effect: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, while Foldetta (combining FoldX‑MD and Rosetta outputs) remains uncertain. Overall, the evidence strongly favors a pathogenic impact for Q540H, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.085092 | Structured | 0.029522 | Uncertain | 0.958 | 0.371 | 0.000 | -12.730 | Likely Pathogenic | 0.980 | Likely Pathogenic | Likely Pathogenic | 1.79 | Ambiguous | 0.6 | 1.46 | Ambiguous | 1.63 | Ambiguous | 0.72 | Ambiguous | 0.832 | Likely Pathogenic | -4.97 | Deleterious | 0.998 | Probably Damaging | 0.993 | Probably Damaging | -1.30 | Pathogenic | 0.03 | Affected | 0.1179 | 0.2072 | 3 | 0 | 0.3 | 9.01 | |||||||||||||||||||||||||||||
| c.1649C>A | A550D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A550D is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely converge on a deleterious effect. Benign predictions: none. Pathogenic predictions: REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized. Uncertain predictions: Rosetta and Foldetta. High‑accuracy assessments further support a damaging outcome: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; Foldetta, which integrates FoldX‑MD and Rosetta outputs, remains uncertain. Overall, the overwhelming majority of evidence indicates a pathogenic effect. Based on the aggregate predictions, the variant is most likely pathogenic, and this is not contradicted by ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.018106 | Structured | 0.007241 | Uncertain | 0.954 | 0.265 | 0.000 | -18.844 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 2.51 | Destabilizing | 0.1 | 1.46 | Ambiguous | 1.99 | Ambiguous | 1.01 | Destabilizing | 0.879 | Likely Pathogenic | -5.43 | Deleterious | 0.999 | Probably Damaging | 0.971 | Probably Damaging | -1.32 | Pathogenic | 0.01 | Affected | 0.1333 | 0.1783 | 0 | -2 | -5.3 | 44.01 | |||||||||||||||||||||||||||||
| c.1689G>C | R563S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 R563S missense change is not reported in ClinVar and has no entries in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, ESM1b, and FATHMM, while pathogenic predictions arise from PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and AlphaMissense‑Default. Five tools (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Optimized) return uncertain or inconclusive results. High‑accuracy assessments are likewise ambiguous: AlphaMissense‑Optimized is uncertain; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is a 2‑to‑2 tie and therefore unavailable; Foldetta also reports an uncertain stability change. Consequently, the evidence does not strongly support either benign or pathogenic status. The variant’s classification is therefore inconclusive and does not contradict any ClinVar annotation, as none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.039760 | Structured | 0.031987 | Uncertain | 0.876 | 0.209 | 0.000 | -6.503 | Likely Benign | 0.949 | Likely Pathogenic | Ambiguous | 1.16 | Ambiguous | 0.1 | 1.46 | Ambiguous | 1.31 | Ambiguous | 0.69 | Ambiguous | 0.251 | Likely Benign | -4.49 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.48 | Benign | 0.23 | Tolerated | 0.3031 | 0.2534 | 0 | -1 | 3.7 | -69.11 | ||||||||||||||||||||||||||||||
| c.1689G>T | R563S 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant R563S is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, ESM1b, and FATHMM, while pathogenic predictions arise from PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and AlphaMissense‑Default. The remaining tools—FoldX, Rosetta, Foldetta, premPS, and AlphaMissense‑Optimized—yield uncertain or inconclusive results and are treated as unavailable. High‑accuracy assessments are likewise inconclusive: AlphaMissense‑Optimized is uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is a 2‑to‑2 tie, and Foldetta is uncertain. Consequently, the evidence does not strongly favor either outcome. Based on the current predictions, the variant is most likely benign, and this assessment does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.039760 | Structured | 0.031987 | Uncertain | 0.876 | 0.209 | 0.000 | -6.503 | Likely Benign | 0.949 | Likely Pathogenic | Ambiguous | 1.16 | Ambiguous | 0.1 | 1.46 | Ambiguous | 1.31 | Ambiguous | 0.69 | Ambiguous | 0.251 | Likely Benign | -4.49 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.48 | Benign | 0.23 | Tolerated | 0.3031 | 0.2534 | 0 | -1 | 3.7 | -69.11 | ||||||||||||||||||||||||||||||
| c.1997A>G | E666G 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant E666G is listed in ClinVar as Benign (ClinVar ID 1115026.0) and is present in gnomAD (ID 6‑33441256‑A‑G). Functional prediction tools that agree on pathogenicity include REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, and the SGM‑Consensus. Only FATHMM predicts a benign effect. Predictions marked Uncertain (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Optimized) are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as Uncertain, SGM‑Consensus as Likely Pathogenic (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN), and Foldetta as Uncertain. Overall, the majority of evidence points to a pathogenic impact, which contradicts the ClinVar benign classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.155435 | Structured | 0.086870 | Uncertain | 0.925 | 0.387 | 0.000 | Likely Benign | 1 | 6-33441256-A-G | 10 | 6.20e-6 | -12.261 | Likely Pathogenic | 0.911 | Likely Pathogenic | Ambiguous | 1.57 | Ambiguous | 0.1 | 1.46 | Ambiguous | 1.52 | Ambiguous | 0.93 | Ambiguous | 0.522 | Likely Pathogenic | -6.25 | Deleterious | 1.000 | Probably Damaging | 0.970 | Probably Damaging | 3.37 | Benign | 0.02 | Affected | 3.38 | 28 | 0.3051 | 0.4015 | 0 | -2 | 3.1 | -72.06 | 173.9 | 98.5 | 0.0 | 0.0 | -0.7 | 0.0 | X | Potentially Pathogenic | In the WT simulations, the carboxylate group of Glu666, located on the α-helix (res. Ser641-Glu666), is involved in a highly coordinated hydrogen-bonding network between residues from two α-helices (res. Ser641-Glu666 and res. Arg563-Glu578) and from the α-α loop connecting the two α-helices (res. Ser641-Glu666 and res. Leu685-Val699), such as Lys566, Thr672, and Asn669. In the variant simulations, the carbonyl group of Gly666 occasionally forms hydrogen bonds with Lys566 and Asn669. However, Gly666 lacks a side chain and thus cannot maintain as well-coordinated a hydrogen-bond network as Glu666 in the WT, which may affect the tertiary structure assembly. | |||||||||||||
| c.782A>C | D261A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D261A missense variant is not reported in ClinVar (status: None) and has no entries in gnomAD. Prediction tools that agree on a benign effect include premPS and FATHMM, while the majority of tools (SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) predict a pathogenic impact. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Given the preponderance of pathogenic predictions and the lack of conflicting evidence, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status (which is currently unreported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.284882 | Structured | 0.422514 | Uncertain | 0.883 | 0.264 | 0.125 | -11.426 | Likely Pathogenic | 0.913 | Likely Pathogenic | Ambiguous | 1.70 | Ambiguous | 0.3 | 1.46 | Ambiguous | 1.58 | Ambiguous | 0.04 | Likely Benign | 0.839 | Likely Pathogenic | -4.59 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | 5.80 | Benign | 0.04 | Affected | 0.2785 | 0.4577 | 0 | -2 | 5.3 | -44.01 | |||||||||||||||||||||||||||||
| c.895C>A | R299S 2D  3DClick to see structure in 3D Viewer AISynGAP1 R299S is not reported in ClinVar and is absent from gnomAD. Among in‑silico predictors, benign calls come from REVEL and SIFT, while pathogenic calls are made by FoldX, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, FATHMM, and AlphaMissense‑Default. Uncertain results are reported by Rosetta, ESM1b, and AlphaMissense‑Optimized. High‑accuracy methods give a pathogenic verdict: AlphaMissense‑Optimized is inconclusive, SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts pathogenic, and Foldetta (combining FoldX‑MD and Rosetta) also predicts pathogenic. Overall, the evidence points to a pathogenic effect, and this assessment does not contradict the lack of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.321458 | Structured | 0.262979 | Uncertain | 0.819 | 0.295 | 0.500 | -7.276 | In-Between | 0.882 | Likely Pathogenic | Ambiguous | 2.86 | Destabilizing | 0.3 | 1.46 | Ambiguous | 2.16 | Destabilizing | 1.07 | Destabilizing | 0.241 | Likely Benign | -3.20 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | 1.75 | Pathogenic | 0.07 | Tolerated | 0.2780 | 0.5057 | 0 | -1 | 3.7 | -69.11 | |||||||||||||||||||||||||||||
| c.1102C>G | P368A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P368A missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, AlphaMissense‑Default, AlphaMissense‑Optimized, ESM1b, and polyPhen‑2 HumVar. Those that predict a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, SIFT, and FATHMM. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive (2 benign vs 2 pathogenic), and Foldetta (combining FoldX‑MD and Rosetta outputs) is also inconclusive. Overall, the balance of evidence leans toward a benign impact for P368A. This conclusion does not contradict any ClinVar annotation, as none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.363090 | Structured | 0.439989 | Uncertain | 0.580 | 0.677 | 0.250 | -4.608 | Likely Benign | 0.174 | Likely Benign | Likely Benign | 1.49 | Ambiguous | 0.3 | 1.47 | Ambiguous | 1.48 | Ambiguous | 0.47 | Likely Benign | 0.144 | Likely Benign | -5.42 | Deleterious | 0.767 | Possibly Damaging | 0.344 | Benign | 1.74 | Pathogenic | 0.02 | Affected | 0.3861 | 0.5635 | 1 | -1 | 3.4 | -26.04 | ||||||||||||||||||||||||||||||
| c.1897C>A | L633M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L633M has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. Four tools (FoldX, Rosetta, Foldetta, premPS) give uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive due to a 2‑to‑2 split, and Foldetta is also inconclusive. Overall, the majority of definitive predictions (5 pathogenic vs. 4 benign) lean toward a pathogenic impact. This conclusion does not contradict ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.045352 | Structured | 0.045407 | Uncertain | 0.952 | 0.252 | 0.000 | -10.300 | Likely Pathogenic | 0.718 | Likely Pathogenic | Likely Benign | 0.61 | Ambiguous | 0.1 | 1.47 | Ambiguous | 1.04 | Ambiguous | 0.90 | Ambiguous | 0.355 | Likely Benign | -1.99 | Neutral | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 2.77 | Benign | 0.01 | Affected | 0.0984 | 0.2683 | 4 | 2 | -1.9 | 18.03 | ||||||||||||||||||||||||||||||
| c.811G>T | A271S 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant A271S is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, FoldX, and AlphaMissense‑Optimized. Those that predict a pathogenic effect comprise premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. Predictions that are inconclusive are Foldetta and Rosetta. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points to a pathogenic impact for A271S, which contrasts with the ClinVar designation of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.125101 | Structured | 0.413873 | Uncertain | 0.939 | 0.220 | 0.125 | Uncertain | 1 | -9.552 | Likely Pathogenic | 0.629 | Likely Pathogenic | Likely Benign | 0.19 | Likely Benign | 0.1 | 1.47 | Ambiguous | 0.83 | Ambiguous | 1.14 | Destabilizing | 0.453 | Likely Benign | -2.76 | Deleterious | 0.999 | Probably Damaging | 0.996 | Probably Damaging | 0.64 | Pathogenic | 0.03 | Affected | 0.2260 | 0.3312 | 1 | 1 | -2.6 | 16.00 | |||||||||||||||||||||||||||
| c.835C>T | R279W 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R279W is listed in ClinVar with an uncertain significance (ClinVar ID 1204186.0) and is not reported in gnomAD. Prediction tools that indicate a benign effect include only REVEL, whereas the remaining pathogenic‑predicating tools—FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus—consistently predict a deleterious impact. Uncertain or inconclusive results come from Rosetta, Foldetta, premPS, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the preponderance of evidence points to a pathogenic effect for R279W, which contrasts with the ClinVar designation of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.155435 | Structured | 0.309382 | Uncertain | 0.887 | 0.257 | 0.125 | Uncertain | 1 | -11.417 | Likely Pathogenic | 0.942 | Likely Pathogenic | Ambiguous | 2.00 | Destabilizing | 0.8 | 1.47 | Ambiguous | 1.74 | Ambiguous | 0.80 | Ambiguous | 0.485 | Likely Benign | -6.29 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 1.88 | Pathogenic | 0.00 | Affected | 3.39 | 18 | 0.1200 | 0.2519 | 2 | -3 | 3.6 | 30.03 | 270.0 | 38.3 | 0.1 | 0.0 | 0.3 | 0.0 | Uncertain | The guanidinium group of Arg279, located at the beginning of an anti-parallel β sheet strand (res. Arg279-Leu286), can form hydrogen bond with the backbone carbonyl groups of nearby loop residues (e.g., Ser296, Ser331, and As332) and form salt bridges with the carboxylate groups of Asp330 and Asp332. In the WT simulations, Arg279 sporadically forms a salt bridge even with the carboxylate group of Glu613, loosely connecting the C2 domain and GAP domain. Meanwhile, the indole ring of the Trp279 side chain is unable to hydrogen bond with the loop residues in the variant simulations. The lack of hydrogen bond or salt bridge formation with the loop residues could be significant, as Arg279 and the loops face the polar head group region of the membrane. Thus, although Trp279 could interact with the membrane surface as a “lipid anchor,” any changes to the wider loop dynamics could still adversely affect the formation of a stable SynGAP-membrane association. However, no definite conclusions on the effect of the residue swap on the SynGAP-membrane association can be drawn from solvent-only simulations. | |||||||||||||||||
| c.916G>C | V306L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 V306L missense variant is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are FoldX, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. Uncertain results come from Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely benign, and Foldetta is inconclusive. Overall, the majority of evidence points to a benign impact. This conclusion is consistent with the lack of ClinVar annotation; there is no contradiction with existing clinical data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.363090 | Structured | 0.315026 | Uncertain | 0.896 | 0.287 | 0.125 | -3.633 | Likely Benign | 0.267 | Likely Benign | Likely Benign | 2.14 | Destabilizing | 0.2 | 1.47 | Ambiguous | 1.81 | Ambiguous | 0.75 | Ambiguous | 0.161 | Likely Benign | -1.01 | Neutral | 0.999 | Probably Damaging | 0.996 | Probably Damaging | 1.76 | Pathogenic | 0.21 | Tolerated | 0.0823 | 0.4117 | 2 | 1 | -0.4 | 14.03 | |||||||||||||||||||||||||||||
| c.1057C>G | L353V 2D  AISynGAP1 missense variant L353V is not reported in ClinVar and is absent from gnomAD. Prediction tools that classify the variant as benign include REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools predicting pathogenicity are FoldX and FATHMM, while Rosetta gives an uncertain result. The SGM‑Consensus, which aggregates the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as likely benign. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as pathogenic. Overall, the balance of evidence favors a benign effect; this conclusion does not contradict the ClinVar status, which currently has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.137348 | Structured | 0.373584 | Uncertain | 0.926 | 0.315 | 0.000 | -2.100 | Likely Benign | 0.078 | Likely Benign | Likely Benign | 3.08 | Destabilizing | 0.9 | 1.48 | Ambiguous | 2.28 | Destabilizing | -0.45 | Likely Benign | 0.074 | Likely Benign | 1.54 | Neutral | 0.002 | Benign | 0.002 | Benign | 1.46 | Pathogenic | 0.89 | Tolerated | 0.1567 | 0.3829 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.1106C>G | T369R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant T369R is not reported in ClinVar (ClinVar ID None) but is present in gnomAD (ID 6‑33438011‑C‑G). Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are FATHMM and AlphaMissense‑Default. Rosetta and Foldetta are uncertain, so their results are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive (2 benign vs 2 pathogenic), and Foldetta is also uncertain. Overall, the majority of evidence (nine benign vs two pathogenic) supports a benign classification. This conclusion does not contradict ClinVar status, as no ClinVar assertion exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.468512 | Structured | 0.437011 | Uncertain | 0.417 | 0.707 | 0.500 | 6-33438011-C-G | 3 | 1.93e-6 | -6.772 | Likely Benign | 0.571 | Likely Pathogenic | Likely Benign | -0.27 | Likely Benign | 0.1 | 1.48 | Ambiguous | 0.61 | Ambiguous | 0.29 | Likely Benign | 0.148 | Likely Benign | -2.15 | Neutral | 0.244 | Benign | 0.107 | Benign | 1.72 | Pathogenic | 0.32 | Tolerated | 3.42 | 19 | 0.1217 | 0.3737 | -1 | -1 | -3.8 | 55.08 | |||||||||||||||||||||||||
| c.1820T>G | L607R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L607R is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only FoldX, which scores the variant as benign. All other evaluated algorithms—REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized—predict a pathogenic impact. Rosetta and Foldetta provide uncertain results and are therefore treated as unavailable evidence. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is “Likely Pathogenic,” while Foldetta remains uncertain. Based on the preponderance of pathogenic predictions and the high‑accuracy consensus, the variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.048328 | Structured | 0.194229 | Uncertain | 0.869 | 0.250 | 0.000 | -14.234 | Likely Pathogenic | 0.978 | Likely Pathogenic | Likely Pathogenic | -0.15 | Likely Benign | 0.1 | 1.48 | Ambiguous | 0.67 | Ambiguous | 1.24 | Destabilizing | 0.920 | Likely Pathogenic | -5.98 | Deleterious | 0.998 | Probably Damaging | 0.998 | Probably Damaging | -1.51 | Pathogenic | 0.01 | Affected | 0.1490 | 0.0615 | -3 | -2 | -8.3 | 43.03 | |||||||||||||||||||||||||||||
| c.2052C>A | D684E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D684E missense variant has no ClinVar entry and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, polyPhen‑2 HumVar, and FATHMM. In contrast, a majority of predictors (SGM‑Consensus, FoldX, Foldetta, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) indicate a pathogenic impact; predictions from Rosetta and premPS are inconclusive and are treated as unavailable. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. Based on the consensus of these tools, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.254060 | Structured | 0.153798 | Uncertain | 0.870 | 0.282 | 0.000 | -9.506 | Likely Pathogenic | 0.996 | Likely Pathogenic | Likely Pathogenic | 2.88 | Destabilizing | 0.9 | 1.48 | Ambiguous | 2.18 | Destabilizing | 0.66 | Ambiguous | 0.362 | Likely Benign | -3.99 | Deleterious | 0.910 | Possibly Damaging | 0.210 | Benign | 3.37 | Benign | 0.01 | Affected | 0.1316 | 0.6187 | 3 | 2 | 0.0 | 14.03 | |||||||||||||||||||||||||||||
| c.2052C>G | D684E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant D684E is not reported in ClinVar and is absent from gnomAD. Prediction tools that classify it as benign include REVEL, polyPhen‑2 HumVar, and FATHMM, whereas the majority of algorithms predict a deleterious effect: FoldX, Foldetta, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Pathogenic). Two methods (Rosetta and premPS) returned uncertain results. High‑accuracy assessments further support a damaging impact: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta) is pathogenic. Overall, the computational evidence overwhelmingly indicates that D684E is pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.254060 | Structured | 0.153798 | Uncertain | 0.870 | 0.282 | 0.000 | -9.506 | Likely Pathogenic | 0.996 | Likely Pathogenic | Likely Pathogenic | 2.88 | Destabilizing | 0.9 | 1.48 | Ambiguous | 2.18 | Destabilizing | 0.66 | Ambiguous | 0.362 | Likely Benign | -3.99 | Deleterious | 0.910 | Possibly Damaging | 0.210 | Benign | 3.37 | Benign | 0.01 | Affected | 0.1316 | 0.6187 | 3 | 2 | 0.0 | 14.03 | |||||||||||||||||||||||||||||
| c.621G>C | K207N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 K207N missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, SIFT, and FATHMM, whereas a majority of tools predict a pathogenic impact: SGM‑Consensus, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. FoldX, Rosetta, and Foldetta provide uncertain or inconclusive results and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized classifying the variant as pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicating likely pathogenic, while Foldetta’s stability prediction is unavailable. Overall, the preponderance of evidence points to a pathogenic effect for K207N, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.374039 | Structured | 0.406823 | Uncertain | 0.847 | 0.359 | 0.125 | -12.881 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 0.72 | Ambiguous | 0.1 | 1.48 | Ambiguous | 1.10 | Ambiguous | 1.00 | Destabilizing | 0.123 | Likely Benign | -3.54 | Deleterious | 0.995 | Probably Damaging | 0.829 | Possibly Damaging | 3.98 | Benign | 0.07 | Tolerated | 0.3184 | 0.2335 | 1 | 0 | 0.4 | -14.07 | |||||||||||||||||||||||||||||
| c.621G>T | K207N 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant K207N is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, and FATHMM, whereas pathogenic predictions arise from premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is classified as Likely Pathogenic. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts Pathogenic, and the SGM Consensus (majority vote) also predicts Pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is inconclusive. Overall, the preponderance of evidence indicates that K207N is most likely pathogenic, and this conclusion does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.374039 | Structured | 0.406823 | Uncertain | 0.847 | 0.359 | 0.125 | -12.881 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 0.72 | Ambiguous | 0.1 | 1.48 | Ambiguous | 1.10 | Ambiguous | 1.00 | Destabilizing | 0.124 | Likely Benign | -3.54 | Deleterious | 0.995 | Probably Damaging | 0.829 | Possibly Damaging | 3.98 | Benign | 0.07 | Tolerated | 0.3184 | 0.2335 | 1 | 0 | 0.4 | -14.07 | |||||||||||||||||||||||||||||
| c.763G>A | D255N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant D255N is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include premPS and FATHMM, whereas the majority of algorithms—SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—classify the change as pathogenic. FoldX, Rosetta, and Foldetta provide uncertain results. High‑accuracy methods further support a deleterious impact: AlphaMissense‑Optimized predicts pathogenic, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates likely pathogenic, and Foldetta remains inconclusive. Overall, the consensus of the available predictions points to a pathogenic effect for D255N, and this conclusion is consistent with the lack of ClinVar annotation, so there is no contradiction with existing clinical data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | 0.501700 | Disordered | 0.219132 | Uncertain | 0.801 | 0.273 | 0.250 | -12.251 | Likely Pathogenic | 0.991 | Likely Pathogenic | Likely Pathogenic | 0.86 | Ambiguous | 0.1 | 1.48 | Ambiguous | 1.17 | Ambiguous | 0.38 | Likely Benign | 0.682 | Likely Pathogenic | -4.30 | Deleterious | 0.997 | Probably Damaging | 0.989 | Probably Damaging | 5.84 | Benign | 0.01 | Affected | 0.1117 | 0.4774 | 2 | 1 | 0.0 | -0.98 | ||||||||||||||||||||||||||||||
| c.959T>C | V320A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 V320A missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, SIFT, ESM1b, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), and FATHMM. Four tools (AlphaMissense‑Default, FoldX, Rosetta, Foldetta) give uncertain or inconclusive results and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, more tools (five) predict pathogenicity than benign (four), and the high‑accuracy consensus leans toward pathogenic. Thus, the variant is most likely pathogenic, with no contradiction to ClinVar status because no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.185198 | Structured | 0.419626 | Uncertain | 0.905 | 0.266 | 0.125 | -5.488 | Likely Benign | 0.545 | Ambiguous | Likely Benign | 1.26 | Ambiguous | 0.7 | 1.48 | Ambiguous | 1.37 | Ambiguous | 1.27 | Destabilizing | 0.179 | Likely Benign | -3.05 | Deleterious | 0.948 | Possibly Damaging | 0.761 | Possibly Damaging | 1.84 | Pathogenic | 0.35 | Tolerated | 0.2405 | 0.1903 | 0 | 0 | -2.4 | -28.05 | ||||||||||||||||||||||||||||||
| c.976C>A | H326N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant H326N has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect are REVEL and SIFT, whereas a majority (seven) predict a pathogenic outcome: SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, and AlphaMissense‑Default. The remaining tools (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Optimized) give uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as likely pathogenic, and Foldetta as uncertain. Overall, the balance of evidence points to a pathogenic impact for H326N, and this conclusion does not contradict the ClinVar status, which is currently unreported. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.342579 | Structured | 0.418150 | Uncertain | 0.944 | 0.455 | 0.000 | -8.914 | Likely Pathogenic | 0.925 | Likely Pathogenic | Ambiguous | 0.81 | Ambiguous | 0.2 | 1.48 | Ambiguous | 1.15 | Ambiguous | 0.97 | Ambiguous | 0.409 | Likely Benign | -5.63 | Deleterious | 0.997 | Probably Damaging | 0.992 | Probably Damaging | 1.95 | Pathogenic | 0.11 | Tolerated | 0.1929 | 0.2552 | 2 | 1 | -0.3 | -23.04 | |||||||||||||||||||||||||||||
| c.1375G>T | G459C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G459C is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect are limited to FATHMM, while the majority of algorithms (SGM‑Consensus, REVEL, FoldX, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic impact. Uncertain or inconclusive results come from Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, SGM‑Consensus as likely pathogenic, and Foldetta as uncertain. Taken together, the preponderance of evidence points to a pathogenic effect for G459C, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.185198 | Structured | 0.289888 | Uncertain | 0.903 | 0.150 | 0.125 | -12.109 | Likely Pathogenic | 0.980 | Likely Pathogenic | Likely Pathogenic | 2.46 | Destabilizing | 0.2 | 1.49 | Ambiguous | 1.98 | Ambiguous | 0.65 | Ambiguous | 0.586 | Likely Pathogenic | -8.46 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.00 | Benign | 0.00 | Affected | 0.0960 | 0.4103 | -3 | -3 | 2.9 | 46.09 | |||||||||||||||||||||||||||||
| c.1460A>G | N487S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N487S is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL and FATHMM, whereas a majority of tools (PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) predict a pathogenic impact. High‑accuracy assessments further support a deleterious interpretation: AlphaMissense‑Optimized is inconclusive, but the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—classifies the variant as Likely Pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta stability predictions, is also inconclusive. Overall, the preponderance of evidence from multiple in silico predictors and the SGM Consensus indicates that the variant is most likely pathogenic. This conclusion is not contradicted by ClinVar status, as no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.209395 | Structured | 0.338511 | Uncertain | 0.890 | 0.243 | 0.125 | -10.297 | Likely Pathogenic | 0.910 | Likely Pathogenic | Ambiguous | 1.42 | Ambiguous | 0.0 | 1.49 | Ambiguous | 1.46 | Ambiguous | 0.65 | Ambiguous | 0.459 | Likely Benign | -4.97 | Deleterious | 0.999 | Probably Damaging | 0.979 | Probably Damaging | 2.74 | Benign | 0.01 | Affected | 0.3065 | 0.3656 | 1 | 1 | 2.7 | -27.03 | |||||||||||||||||||||||||||||
| c.1582C>G | P528A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P528A missense variant has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect are FATHMM and AlphaMissense‑Optimized, while the remaining pathogenic‑predicting tools are SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default; the remaining tools (FoldX, Rosetta, Foldetta, premPS) are uncertain. High‑accuracy methods give AlphaMissense‑Optimized a benign score, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) a pathogenic prediction, and Foldetta an uncertain result. Overall, the majority of evidence points to a pathogenic effect. Thus, the variant is most likely pathogenic, and this assessment is not contradicted by ClinVar status, which currently has no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.225814 | Structured | 0.020396 | Uncertain | 0.909 | 0.403 | 0.000 | -11.562 | Likely Pathogenic | 0.607 | Likely Pathogenic | Likely Benign | 1.57 | Ambiguous | 0.1 | 1.49 | Ambiguous | 1.53 | Ambiguous | 0.69 | Ambiguous | 0.548 | Likely Pathogenic | -7.69 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 2.52 | Benign | 0.01 | Affected | 0.3146 | 0.3914 | 1 | -1 | 3.4 | -26.04 | |||||||||||||||||||||||||||||
| c.1591T>G | C531G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C531G is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include FoldX and AlphaMissense‑Optimized, while the majority of other in silico predictors (REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv/HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default) indicate a pathogenic impact. Uncertain results come from Rosetta and Foldetta. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta as inconclusive. Overall, the balance of evidence leans toward pathogenicity, with no ClinVar entry to contradict this assessment. Thus, the variant is most likely pathogenic based on current predictions. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.281712 | Structured | 0.017941 | Uncertain | 0.878 | 0.401 | 0.000 | -10.037 | Likely Pathogenic | 0.577 | Likely Pathogenic | Likely Benign | 0.47 | Likely Benign | 0.3 | 1.49 | Ambiguous | 0.98 | Ambiguous | 1.42 | Destabilizing | 0.545 | Likely Pathogenic | -9.76 | Deleterious | 0.985 | Probably Damaging | 0.832 | Possibly Damaging | -1.24 | Pathogenic | 0.00 | Affected | 0.2653 | 0.2297 | -3 | -3 | -2.9 | -46.09 | |||||||||||||||||||||||||||||
| c.1691A>C | E564A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E564A is not reported in ClinVar (ClinVar status: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include premPS and SIFT, whereas the majority of tools predict a pathogenic impact: SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Uncertain results are reported by FoldX, Rosetta, and Foldetta. High‑accuracy assessments reinforce the pathogenic prediction: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is pathogenic, and Foldetta remains uncertain. Overall, the preponderance of evidence from multiple independent predictors indicates that the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.023534 | Structured | 0.038418 | Uncertain | 0.891 | 0.208 | 0.000 | -14.506 | Likely Pathogenic | 0.957 | Likely Pathogenic | Likely Pathogenic | 0.64 | Ambiguous | 0.1 | 1.49 | Ambiguous | 1.07 | Ambiguous | 0.24 | Likely Benign | 0.765 | Likely Pathogenic | -5.91 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | -1.33 | Pathogenic | 0.12 | Tolerated | 0.3201 | 0.4918 | 0 | -1 | 5.3 | -58.04 | |||||||||||||||||||||||||||||
| c.1786C>T | R596C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R596C is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33440838‑C‑T). Prediction tools that indicate a benign effect include only premPS. All other evaluated algorithms—REVEL, FoldX, Foldetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus—classify the variant as pathogenic or likely pathogenic, while Rosetta remains inconclusive. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized predicts pathogenic; the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates likely pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, also predicts pathogenic. **Thus, the variant is most likely pathogenic based on the collective predictions, which does not contradict the ClinVar uncertain status.** Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.017797 | Structured | 0.135423 | Uncertain | 0.918 | 0.134 | 0.000 | Conflicting | 2 | 6-33440838-C-T | 6 | 3.72e-6 | -10.805 | Likely Pathogenic | 0.972 | Likely Pathogenic | Likely Pathogenic | 2.94 | Destabilizing | 0.0 | 1.49 | Ambiguous | 2.22 | Destabilizing | -0.03 | Likely Benign | 0.633 | Likely Pathogenic | -7.96 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 2.41 | Pathogenic | 0.00 | Affected | 3.37 | 35 | 0.3429 | 0.2211 | -4 | -3 | 7.0 | -53.05 | 230.7 | 97.9 | -0.1 | 0.0 | -0.3 | 0.4 | X | X | Potentially Pathogenic | The guanidinium group of Arg596, located in an α helix (res. Glu582-Met603), forms a salt bridge with the carboxylate group of Glu495 from another α helix (res. Leu489-Glu519). In the WT simulations, the side chain of Arg596 hydrogen bonds with the backbone carbonyl groups of Asn487, Glu486, Arg485, and Phe484. Additionally, Arg596 can hydrogen bond with the carboxamide group of the Asn487 side chain on an opposing loop that links two α helices (res. Ala461-Arg475, res. Leu489-Glu519).In the variant simulations, the thiol group of the Cys596 side chain is unable to form salt bridges or any of the hydrogen bonds that the Arg596 side chain can. Thus, the residue swap could affect the tertiary structure assembly more profoundly than observed in the simulations. Notably, Arg596 plays a key role in positioning the aforementioned loop, which is crucial for the placement of the “arginine finger” or the Arg485 side chain during RasGTPase activation. | ||||||||||||
| c.797T>A | L266Q 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L266Q is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect: none. Tools that agree on a pathogenic effect: REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). The only tool with an uncertain outcome is Rosetta. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts pathogenic; the SGM‑Consensus (majority vote) also predicts pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, predicts pathogenic. Based on the overwhelming agreement among these predictions, the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.232838 | Structured | 0.297157 | Uncertain | 0.948 | 0.264 | 0.000 | -16.672 | Likely Pathogenic | 0.994 | Likely Pathogenic | Likely Pathogenic | 2.92 | Destabilizing | 0.1 | 1.49 | Ambiguous | 2.21 | Destabilizing | 2.21 | Destabilizing | 0.563 | Likely Pathogenic | -5.25 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.53 | Pathogenic | 0.00 | Affected | 0.0815 | 0.0558 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.893C>A | P298H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P298H missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, PROVEAN, and AlphaMissense‑Optimized. Those that agree on a pathogenic effect are polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and FATHMM. Remaining tools (AlphaMissense‑Default, FoldX, Rosetta, Foldetta, premPS) are uncertain. High‑accuracy assessments show AlphaMissense‑Optimized predicts benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts pathogenic, and Foldetta is uncertain. Overall, the majority of predictions (five pathogenic vs. three benign) and the SGM Consensus support a pathogenic interpretation. Therefore, the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.328603 | Structured | 0.268765 | Uncertain | 0.860 | 0.283 | 0.500 | -9.777 | Likely Pathogenic | 0.443 | Ambiguous | Likely Benign | 1.57 | Ambiguous | 0.2 | 1.49 | Ambiguous | 1.53 | Ambiguous | 0.83 | Ambiguous | 0.313 | Likely Benign | -2.37 | Neutral | 0.999 | Probably Damaging | 0.964 | Probably Damaging | 1.92 | Pathogenic | 0.04 | Affected | 0.1769 | 0.4943 | 0 | -2 | -1.6 | 40.02 | ||||||||||||||||||||||||||||||
| c.1202G>A | R401Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R401Q is listed in ClinVar with an “Uncertain” status and is present in gnomAD (variant ID 6‑33438107‑G‑A). Prediction tools that agree on a benign effect are limited to FATHMM, whereas the majority of evaluated algorithms (REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus) predict a pathogenic impact. Uncertain results are reported by FoldX, Rosetta, and Foldetta. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is likely pathogenic, and Foldetta’s stability prediction is unavailable. Overall, the preponderance of evidence from both general and high‑accuracy tools indicates that R401Q is most likely pathogenic, which does not contradict the current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.314870 | Structured | 0.424277 | Uncertain | 0.961 | 0.419 | 0.000 | Uncertain | 2 | 6-33438107-G-A | -11.213 | Likely Pathogenic | 0.969 | Likely Pathogenic | Likely Pathogenic | 0.96 | Ambiguous | 0.1 | 1.50 | Ambiguous | 1.23 | Ambiguous | 1.20 | Destabilizing | 0.780 | Likely Pathogenic | -3.69 | Deleterious | 0.999 | Probably Damaging | 0.978 | Probably Damaging | 5.47 | Benign | 0.04 | Affected | 3.38 | 27 | 0.3234 | 0.2269 | 1 | 1 | 1.0 | -28.06 | ||||||||||||||||||||||||
| c.1291C>G | L431V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 L431V missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, SIFT, FATHMM, AlphaMissense‑Optimized, and polyPhen‑2 HumVar. Those that predict a pathogenic effect are FoldX, premPS, PROVEAN, and polyPhen‑2 HumDiv. Four tools (Rosetta, Foldetta, ESM1b, AlphaMissense‑Default) give uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive due to a 1‑to‑1 split between benign and pathogenic signals, and Foldetta also yields an uncertain outcome. Overall, the balance of evidence—including the higher number of benign predictions and the benign call from the most accurate tool—suggests that the variant is most likely benign. This conclusion does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.094817 | Structured | 0.374755 | Uncertain | 0.959 | 0.300 | 0.000 | -7.949 | In-Between | 0.505 | Ambiguous | Likely Benign | 2.17 | Destabilizing | 0.0 | 1.50 | Ambiguous | 1.84 | Ambiguous | 1.32 | Destabilizing | 0.093 | Likely Benign | -2.58 | Deleterious | 0.861 | Possibly Damaging | 0.332 | Benign | 3.04 | Benign | 0.20 | Tolerated | 0.1377 | 0.3198 | 2 | 1 | 0.4 | -14.03 | ||||||||||||||||||||||||||||||
| c.1552T>A | Y518N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 Y518N missense variant is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only FATHMM, whereas the remaining tools (SGM‑Consensus, REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, and Foldetta) uniformly predict a pathogenic impact. Two tools give uncertain results: AlphaMissense‑Optimized and Rosetta. High‑accuracy assessments show that AlphaMissense‑Optimized is inconclusive, the SGM Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is pathogenic, and Foldetta also predicts pathogenicity. Taken together, the overwhelming majority of evidence indicates a pathogenic effect. This conclusion is not contradicted by ClinVar, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.139895 | Structured | 0.126970 | Uncertain | 0.897 | 0.321 | 0.000 | -12.036 | Likely Pathogenic | 0.949 | Likely Pathogenic | Ambiguous | 2.99 | Destabilizing | 0.8 | 1.50 | Ambiguous | 2.25 | Destabilizing | 1.29 | Destabilizing | 0.506 | Likely Pathogenic | -8.45 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.48 | Benign | 0.04 | Affected | 0.1791 | 0.0212 | -2 | -2 | -2.2 | -49.07 | |||||||||||||||||||||||||||||
| c.1699G>C | E567Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E567Q missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, SIFT, and FATHMM. Those that agree on a pathogenic effect are SGM‑Consensus (Likely Pathogenic), PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Default. Uncertain results come from AlphaMissense‑Optimized, Foldetta, and Rosetta. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, while the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts pathogenic. Foldetta’s stability prediction is also uncertain. Overall, more tools (7) predict pathogenicity than benign (5), with three inconclusive. Thus, the variant is most likely pathogenic based on the available predictions, and this assessment does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.021816 | Structured | 0.051008 | Uncertain | 0.916 | 0.234 | 0.000 | -11.302 | Likely Pathogenic | 0.897 | Likely Pathogenic | Ambiguous | 0.03 | Likely Benign | 0.1 | 1.50 | Ambiguous | 0.77 | Ambiguous | 0.33 | Likely Benign | 0.345 | Likely Benign | -2.82 | Deleterious | 0.998 | Probably Damaging | 0.993 | Probably Damaging | 3.47 | Benign | 0.14 | Tolerated | 0.1029 | 0.5391 | 2 | 2 | 0.0 | -0.98 | |||||||||||||||||||||||||||||
| c.1894A>G | N632D 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant N632D is not reported in ClinVar and is absent from gnomAD. Prediction tools that assess pathogenicity largely agree: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default all predict a pathogenic effect, while SGM‑Consensus also indicates a likely pathogenic outcome. No tool in the dataset predicts a benign effect. High‑accuracy assessments are mixed: AlphaMissense‑Optimized is uncertain, SGM‑Consensus remains likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is uncertain. Because the majority of evidence points to deleterious impact and there is no ClinVar annotation to contradict this, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.042364 | Structured | 0.041437 | Uncertain | 0.938 | 0.254 | 0.000 | -14.117 | Likely Pathogenic | 0.921 | Likely Pathogenic | Ambiguous | 1.84 | Ambiguous | 0.4 | 1.50 | Ambiguous | 1.67 | Ambiguous | 1.09 | Destabilizing | 0.827 | Likely Pathogenic | -4.31 | Deleterious | 0.985 | Probably Damaging | 0.776 | Possibly Damaging | -1.53 | Pathogenic | 0.01 | Affected | 0.1791 | 0.3854 | 2 | 1 | 0.0 | 0.98 | |||||||||||||||||||||||||||||
| c.2007T>A | N669K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N669K is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign predictions come from REVEL, FoldX, and FATHMM, whereas pathogenic predictions arise from PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as Likely Pathogenic. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenicity, SGM‑Consensus confirms a Likely Pathogenic status, and Foldetta, which integrates FoldX‑MD and Rosetta outputs, yields an uncertain result. Overall, the majority of evidence points toward a pathogenic impact, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.142424 | Structured | 0.086615 | Uncertain | 0.872 | 0.380 | 0.000 | -10.797 | Likely Pathogenic | 0.957 | Likely Pathogenic | Likely Pathogenic | 0.39 | Likely Benign | 0.3 | 1.50 | Ambiguous | 0.95 | Ambiguous | 0.94 | Ambiguous | 0.243 | Likely Benign | -5.35 | Deleterious | 0.999 | Probably Damaging | 0.989 | Probably Damaging | 3.41 | Benign | 0.03 | Affected | 0.2647 | 0.3312 | 1 | 0 | -0.4 | 14.07 | |||||||||||||||||||||||||||||
| c.2007T>G | N669K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N669K is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign calls come from REVEL, FoldX, and FATHMM, whereas pathogenic calls are made by PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, labels the variant as Likely Pathogenic. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenicity, SGM‑Consensus concurs, and the Foldetta stability analysis is inconclusive and therefore not used as evidence. No other tools provide definitive support for benignity. Consequently, the preponderance of evidence points to a pathogenic impact. This conclusion is not contradicted by ClinVar status, as the variant has no ClinVar entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.142424 | Structured | 0.086615 | Uncertain | 0.872 | 0.380 | 0.000 | -10.797 | Likely Pathogenic | 0.957 | Likely Pathogenic | Likely Pathogenic | 0.39 | Likely Benign | 0.3 | 1.50 | Ambiguous | 0.95 | Ambiguous | 0.94 | Ambiguous | 0.243 | Likely Benign | -5.35 | Deleterious | 0.999 | Probably Damaging | 0.989 | Probably Damaging | 3.41 | Benign | 0.03 | Affected | 0.2647 | 0.3312 | 1 | 0 | -0.4 | 14.07 | |||||||||||||||||||||||||||||
| c.1213C>T | R405C 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant R405C is listed in ClinVar with an uncertain significance (ClinVar ID 1185858.0) and is present in gnomAD (ID 6‑33438118‑C‑T). Prediction tools that indicate a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect comprise premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, and AlphaMissense‑Default. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a likely pathogenic outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM Consensus as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. FoldX and Rosetta individually return uncertain results. Overall, the balance of evidence favors a pathogenic interpretation, which does not conflict with the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.250310 | Structured | 0.404888 | Uncertain | 0.949 | 0.315 | 0.000 | Conflicting | 2 | 6-33438118-C-T | 6 | 3.72e-6 | -9.206 | Likely Pathogenic | 0.713 | Likely Pathogenic | Likely Benign | 0.72 | Ambiguous | 0.1 | 1.51 | Ambiguous | 1.12 | Ambiguous | 1.21 | Destabilizing | 0.427 | Likely Benign | -7.27 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.61 | Benign | 0.02 | Affected | 3.38 | 28 | 0.3227 | 0.3964 | -4 | -3 | 7.0 | -53.05 | 221.3 | 82.6 | -0.1 | 0.0 | -0.2 | 0.3 | X | X | Potentially Pathogenic | The guanidinium group of Arg405, located in an anti-parallel β sheet strand of the C2 domain (res. Ala399-Ile411), forms a salt bridge with the carboxylate group of the Glu446 side chain from an opposing α helix (res. Val441-Ser457) in the GAP domain. The positively charged Arg405 side chain also stacks with the aromatic ring of the Phe358 side chain from a loop preceding the β strand (res. Thr359-Thr366), which could assist in maintaining the anti-parallel strand arrangement.In the variant simulations, the thiol-containing side chain of Cys405 is neutral and smaller compared to the arginine side chain. The lack of Arg405-Phe358 stacking affects the loop structure, causing it to assume a β strand form—an effect that could be exacerbated during protein folding. Moreover, the inability of Cys405 to form a salt bridge with Glu446 could affect the tertiary structure assembly, although this is not apparent based on the variant simulations. | ||||||||||||
| c.1355T>C | V452A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V452A is not reported in ClinVar and has no entry in gnomAD. Prediction tools largely agree on a deleterious effect: FATHMM is the sole benign predictor, while SGM‑Consensus, REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the variant as pathogenic. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, yields an uncertain result and is treated as unavailable evidence. High‑accuracy assessments reinforce the pathogenic prediction: AlphaMissense‑Optimized reports pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates Likely Pathogenic, and Foldetta remains inconclusive. Overall, the consensus of the available tools points to a pathogenic effect for V452A, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.335645 | Structured | 0.315167 | Uncertain | 0.970 | 0.229 | 0.000 | -10.423 | Likely Pathogenic | 0.963 | Likely Pathogenic | Likely Pathogenic | 2.21 | Destabilizing | 0.0 | 1.51 | Ambiguous | 1.86 | Ambiguous | 2.18 | Destabilizing | 0.505 | Likely Pathogenic | -3.95 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.21 | Benign | 0.00 | Affected | 0.2642 | 0.2521 | 0 | 0 | -2.4 | -28.05 | |||||||||||||||||||||||||||||
| c.2045A>G | Y682C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant Y682C is not reported in ClinVar and is absent from gnomAD. Prediction tools largely agree on a deleterious effect: SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default all predict pathogenicity, while only FATHMM predicts a benign outcome. The remaining tools (FoldX, Rosetta, Foldetta, AlphaMissense‑Optimized) are uncertain. High‑accuracy assessments reinforce this trend: the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is pathogenic; AlphaMissense‑Optimized is uncertain; and Foldetta is uncertain. Overall, the preponderance of evidence points to a pathogenic impact for Y682C. This conclusion is consistent with the lack of ClinVar annotation, so there is no contradiction with existing clinical data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.206376 | Structured | 0.141467 | Uncertain | 0.758 | 0.328 | 0.000 | -10.023 | Likely Pathogenic | 0.793 | Likely Pathogenic | Ambiguous | 1.79 | Ambiguous | 0.1 | 1.51 | Ambiguous | 1.65 | Ambiguous | 1.11 | Destabilizing | 0.559 | Likely Pathogenic | -8.71 | Deleterious | 1.000 | Probably Damaging | 0.996 | Probably Damaging | 3.33 | Benign | 0.01 | Affected | 0.2949 | 0.2399 | 0 | -2 | 3.8 | -60.04 | |||||||||||||||||||||||||||||
| c.1332G>C | K444N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K444N is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect are REVEL and FATHMM, whereas the majority of tools (FoldX, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus) predict a pathogenic impact. The high‑accuracy methods give the following results: AlphaMissense‑Optimized predicts pathogenic; the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates likely pathogenic; Foldetta’s stability assessment is uncertain and therefore not used as evidence. Overall, the preponderance of evidence points to a pathogenic effect for K444N. This conclusion is not contradicted by ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.203355 | Structured | 0.262172 | Uncertain | 0.955 | 0.213 | 0.000 | -14.797 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 2.41 | Destabilizing | 0.0 | 1.52 | Ambiguous | 1.97 | Ambiguous | 1.18 | Destabilizing | 0.286 | Likely Benign | -4.77 | Deleterious | 1.000 | Probably Damaging | 0.996 | Probably Damaging | 3.39 | Benign | 0.01 | Affected | 0.3282 | 0.1213 | 1 | 0 | 0.4 | -14.07 | |||||||||||||||||||||||||||||
| c.1332G>T | K444N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K444N is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect are REVEL and FATHMM. Tools that predict a pathogenic effect include FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Uncertain predictions come from Rosetta and Foldetta. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points to a pathogenic impact. There is no ClinVar annotation to contradict this assessment, so the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.203355 | Structured | 0.262172 | Uncertain | 0.955 | 0.213 | 0.000 | -14.797 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 2.41 | Destabilizing | 0.0 | 1.52 | Ambiguous | 1.97 | Ambiguous | 1.18 | Destabilizing | 0.286 | Likely Benign | -4.77 | Deleterious | 1.000 | Probably Damaging | 0.996 | Probably Damaging | 3.39 | Benign | 0.01 | Affected | 0.3282 | 0.1213 | 1 | 0 | 0.4 | -14.07 | |||||||||||||||||||||||||||||
| c.1444C>G | L482V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L482V has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include SIFT and AlphaMissense‑Optimized, whereas a majority of tools (SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default) predict a pathogenic impact. Stability‑based methods (FoldX, Rosetta, Foldetta, premPS) are inconclusive and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—indicates pathogenicity; Foldetta remains uncertain. Overall, the preponderance of evidence points to a pathogenic effect for the variant, and this conclusion is not contradicted by any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.254060 | Structured | 0.426236 | Uncertain | 0.795 | 0.248 | 0.000 | -11.676 | Likely Pathogenic | 0.734 | Likely Pathogenic | Likely Benign | 1.97 | Ambiguous | 0.1 | 1.52 | Ambiguous | 1.75 | Ambiguous | 0.76 | Ambiguous | 0.709 | Likely Pathogenic | -2.95 | Deleterious | 0.989 | Probably Damaging | 0.984 | Probably Damaging | -1.30 | Pathogenic | 0.10 | Tolerated | 0.1074 | 0.2027 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.1848T>A | D616E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D616E missense variant is catalogued in gnomAD (ID 6‑33440900‑T‑A) but has no ClinVar submission. Functional prediction tools show a split assessment: benign calls come from REVEL, both polyPhen‑2 HumDiv and HumVar, FATHMM, and AlphaMissense‑Optimized, whereas pathogenic calls arise from PROVEAN, SIFT, and AlphaMissense‑Default. The remaining predictors (FoldX, Rosetta, Foldetta, premPS, ESM1b) are inconclusive. A high‑accuracy consensus (SGM) that aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN yields a pathogenic verdict. AlphaMissense‑Optimized remains benign, and Foldetta, which evaluates protein‑folding stability, is uncertain. Overall, the majority of evidence leans toward pathogenicity, and this conclusion does not conflict with ClinVar because no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.129801 | Structured | 0.166689 | Uncertain | 0.867 | 0.252 | 0.000 | 6-33440900-T-A | 1 | 6.20e-7 | -7.250 | In-Between | 0.695 | Likely Pathogenic | Likely Benign | 0.96 | Ambiguous | 0.1 | 1.52 | Ambiguous | 1.24 | Ambiguous | 0.58 | Ambiguous | 0.092 | Likely Benign | -2.85 | Deleterious | 0.421 | Benign | 0.232 | Benign | 3.32 | Benign | 0.03 | Affected | 3.37 | 35 | 0.1225 | 0.4128 | 2 | 3 | 0.0 | 14.03 | |||||||||||||||||||||||||
| c.1848T>G | D616E 2D  3DClick to see structure in 3D Viewer AISynGAP1 D616E is not reported in ClinVar but is present in gnomAD (ID 6‑33440900‑T‑G). Functional prediction tools that agree on benign impact include REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, FATHMM, and AlphaMissense‑Optimized. Those that predict pathogenicity are PROVEAN, SIFT, and AlphaMissense‑Default. The remaining tools (FoldX, Rosetta, Foldetta, premPS, ESM1b) are inconclusive. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the majority of conventional predictors lean toward a benign effect, but the high‑accuracy consensus is split, leaving the variant’s clinical significance unresolved. Thus, the variant is most likely benign based on the bulk of predictions, and this does not contradict any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.129801 | Structured | 0.166689 | Uncertain | 0.867 | 0.252 | 0.000 | 6-33440900-T-G | 3 | 1.86e-6 | -7.250 | In-Between | 0.695 | Likely Pathogenic | Likely Benign | 0.96 | Ambiguous | 0.1 | 1.52 | Ambiguous | 1.24 | Ambiguous | 0.58 | Ambiguous | 0.092 | Likely Benign | -2.85 | Deleterious | 0.421 | Benign | 0.232 | Benign | 3.32 | Benign | 0.03 | Affected | 3.37 | 35 | 0.1225 | 0.4128 | 2 | 3 | 0.0 | 14.03 | |||||||||||||||||||||||||
| c.1910C>A | S637Y 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S637Y is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that indicate a benign effect include REVEL, FoldX, premPS, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect comprise SGM‑Consensus, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely pathogenic (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN), and Foldetta as uncertain. Overall, the majority of individual predictors (seven versus five) lean toward pathogenicity, and the consensus‑based SGM‑Consensus also supports a likely pathogenic classification. Therefore, the variant is most likely pathogenic based on the available predictions, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.076542 | Structured | 0.083482 | Uncertain | 0.920 | 0.253 | 0.000 | -12.633 | Likely Pathogenic | 0.770 | Likely Pathogenic | Likely Benign | 0.13 | Likely Benign | 0.1 | 1.52 | Ambiguous | 0.83 | Ambiguous | 0.50 | Likely Benign | 0.209 | Likely Benign | -3.78 | Deleterious | 0.985 | Probably Damaging | 0.681 | Possibly Damaging | 3.35 | Benign | 0.00 | Affected | 0.0968 | 0.4071 | -3 | -2 | -0.5 | 76.10 | |||||||||||||||||||||||||||||
| c.2015C>T | T672M 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant T672M is listed in ClinVar with an uncertain significance and is present in gnomAD (ID 6‑33441274‑C‑T). Prediction tools that classify the variant as benign include REVEL, FoldX, premPS, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict pathogenicity are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and ESM1b. Rosetta and Foldetta report uncertain results, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is inconclusive due to a 2‑to‑2 split. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while SGM Consensus and Foldetta remain unavailable. Overall, the balance of evidence favors a benign effect, and this conclusion does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.116183 | Structured | 0.102069 | Uncertain | 0.586 | 0.362 | 0.000 | Conflicting | 3 | 6-33441274-C-T | 19 | 1.18e-5 | -9.472 | Likely Pathogenic | 0.174 | Likely Benign | Likely Benign | 0.31 | Likely Benign | 0.4 | 1.52 | Ambiguous | 0.92 | Ambiguous | 0.41 | Likely Benign | 0.127 | Likely Benign | -4.34 | Deleterious | 0.993 | Probably Damaging | 0.520 | Possibly Damaging | 3.39 | Benign | 0.00 | Affected | 3.40 | 25 | 0.1332 | 0.6677 | -1 | -1 | 2.6 | 30.09 | 231.9 | -52.9 | 1.1 | 0.1 | 0.5 | 0.0 | X | X | Potentially Pathogenic | The hydroxyl group of Thr672, located in an entangled α-α loop connecting the two α-helices (res. Ser641-Glu666 and res. Leu685-Val699), is involved in a highly coordinated hydrogen-bonding network between residues from two α-helices (res. Ser641-Glu666 and res. Arg563-Glu578) and from the α-α loop itself, such as Lys566, Glu666, and Asn669. Met672 can only form a hydrogen bond with the amino group of the Lys566 side chain via its backbone carbonyl group. Nevertheless, the Lys566-Glu666 salt bridge forms intermittently. This is possible because Asn669 keeps the carboxylate group of Glu666 in the vicinity through hydrogen bonding, and the hydrophobic side chain of Met stays mostly rotated away from the salt bridge. Consequently, no drastic disruption of the hydrogen-bond network that keeps the loop close to the helices occurs in the variant simulations. | |||||||||||||
| c.2120C>G | A707G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 A707G missense variant has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict a pathogenic outcome are PROVEAN, polyPhen‑2 (HumDiv and HumVar), and SIFT. The remaining tools (FoldX, Rosetta, Foldetta, premPS, ESM1b) give uncertain or inconclusive results and are treated as unavailable evidence. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also resolves to benign, while Foldetta remains uncertain. Overall, the majority of definitive predictions lean toward a benign effect, and there is no ClinVar status to contradict this assessment. Thus, the variant is most likely benign. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.203355 | Structured | 0.371229 | Uncertain | 0.927 | 0.365 | 0.000 | -7.480 | In-Between | 0.224 | Likely Benign | Likely Benign | 1.45 | Ambiguous | 0.0 | 1.52 | Ambiguous | 1.49 | Ambiguous | 0.62 | Ambiguous | 0.198 | Likely Benign | -2.60 | Deleterious | 0.991 | Probably Damaging | 0.960 | Probably Damaging | 3.42 | Benign | 0.01 | Affected | 0.1678 | 0.2388 | 1 | 0 | -2.2 | -14.03 | ||||||||||||||||||||||||||||||
| c.728T>C | I243T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I243T is not reported in ClinVar and is absent from gnomAD. Prediction tools largely agree on a deleterious effect: SGM‑Consensus, REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default all classify the variant as pathogenic. Only FATHMM predicts a benign outcome, while Foldetta, AlphaMissense‑Optimized, and Rosetta return uncertain results, which are treated as unavailable evidence. High‑accuracy assessments further support a pathogenic interpretation: AlphaMissense‑Optimized is uncertain, SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is Likely Pathogenic, and Foldetta remains uncertain. Overall, the preponderance of evidence indicates that I243T is most likely pathogenic, and this conclusion does not contradict the current ClinVar status, which contains no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.363090 | Structured | 0.344471 | Uncertain | 0.842 | 0.347 | 0.000 | -9.102 | Likely Pathogenic | 0.922 | Likely Pathogenic | Ambiguous | 2.15 | Destabilizing | 0.2 | 1.52 | Ambiguous | 1.84 | Ambiguous | 1.72 | Destabilizing | 0.816 | Likely Pathogenic | -3.06 | Deleterious | 0.982 | Probably Damaging | 0.702 | Possibly Damaging | 5.55 | Benign | 0.01 | Affected | 0.1027 | 0.0541 | 0 | -1 | -5.2 | -12.05 | |||||||||||||||||||||||||||||
| c.736C>G | L246V 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L246V is reported in gnomAD (variant ID 6‑33435587‑C‑G) but has no ClinVar entry. Functional prediction tools largely agree on a deleterious effect: pathogenic calls are made by SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. Only FATHMM predicts a benign outcome. Uncertain results are reported by Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as likely pathogenic, and Foldetta as uncertain. The overwhelming majority of evidence points to a pathogenic impact, and this conclusion is not contradicted by ClinVar, which contains no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.472492 | Structured | 0.302312 | Uncertain | 0.859 | 0.364 | 0.000 | 6-33435587-C-G | 1 | 6.20e-7 | -12.092 | Likely Pathogenic | 0.935 | Likely Pathogenic | Ambiguous | 2.09 | Destabilizing | 0.1 | 1.52 | Ambiguous | 1.81 | Ambiguous | 1.13 | Destabilizing | 0.736 | Likely Pathogenic | -2.60 | Deleterious | 0.930 | Possibly Damaging | 0.504 | Possibly Damaging | 4.71 | Benign | 0.01 | Affected | 3.41 | 14 | 0.1434 | 0.3607 | 1 | 2 | 0.4 | -14.03 | ||||||||||||||||||||||||
| c.923G>T | W308L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant W308L is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that assess pathogenicity unanimously classify the variant as deleterious: REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict pathogenicity. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also indicates “Likely Pathogenic.” Only Rosetta and premPS yield uncertain results, which are treated as unavailable evidence. High‑accuracy assessments reinforce the pathogenic prediction: AlphaMissense‑Optimized is pathogenic, SGM‑Consensus is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. No tools predict benign effects. **Based on the collective predictions, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none is available).** Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.328603 | Structured | 0.333942 | Uncertain | 0.907 | 0.314 | 0.125 | -13.349 | Likely Pathogenic | 0.991 | Likely Pathogenic | Likely Pathogenic | 3.53 | Destabilizing | 0.5 | 1.52 | Ambiguous | 2.53 | Destabilizing | 0.53 | Ambiguous | 0.811 | Likely Pathogenic | -11.95 | Deleterious | 0.999 | Probably Damaging | 0.996 | Probably Damaging | 0.49 | Pathogenic | 0.00 | Affected | 0.2731 | 0.2754 | -2 | -2 | 4.7 | -73.05 | |||||||||||||||||||||||||||||
| c.1786C>A | R596S 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant R596S is not reported in ClinVar and is absent from gnomAD. In silico assessment shows a consensus of pathogenicity: all evaluated tools (REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 HumDiv/HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a deleterious effect, while Rosetta remains uncertain. Grouping by agreement, no tool predicts benign; the pathogenic group includes 13 predictions, with Rosetta excluded as inconclusive. High‑accuracy methods further support a damaging outcome: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. Consequently, the variant is most likely pathogenic based on the available predictions, and this assessment does not contradict ClinVar status, as no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.017797 | Structured | 0.135423 | Uncertain | 0.918 | 0.134 | 0.000 | -13.921 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 3.51 | Destabilizing | 0.2 | 1.53 | Ambiguous | 2.52 | Destabilizing | 1.17 | Destabilizing | 0.626 | Likely Pathogenic | -5.97 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 2.42 | Pathogenic | 0.00 | Affected | 0.3129 | 0.2680 | 0 | -1 | 3.7 | -69.11 | |||||||||||||||||||||||||||||
| c.2012A>G | D671G 2D  AIThe SynGAP1 D671G missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, and FATHMM, whereas a majority of tools predict a pathogenic impact: SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as Likely Pathogenic (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN), and Foldetta as uncertain. No evidence from these tools contradicts the pathogenic prediction. Overall, the balance of evidence favors a pathogenic classification for D671G, and this conclusion is consistent with the absence of a ClinVar entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.194234 | Structured | 0.096749 | Uncertain | 0.677 | 0.370 | 0.000 | -10.346 | Likely Pathogenic | 0.857 | Likely Pathogenic | Ambiguous | 0.50 | Ambiguous | 0.5 | 1.53 | Ambiguous | 1.02 | Ambiguous | 0.00 | Likely Benign | 0.279 | Likely Benign | -4.78 | Deleterious | 0.995 | Probably Damaging | 0.946 | Probably Damaging | 3.45 | Benign | 0.01 | Affected | 0.4032 | 0.5823 | 1 | -1 | 3.1 | -58.04 | |||||||||||||||||||||||||||||
| c.622C>T | P208S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P208S missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. In contrast, the majority of tools predict a pathogenic impact: SGM‑Consensus, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and Foldetta. Rosetta’s output is uncertain and therefore not counted as evidence. High‑accuracy methods give a consistent pathogenic signal: AlphaMissense‑Optimized predicts benign, but the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) resolves to pathogenic, and Foldetta also predicts pathogenic. With no ClinVar annotation to contradict, the overall evidence strongly favors a pathogenic classification for P208S. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.271506 | Structured | 0.399506 | Uncertain | 0.864 | 0.345 | 0.125 | -8.363 | Likely Pathogenic | 0.587 | Likely Pathogenic | Likely Benign | 2.68 | Destabilizing | 0.3 | 1.53 | Ambiguous | 2.11 | Destabilizing | 1.21 | Destabilizing | 0.270 | Likely Benign | -6.78 | Deleterious | 1.000 | Probably Damaging | 0.994 | Probably Damaging | 3.79 | Benign | 0.02 | Affected | 0.3808 | 0.4537 | 1 | -1 | 0.8 | -10.04 | |||||||||||||||||||||||||||||
| c.655T>A | C219S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C219S is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: benign calls come from FoldX, polyPhen‑2 HumVar, and FATHMM, while pathogenic predictions are made by SGM‑Consensus (Likely Pathogenic), REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments reinforce this trend: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates pathogenicity, and Foldetta, which integrates FoldX‑MD and Rosetta outputs, is inconclusive. With the majority of evidence pointing to a damaging effect and no ClinVar annotation to contradict, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.254060 | Structured | 0.426845 | Uncertain | 0.903 | 0.279 | 0.000 | -12.962 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 0.09 | Likely Benign | 0.4 | 1.53 | Ambiguous | 0.81 | Ambiguous | 1.56 | Destabilizing | 0.892 | Likely Pathogenic | -8.35 | Deleterious | 0.900 | Possibly Damaging | 0.380 | Benign | 5.95 | Benign | 0.04 | Affected | 0.3568 | 0.1454 | 0 | -1 | -3.3 | -16.06 | |||||||||||||||||||||||||||||
| c.656G>C | C219S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C219S is not reported in ClinVar and is absent from gnomAD. Prediction tools that classify the variant as benign include FoldX, polyPhen‑2 HumVar, and FATHMM, whereas the majority of tools predict it to be pathogenic: SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates pathogenicity, and Foldetta’s stability analysis is inconclusive (treated as unavailable). Taken together, the preponderance of evidence from both general and high‑accuracy predictors points to a pathogenic impact for C219S. This conclusion is not contradicted by ClinVar status, which has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.254060 | Structured | 0.426845 | Uncertain | 0.903 | 0.279 | 0.000 | -12.962 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 0.09 | Likely Benign | 0.4 | 1.53 | Ambiguous | 0.81 | Ambiguous | 1.56 | Destabilizing | 0.817 | Likely Pathogenic | -8.35 | Deleterious | 0.900 | Possibly Damaging | 0.380 | Benign | 5.95 | Benign | 0.04 | Affected | 0.3568 | 0.1454 | 0 | -1 | -3.3 | -16.06 | |||||||||||||||||||||||||||||
| c.775C>G | R259G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R259G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: FATHMM is the sole benign predictor, whereas the remaining 13 tools—including SGM‑Consensus, REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—classify the change as pathogenic. The high‑accuracy subset reinforces this assessment: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) also indicates pathogenic. Rosetta alone is uncertain and is treated as unavailable. Overall, the consensus of available predictions indicates that R259G is most likely pathogenic, and this conclusion does not contradict any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.222385 | Structured | 0.338208 | Uncertain | 0.885 | 0.255 | 0.250 | -15.389 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 3.17 | Destabilizing | 0.6 | 1.53 | Ambiguous | 2.35 | Destabilizing | 1.14 | Destabilizing | 0.762 | Likely Pathogenic | -6.43 | Deleterious | 0.997 | Probably Damaging | 0.987 | Probably Damaging | 5.78 | Benign | 0.00 | Affected | 0.3342 | 0.4139 | -3 | -2 | 4.1 | -99.14 | |||||||||||||||||||||||||||||
| c.893C>T | P298L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P298L missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, SIFT, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict a pathogenic outcome are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. The remaining tools—FoldX, Rosetta, Foldetta, and ESM1b—return uncertain or inconclusive results. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized is benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) yields a benign outcome; and Foldetta’s stability prediction is unavailable. Overall, the majority of evidence points to a benign effect, and this conclusion does not contradict any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.328603 | Structured | 0.268765 | Uncertain | 0.860 | 0.283 | 0.500 | -7.334 | In-Between | 0.107 | Likely Benign | Likely Benign | 0.60 | Ambiguous | 0.2 | 1.53 | Ambiguous | 1.07 | Ambiguous | -0.16 | Likely Benign | 0.267 | Likely Benign | -0.82 | Neutral | 0.885 | Possibly Damaging | 0.589 | Possibly Damaging | 1.91 | Pathogenic | 0.21 | Tolerated | 0.2137 | 0.6795 | -3 | -3 | 5.4 | 16.04 | ||||||||||||||||||||||||||||||
| c.1103C>G | P368R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P368R missense variant is not reported in ClinVar (status: None) and has no entry in gnomAD. Prediction tools that agree on a benign effect include REVEL and AlphaMissense‑Optimized. Tools that agree on a pathogenic effect include SGM‑Consensus, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. Uncertain or inconclusive results come from FoldX, Rosetta, Foldetta, and premPS. High‑accuracy methods give a mixed picture: AlphaMissense‑Optimized predicts benign, whereas the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts likely pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, is uncertain. Overall, the majority of evidence—including the SGM‑Consensus and several individual high‑accuracy tools—points to a pathogenic effect. This conclusion is not contradicted by ClinVar status, which is currently unreported. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.363090 | Structured | 0.439989 | Uncertain | 0.580 | 0.677 | 0.250 | -9.564 | Likely Pathogenic | 0.736 | Likely Pathogenic | Likely Benign | 1.57 | Ambiguous | 1.0 | 1.54 | Ambiguous | 1.56 | Ambiguous | 0.58 | Ambiguous | 0.263 | Likely Benign | -6.07 | Deleterious | 0.991 | Probably Damaging | 0.881 | Possibly Damaging | 1.78 | Pathogenic | 0.00 | Affected | 0.1439 | 0.3922 | 0 | -2 | -2.9 | 59.07 | |||||||||||||||||||||||||||||
| c.1767C>G | I589M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I589M is listed in ClinVar with an uncertain significance (ClinVar ID 964298.0) and is not reported in gnomAD. Functional prediction tools that provide a definitive call overwhelmingly predict a deleterious effect: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default all indicate pathogenicity, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also reports a likely pathogenic outcome. Tools that are inconclusive—FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized—are listed as uncertain and do not influence the overall assessment. High‑accuracy methods specifically show AlphaMissense‑Optimized as uncertain, SGM Consensus as likely pathogenic, and Foldetta as uncertain. Taken together, the majority of available predictions support a pathogenic effect, which is consistent with the ClinVar uncertain designation rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.018415 | Structured | 0.084536 | Uncertain | 0.927 | 0.214 | 0.000 | Uncertain | 1 | -12.225 | Likely Pathogenic | 0.926 | Likely Pathogenic | Ambiguous | 0.74 | Ambiguous | 0.2 | 1.54 | Ambiguous | 1.14 | Ambiguous | 1.33 | Destabilizing | 0.830 | Likely Pathogenic | -2.99 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.94 | Pathogenic | 0.00 | Affected | 3.37 | 35 | 0.0909 | 0.2552 | 2 | 1 | -2.6 | 18.03 | 267.6 | -24.5 | 0.0 | 0.0 | -0.1 | 0.1 | X | Potentially Benign | A hydrophobic residue, Ile589, located in an α helix (res. Glu582-Met603), is swapped for another hydrophobic residue, methionine. The sec-butyl hydrocarbon side chain of Ile589 packs favourably with multiple residues in the inter-helix hydrophobic space (e.g., Phe569, Ile667, and Leu664).Although the S-methyl thioether group of the Met589 side chain in the variant is longer than the branched side chain of isoleucine, it stacks favourably with the aromatic phenol ring. Additionally, the polar sulphur atom forms a weak hydrogen bond with the guanidinium group of Arg573, which in turn forms a salt bridge with the carboxylate group of Asp586.Overall, the hydrophobic packing in the inter-helix space does not appear to be disrupted in the variant simulations. | ||||||||||||||||
| c.952C>A | P318T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P318T is not reported in ClinVar and is absent from gnomAD. Prediction tools that assess pathogenicity uniformly favor a deleterious effect: REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) all predict pathogenic or likely pathogenic. No tool in the dataset predicts a benign outcome; the only inconclusive results come from Rosetta, Foldetta, and premPS, which are treated as unavailable evidence. High‑accuracy assessments confirm this trend: AlphaMissense‑Optimized is pathogenic, SGM‑Consensus is likely pathogenic, and Foldetta is uncertain. Consequently, the variant is most likely pathogenic based on the available predictions, and this assessment does not contradict any ClinVar status because none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.111485 | Structured | 0.400936 | Uncertain | 0.858 | 0.234 | 0.000 | -10.759 | Likely Pathogenic | 0.962 | Likely Pathogenic | Likely Pathogenic | 2.03 | Destabilizing | 0.2 | 1.54 | Ambiguous | 1.79 | Ambiguous | 0.84 | Ambiguous | 0.680 | Likely Pathogenic | -7.09 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.84 | Pathogenic | 0.01 | Affected | 0.1583 | 0.6044 | 0 | -1 | 0.9 | 3.99 | |||||||||||||||||||||||||||||
| c.1019C>A | A340E 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant A340E is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, SIFT, and polyPhen‑2 HumVar. Those that predict a pathogenic effect are polyPhen‑2 HumDiv, ESM1b, FATHMM, and AlphaMissense‑Default. Predictions that are inconclusive or unavailable are FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. The high‑accuracy consensus from SGM (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) reports a likely pathogenic outcome, while the protein‑folding stability method Foldetta is uncertain. AlphaMissense‑Optimized also yields an uncertain result. Overall, the majority of evidence points toward a pathogenic impact, and this is consistent with the lack of ClinVar annotation; there is no contradiction with ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.390993 | Structured | 0.410781 | Uncertain | 0.558 | 0.485 | 0.250 | -8.225 | Likely Pathogenic | 0.803 | Likely Pathogenic | Ambiguous | 0.63 | Ambiguous | 0.4 | 1.55 | Ambiguous | 1.09 | Ambiguous | 0.06 | Likely Benign | 0.138 | Likely Benign | -0.33 | Neutral | 0.625 | Possibly Damaging | 0.252 | Benign | 1.91 | Pathogenic | 0.39 | Tolerated | 0.1418 | 0.2160 | 0 | -1 | -5.3 | 58.04 | |||||||||||||||||||||||||||||
| c.1060G>C | A354P 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant A354P is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are FoldX, SIFT, and FATHMM, while Rosetta and premPS are inconclusive. The high‑accuracy consensus from AlphaMissense‑Optimized and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) both indicate a benign outcome, whereas Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, predicts a pathogenic impact. Overall, the majority of evidence points to a benign effect for A354P, and this assessment does not contradict any ClinVar annotation because no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.203355 | Structured | 0.381329 | Uncertain | 0.882 | 0.335 | 0.125 | -5.971 | Likely Benign | 0.239 | Likely Benign | Likely Benign | 2.66 | Destabilizing | 0.2 | 1.55 | Ambiguous | 2.11 | Destabilizing | 0.57 | Ambiguous | 0.241 | Likely Benign | -1.66 | Neutral | 0.001 | Benign | 0.002 | Benign | 1.76 | Pathogenic | 0.04 | Affected | 0.2259 | 0.5884 | 1 | -1 | -3.4 | 26.04 | |||||||||||||||||||||||||||||
| c.1359C>A | H453Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant H453Q is not reported in ClinVar (ClinVar ID None) and has no entries in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, SIFT, and FATHMM, whereas a majority of tools predict a pathogenic outcome: SGM‑Consensus (Likely Pathogenic), PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Default. Tools with uncertain or inconclusive results—FoldX, Rosetta, premPS, AlphaMissense‑Optimized, and Foldetta—are treated as unavailable for pathogenicity assessment. High‑accuracy methods specifically show SGM‑Consensus as Likely Pathogenic, AlphaMissense‑Optimized as uncertain, and Foldetta as uncertain. Overall, the preponderance of evidence from consensus and high‑accuracy predictors indicates that H453Q is most likely pathogenic, and this assessment does not contradict the current ClinVar status, which contains no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.352862 | Structured | 0.316097 | Uncertain | 0.946 | 0.200 | 0.000 | -10.894 | Likely Pathogenic | 0.893 | Likely Pathogenic | Ambiguous | 0.90 | Ambiguous | 0.1 | 1.55 | Ambiguous | 1.23 | Ambiguous | 0.92 | Ambiguous | 0.259 | Likely Benign | -7.91 | Deleterious | 1.000 | Probably Damaging | 0.993 | Probably Damaging | 3.45 | Benign | 0.09 | Tolerated | 0.1223 | 0.3112 | 3 | 0 | -0.3 | -9.01 | |||||||||||||||||||||||||||||
| c.1359C>G | H453Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant H453Q is not reported in ClinVar (ClinVar ID None) and has no entries in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, SIFT, and FATHMM, whereas a majority of tools predict a pathogenic outcome: SGM‑Consensus (Likely Pathogenic), PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Default. Tools with uncertain or inconclusive results—FoldX, Rosetta, Foldetta, premPS, and AlphaMissense‑Optimized—are treated as unavailable for pathogenicity assessment. High‑accuracy methods specifically show SGM‑Consensus as Likely Pathogenic, AlphaMissense‑Optimized as Uncertain, and Foldetta as Uncertain. Overall, the preponderance of evidence points to a pathogenic effect for H453Q. This prediction does not contradict ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.352862 | Structured | 0.316097 | Uncertain | 0.946 | 0.200 | 0.000 | -10.894 | Likely Pathogenic | 0.893 | Likely Pathogenic | Ambiguous | 0.90 | Ambiguous | 0.1 | 1.55 | Ambiguous | 1.23 | Ambiguous | 0.92 | Ambiguous | 0.258 | Likely Benign | -7.91 | Deleterious | 1.000 | Probably Damaging | 0.993 | Probably Damaging | 3.45 | Benign | 0.09 | Tolerated | 0.1223 | 0.3112 | 3 | 0 | -0.3 | -9.01 | |||||||||||||||||||||||||||||
| c.1463C>G | T488R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant T488R is not reported in ClinVar and is absent from gnomAD. Prediction tools that indicate a benign effect include only FATHMM, whereas the remaining tools (REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus) all predict a pathogenic outcome. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Based on the collective evidence, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.206376 | Structured | 0.332663 | Uncertain | 0.928 | 0.233 | 0.125 | -14.353 | Likely Pathogenic | 0.988 | Likely Pathogenic | Likely Pathogenic | 1.29 | Ambiguous | 0.3 | 1.55 | Ambiguous | 1.42 | Ambiguous | 0.85 | Ambiguous | 0.726 | Likely Pathogenic | -5.70 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.22 | Benign | 0.00 | Affected | 0.0711 | 0.2048 | -1 | -1 | -3.8 | 55.08 | |||||||||||||||||||||||||||||
| c.1468G>T | A490S 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant A490S is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include AlphaMissense‑Optimized, whereas the majority of other in silico predictors (SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM) indicate a pathogenic impact. High‑accuracy assessments further support this: AlphaMissense‑Optimized classifies the variant as benign, SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts it to be likely pathogenic, and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, yields an inconclusive result. No other tools provide definitive evidence. Overall, the preponderance of pathogenic predictions suggests that A490S is most likely pathogenic, and this assessment does not contradict the current ClinVar status, which has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.120615 | Structured | 0.322979 | Uncertain | 0.938 | 0.210 | 0.125 | -8.307 | Likely Pathogenic | 0.426 | Ambiguous | Likely Benign | 0.76 | Ambiguous | 0.1 | 1.55 | Ambiguous | 1.16 | Ambiguous | 0.89 | Ambiguous | 0.766 | Likely Pathogenic | -2.82 | Deleterious | 0.983 | Probably Damaging | 0.993 | Probably Damaging | -1.41 | Pathogenic | 0.02 | Affected | 0.2338 | 0.3116 | 1 | 1 | -2.6 | 16.00 | |||||||||||||||||||||||||||||
| c.1504G>T | G502C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G502C lies in the GAP domain. ClinVar has no entry for this variant, and it is not reported in gnomAD. Prediction tools that agree on a benign effect include only premPS. The majority of tools predict a pathogenic effect: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus score. Tools with uncertain or inconclusive outputs are FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (derived from a unanimous majority of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta) as uncertain. Overall, the preponderance of evidence points to a pathogenic impact for G502C, and this conclusion does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.083462 | Structured | 0.340113 | Uncertain | 0.882 | 0.152 | 0.000 | -12.086 | Likely Pathogenic | 0.907 | Likely Pathogenic | Ambiguous | 1.02 | Ambiguous | 0.5 | 1.55 | Ambiguous | 1.29 | Ambiguous | 0.30 | Likely Benign | 0.845 | Likely Pathogenic | -8.65 | Deleterious | 1.000 | Probably Damaging | 0.988 | Probably Damaging | -1.67 | Pathogenic | 0.00 | Affected | 0.1279 | 0.2691 | -3 | -3 | 2.9 | 46.09 | |||||||||||||||||||||||||||||
| c.1570T>A | C524S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C524S is listed in gnomAD (variant ID 6‑33438813‑T‑A) but has no ClinVar entry. Prediction tools that assess pathogenicity all converge on a deleterious effect: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all report a pathogenic outcome, while FoldX, Rosetta, and Foldetta are uncertain and therefore not counted as evidence. Grouping by agreement yields a benign‑prediction set that is empty and a pathogenic‑prediction set that contains the eleven tools above. High‑accuracy assessments reinforce this: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic; Foldetta remains uncertain. Consequently, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.134866 | Structured | 0.024729 | Uncertain | 0.916 | 0.385 | 0.125 | 6-33438813-T-A | 1 | 6.20e-7 | -11.174 | Likely Pathogenic | 0.996 | Likely Pathogenic | Likely Pathogenic | 0.80 | Ambiguous | 0.1 | 1.55 | Ambiguous | 1.18 | Ambiguous | 1.58 | Destabilizing | 0.915 | Likely Pathogenic | -9.94 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.38 | Pathogenic | 0.00 | Affected | 3.37 | 35 | 0.5362 | 0.1848 | Weaken | -1 | 0 | -3.3 | -16.06 | |||||||||||||||||||||||
| c.1571G>C | C524S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 C524S variant has no ClinVar entry and is not present in gnomAD. Prediction tools that agree on a benign effect are none; those that predict pathogenicity include REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. FoldX, Rosetta, and Foldetta returned inconclusive results and are treated as unavailable evidence. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely pathogenic, and Foldetta as inconclusive. Overall, the preponderance of evidence points to a pathogenic effect. This conclusion is not contradicted by ClinVar status, which simply lacks an entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.134866 | Structured | 0.024729 | Uncertain | 0.916 | 0.385 | 0.125 | -11.174 | Likely Pathogenic | 0.996 | Likely Pathogenic | Likely Pathogenic | 0.80 | Ambiguous | 0.1 | 1.55 | Ambiguous | 1.18 | Ambiguous | 1.58 | Destabilizing | 0.904 | Likely Pathogenic | -9.94 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.38 | Pathogenic | 0.00 | Affected | 3.37 | 35 | 0.5362 | 0.1848 | Weaken | -1 | 0 | -3.3 | -16.06 | ||||||||||||||||||||||||||
| c.1739G>C | G580A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G580A is not reported in ClinVar (ClinVar status: None) and has no entries in gnomAD (gnomAD ID: None). Prediction tools that assess pathogenicity are unanimous: REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus all predict a deleterious effect. Tools with uncertain or inconclusive results (Rosetta and premPS) are noted as unavailable for pathogenicity inference. High‑accuracy methods reinforce this view: AlphaMissense‑Optimized reports Pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates Likely Pathogenic; and Foldetta, a protein‑folding stability predictor that integrates FoldX‑MD and Rosetta outputs, also classifies the variant as Pathogenic. Consequently, the variant is most likely pathogenic based on the consensus of predictive tools, and this assessment does not contradict any ClinVar status, as none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.104810 | Structured | 0.025952 | Uncertain | 0.853 | 0.236 | 0.000 | -10.841 | Likely Pathogenic | 0.956 | Likely Pathogenic | Likely Pathogenic | 2.84 | Destabilizing | 0.1 | 1.55 | Ambiguous | 2.20 | Destabilizing | 0.64 | Ambiguous | 0.646 | Likely Pathogenic | -5.73 | Deleterious | 0.999 | Probably Damaging | 0.995 | Probably Damaging | -1.22 | Pathogenic | 0.05 | Affected | 0.3657 | 0.2862 | 1 | 0 | 2.2 | 14.03 | |||||||||||||||||||||||||||||
| c.1891C>G | Q631E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant Q631E is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Prediction tools that indicate a benign effect include FoldX, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect comprise SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized classifying the variant as benign, SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) classifying it as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) yielding an uncertain result. Overall, the majority of evidence (eight pathogenic predictions versus three benign) points to a pathogenic impact. This conclusion is not contradicted by ClinVar status, which is currently unavailable. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.041405 | Structured | 0.038963 | Uncertain | 0.948 | 0.230 | 0.000 | -15.628 | Likely Pathogenic | 0.782 | Likely Pathogenic | Likely Benign | 0.04 | Likely Benign | 0.1 | 1.55 | Ambiguous | 0.80 | Ambiguous | 0.95 | Ambiguous | 0.532 | Likely Pathogenic | -2.99 | Deleterious | 0.997 | Probably Damaging | 0.981 | Probably Damaging | 2.78 | Benign | 0.01 | Affected | 0.1068 | 0.1264 | 2 | 2 | 0.0 | 0.98 | |||||||||||||||||||||||||||||
| c.2123T>A | L708H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L708H is not reported in ClinVar and is absent from gnomAD. Prediction tools that indicate a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect comprise premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, and AlphaMissense‑Default; the SGM Consensus score is also labeled Likely Pathogenic. Stability‑based methods FoldX and Rosetta return uncertain results, and Foldetta likewise is inconclusive. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus as Likely Pathogenic, and Foldetta as uncertain. Overall, the majority of conventional predictors lean toward pathogenicity, whereas a few high‑accuracy tools suggest benign or are inconclusive. Thus, the variant is most likely pathogenic based on the prevailing predictions, and this assessment does not contradict any ClinVar annotation because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.250310 | Structured | 0.365875 | Uncertain | 0.931 | 0.378 | 0.000 | -8.059 | Likely Pathogenic | 0.677 | Likely Pathogenic | Likely Benign | 1.84 | Ambiguous | 0.1 | 1.55 | Ambiguous | 1.70 | Ambiguous | 1.44 | Destabilizing | 0.243 | Likely Benign | -4.68 | Deleterious | 1.000 | Probably Damaging | 0.981 | Probably Damaging | 3.25 | Benign | 0.02 | Affected | 0.0824 | 0.0288 | -2 | -3 | -7.0 | 23.98 | |||||||||||||||||||||||||||||
| c.695C>A | A232D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A232D is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include FoldX, polyPhen‑2 HumVar, and FATHMM. Those that predict a pathogenic effect comprise REVEL, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus. Uncertain or inconclusive results are reported for Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also as pathogenic, while Foldetta remains uncertain. Overall, the majority of available predictions support a pathogenic impact. Thus, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which currently has no classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.254060 | Structured | 0.307228 | Uncertain | 0.878 | 0.305 | 0.000 | -13.956 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 0.14 | Likely Benign | 0.2 | 1.55 | Ambiguous | 0.85 | Ambiguous | 0.77 | Ambiguous | 0.725 | Likely Pathogenic | -2.50 | Deleterious | 0.845 | Possibly Damaging | 0.348 | Benign | 5.78 | Benign | 0.02 | Affected | 0.2066 | 0.2896 | 0 | -2 | -5.3 | 44.01 | |||||||||||||||||||||||||||||
| c.913A>G | T305A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 T305A variant is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33437818‑A‑G). Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus as benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Overall, the majority of evidence points to a benign impact, and this conclusion does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.359901 | Structured | 0.299706 | Uncertain | 0.872 | 0.274 | 0.125 | Conflicting | 2 | 6-33437818-A-G | 13 | 8.05e-6 | -4.307 | Likely Benign | 0.078 | Likely Benign | Likely Benign | 1.30 | Ambiguous | 0.6 | 1.55 | Ambiguous | 1.43 | Ambiguous | 0.77 | Ambiguous | 0.144 | Likely Benign | -2.10 | Neutral | 0.939 | Possibly Damaging | 0.645 | Possibly Damaging | 1.76 | Pathogenic | 0.12 | Tolerated | 3.40 | 20 | 0.4277 | 0.4403 | 1 | 0 | 2.5 | -30.03 | 177.9 | 43.5 | -0.2 | 0.1 | 0.4 | 0.0 | Uncertain | The hydroxyl group of Thr305, located at the beginning of an anti-parallel β strand (res. Thr305-Asn315), hydrogen bonds with the carboxylate groups of Glu270 and Asp304 in the anti-parallel β strand and the adjacent β hairpin loop, respectively. In the variant simulations, the methyl group of the Ala305 side chain cannot hydrogen bond with either of the acidic residues, which could weaken the integrity of the tertiary structure and the β hairpin loop. Indeed, the guanidinium group of Arg299 does not acquire its central hairpin loop position due to the residue swap.β hairpins are potential nucleation sites during the initial stages of protein folding, so even minor changes in them could be significant. Due to its location near the membrane surface, the residue swap could also affect the C2 loop dynamics and SynGAP-membrane association. However, this is beyond the scope of the solvent-only simulations to unravel. | ||||||||||||||
| c.958G>T | V320F 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant V320F is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on benign impact include REVEL, premPS, and SIFT, whereas tools that agree on pathogenic impact include PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as inconclusive, SGM‑Consensus (derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as inconclusive. Because the majority of available predictions and the SGM‑Consensus favor pathogenicity, the variant is most likely pathogenic. This conclusion does not contradict ClinVar status, as no ClinVar entry exists for V320F. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.185198 | Structured | 0.419626 | Uncertain | 0.905 | 0.266 | 0.125 | -9.958 | Likely Pathogenic | 0.877 | Likely Pathogenic | Ambiguous | 1.49 | Ambiguous | 1.4 | 1.55 | Ambiguous | 1.52 | Ambiguous | 0.44 | Likely Benign | 0.237 | Likely Benign | -3.26 | Deleterious | 0.994 | Probably Damaging | 0.944 | Probably Damaging | 1.79 | Pathogenic | 0.06 | Tolerated | 0.0473 | 0.3064 | -1 | -1 | -1.4 | 48.04 | |||||||||||||||||||||||||||||
| c.1186G>C | G396R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 G396R missense variant is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on benign impact include REVEL, polyPhen‑2 HumVar, SIFT, FATHMM, and AlphaMissense‑Optimized. Those that predict pathogenicity are PROVEAN, polyPhen‑2 HumDiv, ESM1b, AlphaMissense‑Default, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Four tools (FoldX, Rosetta, Foldetta, premPS) returned uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Overall, the variant is more frequently predicted to be pathogenic (five tools) than benign (five tools), and the high‑accuracy consensus leans toward pathogenicity, though Foldetta does not provide a definitive verdict. Thus, the variant is most likely pathogenic based on the current predictions, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.414856 | Structured | 0.394626 | Uncertain | 0.640 | 0.584 | 0.500 | -9.310 | Likely Pathogenic | 0.775 | Likely Pathogenic | Likely Benign | 1.68 | Ambiguous | 1.1 | 1.56 | Ambiguous | 1.62 | Ambiguous | 0.66 | Ambiguous | 0.319 | Likely Benign | -2.65 | Deleterious | 0.718 | Possibly Damaging | 0.216 | Benign | 4.42 | Benign | 0.24 | Tolerated | 0.0986 | 0.4007 | -3 | -2 | -4.1 | 99.14 | |||||||||||||||||||||||||||||
| c.1384A>G | K462E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K462E is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, SIFT, and FATHMM. Tools that agree on a pathogenic effect include SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Rosetta and Foldetta are uncertain and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized predicts pathogenic, SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts likely pathogenic, and Foldetta is uncertain. Overall, the majority of predictions (7 pathogenic vs. 5 benign) and the high‑accuracy tools support a pathogenic classification. Thus, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.264545 | Structured | 0.297737 | Uncertain | 0.921 | 0.159 | 0.125 | -14.696 | Likely Pathogenic | 0.967 | Likely Pathogenic | Likely Pathogenic | 0.34 | Likely Benign | 0.0 | 1.56 | Ambiguous | 0.95 | Ambiguous | 0.33 | Likely Benign | 0.433 | Likely Benign | -3.88 | Deleterious | 0.998 | Probably Damaging | 0.991 | Probably Damaging | 3.49 | Benign | 0.15 | Tolerated | 0.4145 | 0.1022 | 0 | 1 | 0.4 | 0.94 | |||||||||||||||||||||||||||||
| c.1831A>G | M611V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M611V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, PROVEAN, SIFT, AlphaMissense‑Default, and AlphaMissense‑Optimized; pathogenic predictions arise from polyPhen‑2 (HumDiv and HumVar), ESM1b, and FATHMM. High‑accuracy assessments show AlphaMissense‑Optimized classifying the variant as benign, while the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—yields an inconclusive result (two benign, two pathogenic). Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, reports an uncertain effect, so its result is treated as unavailable. Overall, the balance of evidence leans toward a benign impact, and this assessment does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.236433 | Structured | 0.210791 | Uncertain | 0.870 | 0.253 | 0.000 | -8.057 | Likely Pathogenic | 0.176 | Likely Benign | Likely Benign | 1.42 | Ambiguous | 0.4 | 1.56 | Ambiguous | 1.49 | Ambiguous | 0.66 | Ambiguous | 0.315 | Likely Benign | -2.06 | Neutral | 0.960 | Probably Damaging | 0.474 | Possibly Damaging | -1.04 | Pathogenic | 0.49 | Tolerated | 0.2324 | 0.2721 | 2 | 1 | 2.3 | -32.06 | ||||||||||||||||||||||||||||||
| c.1835A>G | Q612R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant Q612R is not reported in ClinVar (ClinVar ID None) and has no entries in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are SIFT and FoldX. Tools that agree on a pathogenic effect are SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, and AlphaMissense‑Default. Predictions that are uncertain or inconclusive are Rosetta, Foldetta, premPS, and AlphaMissense‑Optimized. High‑accuracy assessments show that the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts pathogenicity, while AlphaMissense‑Optimized and Foldetta are uncertain. No high‑accuracy tool provides a benign prediction. Overall, the majority of available evidence points to a pathogenic effect. This conclusion is not contradicted by ClinVar status, which has no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.275179 | Structured | 0.203988 | Uncertain | 0.822 | 0.263 | 0.000 | -10.571 | Likely Pathogenic | 0.837 | Likely Pathogenic | Ambiguous | -0.35 | Likely Benign | 0.2 | 1.56 | Ambiguous | 0.61 | Ambiguous | 0.78 | Ambiguous | 0.683 | Likely Pathogenic | -3.85 | Deleterious | 0.956 | Probably Damaging | 0.969 | Probably Damaging | -1.29 | Pathogenic | 0.10 | Tolerated | 0.1480 | 0.2196 | 1 | 1 | -1.0 | 28.06 | |||||||||||||||||||||||||||||
| c.593T>G | L198R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L198R is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that agree on a benign effect include REVEL and FATHMM, whereas a majority of tools predict a pathogenic impact: PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Predictions from FoldX, Rosetta, Foldetta, and premPS are uncertain and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as likely pathogenic, and Foldetta as uncertain. Overall, the preponderance of evidence from multiple in‑silico predictors indicates that the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status because no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | 0.444081 | Structured | 0.431715 | Uncertain | 0.572 | 0.485 | 0.125 | -15.992 | Likely Pathogenic | 0.990 | Likely Pathogenic | Likely Pathogenic | 0.74 | Ambiguous | 0.2 | 1.56 | Ambiguous | 1.15 | Ambiguous | 0.69 | Ambiguous | 0.410 | Likely Benign | -4.98 | Deleterious | 0.982 | Probably Damaging | 0.648 | Possibly Damaging | 3.34 | Benign | 0.00 | Affected | 0.1180 | 0.0719 | -3 | -2 | -8.3 | 43.03 | ||||||||||||||||||||||||||||||
| c.755C>T | P252L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P252L missense variant has no ClinVar entry and is not reported in gnomAD. Prediction tools that classify the variant as benign are premPS and FATHMM, while the majority of other in silico predictors (SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) indicate a pathogenic effect. Predictions from FoldX, Rosetta, and Foldetta are uncertain and therefore do not contribute to the overall assessment. High‑accuracy methods give the following results: AlphaMissense‑Optimized predicts pathogenicity; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates likely pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, remains uncertain. Based on the preponderance of pathogenic predictions and the lack of benign consensus, the variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.461924 | Structured | 0.211606 | Uncertain | 0.753 | 0.304 | 0.250 | -10.181 | Likely Pathogenic | 0.993 | Likely Pathogenic | Likely Pathogenic | 0.93 | Ambiguous | 0.0 | 1.56 | Ambiguous | 1.25 | Ambiguous | 0.47 | Likely Benign | 0.794 | Likely Pathogenic | -9.19 | Deleterious | 0.991 | Probably Damaging | 0.781 | Possibly Damaging | 5.81 | Benign | 0.00 | Affected | 0.2027 | 0.6475 | -3 | -3 | 5.4 | 16.04 | |||||||||||||||||||||||||||||
| c.1192C>A | P398T 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant P398T is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include polyPhen‑2 HumVar, ESM1b, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are REVEL, FoldX, PROVEAN, polyPhen‑2 HumDiv, and SIFT. The remaining tools (Rosetta, Foldetta, premPS, AlphaMissense‑Default) give uncertain results. High‑accuracy methods give a benign consensus: AlphaMissense‑Optimized is benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is benign, and Foldetta is uncertain. Overall, the majority of high‑confidence predictions lean toward a benign impact, although several other predictors indicate pathogenicity. There is no conflict with ClinVar status, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.436924 | Structured | 0.401041 | Uncertain | 0.891 | 0.525 | 0.250 | -6.670 | Likely Benign | 0.536 | Ambiguous | Likely Benign | 2.11 | Destabilizing | 0.4 | 1.57 | Ambiguous | 1.84 | Ambiguous | 0.78 | Ambiguous | 0.608 | Likely Pathogenic | -5.70 | Deleterious | 0.816 | Possibly Damaging | 0.307 | Benign | 5.51 | Benign | 0.01 | Affected | 0.1671 | 0.6607 | 0 | -1 | 0.9 | 3.99 | ||||||||||||||||||||||||||||||
| c.1336G>A | E446K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E446K is not reported in ClinVar (ClinVar status: not present) and is found in gnomAD (ID 6‑33438241‑G‑A). Prediction tools that agree on a benign effect include only FATHMM. Tools that agree on a pathogenic effect comprise REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, and the SGM‑Consensus (majority vote). Predictions that are uncertain or inconclusive are FoldX, Rosetta, Foldetta, premPS, and AlphaMissense‑Optimized. High‑accuracy methods give the following: AlphaMissense‑Optimized is uncertain; SGM‑Consensus indicates likely pathogenic; Foldetta is uncertain. Overall, the majority of evidence points to a pathogenic impact. Thus, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.271506 | Structured | 0.276479 | Uncertain | 0.940 | 0.216 | 0.000 | 6-33438241-G-A | 1 | 6.19e-7 | -14.140 | Likely Pathogenic | 0.953 | Likely Pathogenic | Ambiguous | 0.80 | Ambiguous | 0.4 | 1.57 | Ambiguous | 1.19 | Ambiguous | 0.81 | Ambiguous | 0.518 | Likely Pathogenic | -3.75 | Deleterious | 0.994 | Probably Damaging | 0.975 | Probably Damaging | 3.36 | Benign | 0.01 | Affected | 3.38 | 31 | 0.2141 | 0.6511 | 1 | 0 | -0.4 | -0.94 | ||||||||||||||||||||||||
| c.1726T>A | C576S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C576S is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign predictions from REVEL and FATHMM, while the majority of other in silico methods (premPS, PROVEAN, polyPhen‑2 HumDiv/HumVar, SIFT, ESM1b, AlphaMissense‑Default) predict pathogenicity. FoldX and Rosetta give uncertain stability changes, and Foldetta likewise reports no definitive effect. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenic, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic. Foldetta remains inconclusive. Overall, the preponderance of evidence from multiple pathogenic‑oriented tools and the high‑accuracy predictions indicates that C576S is most likely pathogenic, which is consistent with the absence of a ClinVar entry and gnomAD data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.113710 | Structured | 0.017684 | Uncertain | 0.913 | 0.245 | 0.000 | -10.474 | Likely Pathogenic | 0.991 | Likely Pathogenic | Likely Pathogenic | 0.77 | Ambiguous | 0.1 | 1.57 | Ambiguous | 1.17 | Ambiguous | 1.61 | Destabilizing | 0.414 | Likely Benign | -8.91 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.40 | Benign | 0.02 | Affected | 0.4968 | 0.1464 | 0 | -1 | -3.3 | -16.06 | |||||||||||||||||||||||||||||
| c.1727G>C | C576S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C576S is not reported in ClinVar and has no entries in gnomAD. Prediction tools that indicate a benign effect are limited to FATHMM, whereas the remaining 11 tools (SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) all predict a pathogenic or likely pathogenic outcome. High‑accuracy methods reinforce this trend: AlphaMissense‑Optimized reports pathogenic; the SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also returns likely pathogenic; Foldetta, a protein‑folding stability predictor combining FoldX‑MD and Rosetta, is inconclusive. Folding‑stability tools FoldX and Rosetta individually yield uncertain results and are treated as unavailable. Taken together, the majority of evidence points to a pathogenic effect. This conclusion is consistent with the absence of ClinVar annotation and gnomAD data, so there is no contradiction with existing database status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.113710 | Structured | 0.017684 | Uncertain | 0.913 | 0.245 | 0.000 | -10.474 | Likely Pathogenic | 0.991 | Likely Pathogenic | Likely Pathogenic | 0.77 | Ambiguous | 0.1 | 1.57 | Ambiguous | 1.17 | Ambiguous | 1.61 | Destabilizing | 0.523 | Likely Pathogenic | -8.91 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.40 | Benign | 0.02 | Affected | 0.4968 | 0.1464 | 0 | -1 | -3.3 | -16.06 | |||||||||||||||||||||||||||||
| c.1795T>A | C599S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C599S is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only SIFT, which scores the variant as benign. All other evaluated algorithms—REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and the SGM Consensus—predict a pathogenic or likely pathogenic outcome. High‑accuracy assessments show the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as Likely Pathogenic, AlphaMissense‑Optimized as Uncertain, and Foldetta (combining FoldX‑MD and Rosetta) as Uncertain. No prediction or folding stability result is missing; all available data are reported. Based on the preponderance of pathogenic predictions, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.009865 | Structured | 0.151725 | Uncertain | 0.960 | 0.151 | 0.000 | -13.336 | Likely Pathogenic | 0.954 | Likely Pathogenic | Ambiguous | 0.85 | Ambiguous | 0.0 | 1.57 | Ambiguous | 1.21 | Ambiguous | 1.27 | Destabilizing | 0.919 | Likely Pathogenic | -9.95 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.45 | Pathogenic | 0.06 | Tolerated | 0.3414 | 0.1415 | 0 | -1 | -3.3 | -16.06 | |||||||||||||||||||||||||||||
| c.1796G>C | C599S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C599S is not reported in ClinVar and has no entries in gnomAD. Prediction tools cluster into two groups: the sole benign prediction comes from SIFT, whereas the remaining nine tools—REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus—indicate pathogenicity. High‑accuracy assessments give a mixed picture: AlphaMissense‑Optimized is uncertain, SGM‑Consensus predicts likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is uncertain. FoldX and Rosetta individually report uncertain stability changes. Overall, the majority of evidence points to a deleterious effect, so the variant is most likely pathogenic, with no ClinVar annotation to contradict this assessment. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.009865 | Structured | 0.151725 | Uncertain | 0.960 | 0.151 | 0.000 | -13.336 | Likely Pathogenic | 0.954 | Likely Pathogenic | Ambiguous | 0.85 | Ambiguous | 0.0 | 1.57 | Ambiguous | 1.21 | Ambiguous | 1.27 | Destabilizing | 0.866 | Likely Pathogenic | -9.95 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.45 | Pathogenic | 0.06 | Tolerated | 0.3414 | 0.1415 | 0 | -1 | -3.3 | -16.06 | |||||||||||||||||||||||||||||
| c.2083C>G | L695V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L695V is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that agree on a pathogenic effect are premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and ESM1b. FoldX, Rosetta, and Foldetta give uncertain results and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive (2 benign vs. 2 pathogenic), and Foldetta is also unavailable. Overall, the majority of available predictions (six pathogenic vs. four benign) indicate that the variant is most likely pathogenic, and this assessment does not contradict the current ClinVar status, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.118441 | Structured | 0.373419 | Uncertain | 0.942 | 0.258 | 0.000 | -10.605 | Likely Pathogenic | 0.317 | Likely Benign | Likely Benign | 1.61 | Ambiguous | 0.0 | 1.57 | Ambiguous | 1.59 | Ambiguous | 1.19 | Destabilizing | 0.274 | Likely Benign | -2.99 | Deleterious | 0.993 | Probably Damaging | 0.694 | Possibly Damaging | 3.20 | Benign | 0.02 | Affected | 0.1158 | 0.2828 | 2 | 1 | 0.4 | -14.03 | ||||||||||||||||||||||||||||||
| c.2129A>C | K710T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K710T is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include REVEL, premPS, SIFT, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect comprise SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as Likely Pathogenic (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN), and Foldetta as uncertain (no definitive stability change). Other stability predictions (FoldX, Rosetta) are also uncertain and thus unavailable for interpretation. Overall, the majority of evidence points to a pathogenic impact. This conclusion does not contradict ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.321458 | Structured | 0.370438 | Uncertain | 0.949 | 0.368 | 0.000 | -10.454 | Likely Pathogenic | 0.759 | Likely Pathogenic | Likely Benign | 1.02 | Ambiguous | 0.0 | 1.57 | Ambiguous | 1.30 | Ambiguous | 0.21 | Likely Benign | 0.305 | Likely Benign | -5.45 | Deleterious | 0.999 | Probably Damaging | 1.000 | Probably Damaging | 3.41 | Benign | 0.06 | Tolerated | 0.1487 | 0.3000 | 0 | -1 | 3.2 | -27.07 | |||||||||||||||||||||||||||||
| c.853T>G | C285G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C285G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely converge on a pathogenic interpretation: SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default all predict deleterious effects. Tools that assess protein stability (FoldX, Rosetta, Foldetta) and AlphaMissense‑Optimized return uncertain results, providing no clear evidence for or against pathogenicity. High‑accuracy assessments further support a likely pathogenic verdict: the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is labeled Likely Pathogenic, while AlphaMissense‑Optimized and Foldetta remain inconclusive. Overall, the consensus of the majority of predictors indicates that the variant is most likely pathogenic, and this conclusion does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.144935 | Structured | 0.375400 | Uncertain | 0.946 | 0.250 | 0.000 | -9.937 | Likely Pathogenic | 0.859 | Likely Pathogenic | Ambiguous | 1.26 | Ambiguous | 0.0 | 1.57 | Ambiguous | 1.42 | Ambiguous | 1.50 | Destabilizing | 0.564 | Likely Pathogenic | -9.86 | Deleterious | 0.999 | Probably Damaging | 0.996 | Probably Damaging | 1.78 | Pathogenic | 0.04 | Affected | 0.3509 | 0.2922 | -3 | -3 | -2.9 | -46.09 | |||||||||||||||||||||||||||||
| c.1115G>C | G372A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G372A is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, AlphaMissense‑Optimized, and ESM1b. Tools that predict a pathogenic effect are SIFT and FATHMM. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, indicates a likely benign outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. FoldX and Rosetta individually also return uncertain results. Overall, the majority of evidence points to a benign impact for G372A, and this conclusion does not contradict any ClinVar status, as the variant is not yet catalogued in ClinVar. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.433034 | Structured | 0.430335 | Uncertain | 0.322 | 0.774 | 0.375 | -5.163 | Likely Benign | 0.087 | Likely Benign | Likely Benign | 1.52 | Ambiguous | 0.2 | 1.58 | Ambiguous | 1.55 | Ambiguous | -0.12 | Likely Benign | 0.465 | Likely Benign | -0.32 | Neutral | 0.000 | Benign | 0.000 | Benign | -0.74 | Pathogenic | 0.03 | Affected | 0.3956 | 0.5247 | 1 | 0 | 2.2 | 14.03 | |||||||||||||||||||||||||||||
| c.1306G>C | E436Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E436Q is not reported in ClinVar (ClinVar status: not listed) and is absent from gnomAD (no gnomAD ID). Prediction tools that indicate a benign effect include premPS and FATHMM, while the majority of tools predict a pathogenic outcome: SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. Tools with uncertain or inconclusive results are FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the consensus of pathogenic predictions outweighs benign ones, and the high‑accuracy tools reinforce a pathogenic classification. Therefore, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.239899 | Structured | 0.321046 | Uncertain | 0.934 | 0.289 | 0.000 | -12.413 | Likely Pathogenic | 0.952 | Likely Pathogenic | Ambiguous | 0.51 | Ambiguous | 0.1 | 1.58 | Ambiguous | 1.05 | Ambiguous | 0.50 | Likely Benign | 0.607 | Likely Pathogenic | -2.76 | Deleterious | 0.992 | Probably Damaging | 0.946 | Probably Damaging | 4.64 | Benign | 0.01 | Affected | 0.1237 | 0.5809 | 2 | 2 | 0.0 | -0.98 | |||||||||||||||||||||||||||||
| c.1792C>G | L598V 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L598V is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Prediction tools that classify the variant as benign include REVEL, FATHMM, and AlphaMissense‑Optimized, whereas pathogenic predictions are made by premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, indicates a likely pathogenic effect. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM Consensus as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as inconclusive. Overall, the majority of evidence points to a pathogenic impact, which contrasts with the ClinVar designation of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.007259 | Structured | 0.147872 | Uncertain | 0.953 | 0.154 | 0.000 | Uncertain | 1 | -10.002 | Likely Pathogenic | 0.578 | Likely Pathogenic | Likely Benign | 1.89 | Ambiguous | 0.1 | 1.58 | Ambiguous | 1.74 | Ambiguous | 1.01 | Destabilizing | 0.221 | Likely Benign | -2.92 | Deleterious | 0.944 | Possibly Damaging | 0.786 | Possibly Damaging | 3.21 | Benign | 0.02 | Affected | 3.37 | 35 | 0.1082 | 0.1795 | 2 | 1 | 0.4 | -14.03 | 218.4 | 29.6 | 0.0 | 0.0 | 0.8 | 0.0 | X | Potentially Benign | The iso-butyl side chain of Leu598, located on an α helix (res. Glu582-Met603), packs hydrophobically with other hydrophobic residues in the inter-helix space (e.g., Ile602, Phe594, Ile510).In the variant simulations, Val598, which has similar size and physicochemical properties to leucine, resides in the inter-helix hydrophobic space in a similar manner to Leu598 in the WT. This causes no negative effects on the protein structure. | ||||||||||||||||
| c.2090G>C | W697S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant W697S is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include REVEL, SIFT, ESM1b, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are premPS, PROVEAN, polyPhen‑2 HumDiv, and polyPhen‑2 HumVar. Four tools (FoldX, Rosetta, Foldetta, AlphaMissense‑Default) give uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as benign, and Foldetta as unavailable due to uncertainty. Overall, the majority of evidence points to a benign impact. This conclusion is not contradicted by any ClinVar annotation, as none exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.175930 | Structured | 0.400169 | Uncertain | 0.945 | 0.297 | 0.000 | -4.900 | Likely Benign | 0.449 | Ambiguous | Likely Benign | 1.90 | Ambiguous | 0.1 | 1.58 | Ambiguous | 1.74 | Ambiguous | 1.13 | Destabilizing | 0.322 | Likely Benign | -8.89 | Deleterious | 1.000 | Probably Damaging | 0.992 | Probably Damaging | 3.53 | Benign | 0.13 | Tolerated | 0.3738 | 0.1089 | -2 | -3 | 0.1 | -99.14 | ||||||||||||||||||||||||||||||
| c.639C>G | I213M 2D  AIThe SynGAP1 missense variant I213M is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are PROVEAN and FATHMM, while a larger group—REVEL, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default—consistently predict a pathogenic impact. The remaining tools (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Optimized) return uncertain or inconclusive results and are therefore treated as unavailable. High‑accuracy assessments are likewise inconclusive: AlphaMissense‑Optimized is uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is a tie, and Foldetta is uncertain. Overall, the majority of available predictions lean toward pathogenicity, and this conclusion does not contradict the lack of ClinVar annotation. Thus, the variant is most likely pathogenic based on current computational evidence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.158265 | Structured | 0.372201 | Uncertain | 0.850 | 0.295 | 0.125 | -10.777 | Likely Pathogenic | 0.906 | Likely Pathogenic | Ambiguous | 0.66 | Ambiguous | 0.5 | 1.58 | Ambiguous | 1.12 | Ambiguous | 0.85 | Ambiguous | 0.680 | Likely Pathogenic | -2.31 | Neutral | 0.995 | Probably Damaging | 0.880 | Possibly Damaging | 5.85 | Benign | 0.01 | Affected | 0.0611 | 0.2524 | 2 | 1 | -2.6 | 18.03 | ||||||||||||||||||||||||||||||
| c.706G>A | A236T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 A236T missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict a pathogenic impact are REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and ESM1b. Predictions that are uncertain or inconclusive are FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is a 2‑vs‑2 tie and thus unavailable. Foldetta, which integrates FoldX‑MD and Rosetta outputs, is also uncertain. Overall, six tools favor pathogenicity versus three favor benign, and no high‑confidence consensus tool contradicts this trend. Therefore, the variant is most likely pathogenic, and this assessment does not conflict with any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.185198 | Structured | 0.329926 | Uncertain | 0.775 | 0.330 | 0.000 | -8.319 | Likely Pathogenic | 0.240 | Likely Benign | Likely Benign | 0.73 | Ambiguous | 0.2 | 1.58 | Ambiguous | 1.16 | Ambiguous | 0.83 | Ambiguous | 0.811 | Likely Pathogenic | -3.35 | Deleterious | 0.982 | Probably Damaging | 0.747 | Possibly Damaging | 5.79 | Benign | 0.03 | Affected | 0.1342 | 0.6926 | 1 | 0 | -2.5 | 30.03 | ||||||||||||||||||||||||||||||
| c.844T>C | C282R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C282R is listed in ClinVar as Pathogenic (ClinVar ID 635755.0) and is not reported in gnomAD. Prediction tools that agree on a benign effect are limited to REVEL, which scores the variant as benign. All other evaluated algorithms predict a pathogenic outcome: FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Rosetta’s output is uncertain and is therefore not counted as evidence. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts pathogenic, the SGM‑Consensus indicates Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta) also predicts pathogenic. Based on the overwhelming agreement among these predictions, the variant is most likely pathogenic, which aligns with its ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.098513 | Structured | 0.348535 | Uncertain | 0.942 | 0.250 | 0.000 | Pathogenic | 2 | -16.378 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 3.13 | Destabilizing | 0.6 | 1.58 | Ambiguous | 2.36 | Destabilizing | 1.70 | Destabilizing | 0.466 | Likely Benign | -11.03 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | 1.63 | Pathogenic | 0.00 | Affected | 3.39 | 18 | 0.1696 | 0.1557 | -4 | -3 | -7.0 | 53.05 | 297.4 | -98.2 | -0.1 | 0.1 | 0.5 | 0.0 | X | X | X | Potentially Pathogenic | The thiol-containing side chain of Cys282, located at the beginning of an anti-parallel β sheet strand (res. Arg279-Leu286), is packed against multiple hydrophobic residues (e.g., Ile268, Leu284, Trp308, Leu327). In the variant simulations, the bulky side chain of Arg282 with its positively charged guanidinium group is not suitable for this hydrophobic niche. Consequently, the hydrophobic residues must either make room to accommodate Arg282 or it must escape the hydrophobic C2 domain core.As a result, new hydrogen bonds are formed with the backbone carbonyl groups of the surrounding β sheet residues Ala399, Leu325, and His326, which decreases the unity of the secondary structure elements. Notably, it is likely that the residue swap causes major problems during the C2 domain folding that are not visible in the variant simulations. In fact, even increased lability in the C2 domain could adversely affect the establishment of a stable SynGAP-membrane association. | ||||||||||||||
| c.985C>G | R329G 2D  3DClick to see structure in 3D Viewer AISynGAP1 R329G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign calls come from REVEL, polyPhen‑2 HumVar, and FATHMM, while pathogenic calls are made by FoldX, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, and AlphaMissense‑Default. Uncertain results are reported by Rosetta, Foldetta, premPS, and AlphaMissense‑Optimized. High‑accuracy assessments give an overall pathogenic signal: AlphaMissense‑Optimized is inconclusive, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta is inconclusive. Taken together, the majority of evidence points to a pathogenic effect, and this conclusion is not contradicted by ClinVar, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.384043 | Structured | 0.376086 | Uncertain | 0.887 | 0.479 | 0.250 | -12.426 | Likely Pathogenic | 0.927 | Likely Pathogenic | Ambiguous | 2.21 | Destabilizing | 0.3 | 1.58 | Ambiguous | 1.90 | Ambiguous | 0.92 | Ambiguous | 0.204 | Likely Benign | -4.78 | Deleterious | 0.653 | Possibly Damaging | 0.293 | Benign | 4.03 | Benign | 0.04 | Affected | 0.3147 | 0.3037 | -3 | -2 | 4.1 | -99.14 | |||||||||||||||||||||||||||||
| c.1457A>G | E486G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E486G missense change is not listed in ClinVar and has no gnomAD entry. Functional prediction tools that agree on a benign effect include REVEL, premPS, SIFT, and FATHMM. Those that predict a damaging outcome are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, and ESM1b. Predictions from FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized are inconclusive. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Overall, the majority of evidence points toward a pathogenic effect. This conclusion is consistent with the absence of a ClinVar annotation, so there is no contradiction with existing clinical data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.196879 | Structured | 0.358545 | Uncertain | 0.833 | 0.245 | 0.125 | -12.488 | Likely Pathogenic | 0.924 | Likely Pathogenic | Ambiguous | 1.09 | Ambiguous | 0.1 | 1.59 | Ambiguous | 1.34 | Ambiguous | -0.14 | Likely Benign | 0.328 | Likely Benign | -5.46 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 3.80 | Benign | 0.40 | Tolerated | 0.2918 | 0.5385 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.1684C>T | P562S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P562S is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that assess pathogenicity unanimously classify the variant as deleterious: REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict pathogenicity. The only tools with uncertain outcomes are Rosetta and premPS, which provide inconclusive results. No tool predicts a benign effect. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. Consequently, the variant is most likely pathogenic based on the collective predictions, and this assessment does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.022306 | Structured | 0.023606 | Uncertain | 0.893 | 0.200 | 0.000 | -14.293 | Likely Pathogenic | 0.992 | Likely Pathogenic | Likely Pathogenic | 2.93 | Destabilizing | 0.1 | 1.59 | Ambiguous | 2.26 | Destabilizing | 0.70 | Ambiguous | 0.708 | Likely Pathogenic | -7.96 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 0.62 | Pathogenic | 0.01 | Affected | 0.3365 | 0.2900 | 1 | -1 | 0.8 | -10.04 | |||||||||||||||||||||||||||||
| c.1880C>T | A627V 2D  AIThe SynGAP1 missense variant A627V is not reported in ClinVar (ClinVar status: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that assess pathogenicity unanimously classify the variant as deleterious: REVEL, FoldX, Foldetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict pathogenicity. No tool in the dataset predicts a benign effect. Two tools return uncertain results: Rosetta and premPS. High‑accuracy assessments reinforce the pathogenic prediction: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; and Foldetta is pathogenic. Based on the collective evidence, the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.100716 | Structured | 0.037862 | Uncertain | 0.970 | 0.210 | 0.000 | -12.150 | Likely Pathogenic | 0.984 | Likely Pathogenic | Likely Pathogenic | 2.64 | Destabilizing | 1.5 | 1.59 | Ambiguous | 2.12 | Destabilizing | 0.74 | Ambiguous | 0.549 | Likely Pathogenic | -3.98 | Deleterious | 0.999 | Probably Damaging | 0.900 | Possibly Damaging | 2.47 | Pathogenic | 0.00 | Affected | 0.0856 | 0.4551 | 0 | 0 | 2.4 | 28.05 | |||||||||||||||||||||||||||||
| c.1102C>T | P368S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P368S is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are PROVEAN and FATHMM. The remaining methods (FoldX, Rosetta, Foldetta, premPS) yield uncertain results and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive (2 benign vs 2 pathogenic), and Foldetta is uncertain, so these do not alter the overall interpretation. Overall, the majority of evidence points to a benign impact. Thus, the variant is most likely benign, and this conclusion does not contradict the current ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.363090 | Structured | 0.439989 | Uncertain | 0.580 | 0.677 | 0.250 | -4.790 | Likely Benign | 0.247 | Likely Benign | Likely Benign | 1.68 | Ambiguous | 0.4 | 1.60 | Ambiguous | 1.64 | Ambiguous | 0.52 | Ambiguous | 0.090 | Likely Benign | -5.12 | Deleterious | 0.384 | Benign | 0.113 | Benign | 1.80 | Pathogenic | 0.10 | Tolerated | 0.3700 | 0.5635 | 1 | -1 | 0.8 | -10.04 | ||||||||||||||||||||||||||||||
| c.1453C>G | R485G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R485G is not reported in ClinVar (ClinVar status: not listed) but is present in gnomAD (gnomAD ID: 6‑33438485‑C‑G). Prediction tools that agree on a pathogenic effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized; no tools predict a benign outcome. Uncertain or inconclusive predictions come from FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta as Uncertain. Overall, the evidence strongly favors a pathogenic classification, and this conclusion does not contradict the ClinVar status, which simply lacks an entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.188120 | Structured | 0.377409 | Uncertain | 0.805 | 0.246 | 0.125 | 6-33438485-C-G | 1 | 6.20e-7 | -15.777 | Likely Pathogenic | 0.991 | Likely Pathogenic | Likely Pathogenic | 0.84 | Ambiguous | 0.1 | 1.60 | Ambiguous | 1.22 | Ambiguous | 0.98 | Ambiguous | 0.631 | Likely Pathogenic | -6.96 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 1.92 | Pathogenic | 0.00 | Affected | 3.37 | 35 | 0.3140 | 0.2678 | -2 | -3 | 4.1 | -99.14 | ||||||||||||||||||||||||
| c.1850A>G | E617G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E617G is not reported in ClinVar and is absent from gnomAD. Prediction tools cluster into two groups: benign predictions come from premPS, SIFT, and AlphaMissense‑Optimized, whereas pathogenic predictions are made by SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), FATHMM, and AlphaMissense‑Default. Uncertain results are reported by FoldX, Rosetta, Foldetta, and ESM1b. High‑accuracy methods give a mixed picture: AlphaMissense‑Optimized predicts benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts pathogenic, and Foldetta is uncertain. Overall, the majority of evidence points to a pathogenic effect. This conclusion is not contradicted by ClinVar, which contains no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.111485 | Structured | 0.155123 | Uncertain | 0.877 | 0.240 | 0.000 | -7.984 | In-Between | 0.777 | Likely Pathogenic | Likely Benign | 0.59 | Ambiguous | 0.2 | 1.60 | Ambiguous | 1.10 | Ambiguous | 0.22 | Likely Benign | 0.701 | Likely Pathogenic | -4.99 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | -1.41 | Pathogenic | 0.18 | Tolerated | 0.2344 | 0.5442 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.1994A>G | Y665C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant Y665C is listed in ClinVar with no assertion (status: None) and is present in gnomAD (ID 6‑33441253‑A‑G). Prediction tools that agree on a benign effect include REVEL, SIFT, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict a pathogenic outcome are premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and ESM1b. FoldX, Rosetta, and Foldetta give uncertain results and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive (2 benign vs 2 pathogenic), and Foldetta is also unavailable. Overall, the evidence is mixed, but the majority of high‑confidence tools lean toward a benign interpretation. Thus, the variant is most likely benign based on current predictions, and this does not contradict the ClinVar status, which remains unclassified. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.098513 | Structured | 0.086641 | Uncertain | 0.922 | 0.361 | 0.000 | 6-33441253-A-G | 1 | 6.20e-7 | -9.007 | Likely Pathogenic | 0.261 | Likely Benign | Likely Benign | 1.05 | Ambiguous | 0.1 | 1.60 | Ambiguous | 1.33 | Ambiguous | 1.12 | Destabilizing | 0.210 | Likely Benign | -3.22 | Deleterious | 1.000 | Probably Damaging | 0.981 | Probably Damaging | 3.45 | Benign | 0.14 | Tolerated | 3.38 | 28 | 0.2562 | 0.2019 | -2 | 0 | 3.8 | -60.04 | |||||||||||||||||||||||||
| c.891G>C | K297N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K297N is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only REVEL, whereas the majority of tools (premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus) predict a pathogenic impact. FoldX, Rosetta, and Foldetta provide uncertain or inconclusive stability results and are therefore not considered evidence for or against pathogenicity. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. Based on the preponderance of pathogenic predictions and the lack of benign consensus, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.422041 | Structured | 0.272593 | Uncertain | 0.880 | 0.285 | 0.375 | -11.328 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 1.08 | Ambiguous | 0.1 | 1.60 | Ambiguous | 1.34 | Ambiguous | 1.15 | Destabilizing | 0.253 | Likely Benign | -4.31 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 1.61 | Pathogenic | 0.01 | Affected | 0.3797 | 0.2265 | 1 | 0 | 0.4 | -14.07 | |||||||||||||||||||||||||||||
| c.891G>T | K297N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K297N is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only REVEL, whereas the majority of tools (premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus) predict a pathogenic impact. FoldX, Rosetta, and Foldetta provide uncertain or inconclusive stability results and are therefore not considered evidence for or against pathogenicity. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. Based on the preponderance of pathogenic predictions and the lack of benign consensus, K297N is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.422041 | Structured | 0.272593 | Uncertain | 0.880 | 0.285 | 0.375 | -11.328 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 1.08 | Ambiguous | 0.1 | 1.60 | Ambiguous | 1.34 | Ambiguous | 1.15 | Destabilizing | 0.253 | Likely Benign | -4.31 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 1.61 | Pathogenic | 0.01 | Affected | 0.3797 | 0.2265 | 1 | 0 | 0.4 | -14.07 | |||||||||||||||||||||||||||||
| c.1102C>A | P368T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P368T missense variant is not listed in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and FATHMM. FoldX, Rosetta, and Foldetta report uncertain or inconclusive stability changes and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is a tie (2 benign, 2 pathogenic) and thus inconclusive, and Foldetta remains uncertain. Overall, the predictions are evenly split between benign and pathogenic, providing no definitive classification. The variant’s status does not contradict ClinVar, which has no entry for it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.363090 | Structured | 0.439989 | Uncertain | 0.580 | 0.677 | 0.250 | -5.308 | Likely Benign | 0.284 | Likely Benign | Likely Benign | 1.95 | Ambiguous | 0.6 | 1.61 | Ambiguous | 1.78 | Ambiguous | 0.45 | Likely Benign | 0.188 | Likely Benign | -5.43 | Deleterious | 0.941 | Possibly Damaging | 0.527 | Possibly Damaging | 1.72 | Pathogenic | 0.01 | Affected | 0.1983 | 0.6155 | 0 | -1 | 0.9 | 3.99 | ||||||||||||||||||||||||||||||
| c.1530T>G | I510M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 I510M missense variant is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign impact include PROVEAN, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict pathogenicity are REVEL, polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM. The remaining tools (FoldX, Rosetta, Foldetta, premPS, ESM1b) returned uncertain results and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus—derived from a majority vote of AlphaMissense‑Default (benign), ESM1b (uncertain), FATHMM (pathogenic), and PROVEAN (benign)—also favors benign. Foldetta, a protein‑folding stability method, yielded an uncertain outcome. Taken together, the consensus of the most reliable predictors indicates a benign effect. This conclusion does not contradict the ClinVar status, which contains no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.025762 | Structured | 0.250630 | Uncertain | 0.945 | 0.273 | 0.000 | -7.988 | In-Between | 0.235 | Likely Benign | Likely Benign | 0.56 | Ambiguous | 0.3 | 1.61 | Ambiguous | 1.09 | Ambiguous | 0.55 | Ambiguous | 0.532 | Likely Pathogenic | -0.97 | Neutral | 0.999 | Probably Damaging | 0.996 | Probably Damaging | -1.42 | Pathogenic | 0.02 | Affected | 0.0674 | 0.1789 | 2 | 1 | -2.6 | 18.03 | ||||||||||||||||||||||||||||||
| c.1952A>G | E651G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E651G missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, SIFT, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and AlphaMissense‑Default. Predictions that are uncertain or unavailable are FoldX, Rosetta, ESM1b, and Foldetta. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as unavailable. Overall, the majority of tools (five benign vs. four pathogenic) lean toward a benign interpretation, and this does not contradict the lack of ClinVar annotation. Thus, based on the available predictions, the variant is most likely benign. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.088832 | Structured | 0.365409 | Uncertain | 0.955 | 0.340 | 0.000 | -7.596 | In-Between | 0.591 | Likely Pathogenic | Likely Benign | 0.66 | Ambiguous | 0.1 | 1.61 | Ambiguous | 1.14 | Ambiguous | 0.32 | Likely Benign | 0.444 | Likely Benign | -4.78 | Deleterious | 0.999 | Probably Damaging | 0.935 | Probably Damaging | 3.31 | Benign | 0.06 | Tolerated | 0.2908 | 0.4488 | 0 | -2 | 3.1 | -72.06 | ||||||||||||||||||||||||||||||
| c.649G>C | E217Q 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant E217Q is reported in gnomAD (ID 6‑33435291‑G‑C) but has no ClinVar entry. Functional prediction tools cluster into two groups: benign predictions come from REVEL, FoldX, PROVEAN, SIFT, ESM1b, and FATHMM, while pathogenic predictions arise from polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. With six benign versus three pathogenic calls and a benign consensus from the high‑accuracy panel, the variant is most likely benign. This conclusion does not contradict ClinVar, which contains no classification for this change. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | PH | 0.278302 | Structured | 0.404912 | Uncertain | 0.823 | 0.284 | 0.000 | 6-33435291-G-C | 1 | 6.20e-7 | -6.810 | Likely Benign | 0.949 | Likely Pathogenic | Ambiguous | 0.25 | Likely Benign | 0.2 | 1.61 | Ambiguous | 0.93 | Ambiguous | 0.67 | Ambiguous | 0.459 | Likely Benign | -1.89 | Neutral | 0.900 | Possibly Damaging | 0.461 | Possibly Damaging | 5.83 | Benign | 0.14 | Tolerated | 3.41 | 13 | 0.1474 | 0.8473 | 2 | 2 | 0.0 | -0.98 | ||||||||||||||||||||||||
| c.732G>C | E244D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E244D missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include FoldX and FATHMM, whereas a majority of tools predict a pathogenic impact: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, AlphaMissense‑Default, and AlphaMissense‑Optimized. Uncertain results are reported by Rosetta, Foldetta, premPS, and ESM1b. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic, and Foldetta remains uncertain. Overall, the balance of evidence favors a pathogenic classification for E244D, and this conclusion does not contradict the current ClinVar status, which has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.450668 | Structured | 0.329406 | Uncertain | 0.778 | 0.360 | 0.000 | -7.839 | In-Between | 0.979 | Likely Pathogenic | Likely Pathogenic | 0.46 | Likely Benign | 0.1 | 1.61 | Ambiguous | 1.04 | Ambiguous | 0.93 | Ambiguous | 0.730 | Likely Pathogenic | -2.53 | Deleterious | 0.976 | Probably Damaging | 0.675 | Possibly Damaging | 5.78 | Benign | 0.03 | Affected | 0.1740 | 0.3783 | 3 | 2 | 0.0 | -14.03 | ||||||||||||||||||||||||||||||
| c.732G>T | E244D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E244D missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include FoldX and FATHMM, whereas a majority of tools predict a pathogenic impact: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, AlphaMissense‑Default, and AlphaMissense‑Optimized. Uncertain results come from Rosetta, Foldetta, premPS, and ESM1b. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic, and Foldetta remains uncertain. Overall, the balance of evidence points to a likely pathogenic effect for E244D, and this conclusion does not contradict the current ClinVar status, which has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.450668 | Structured | 0.329406 | Uncertain | 0.778 | 0.360 | 0.000 | -7.839 | In-Between | 0.979 | Likely Pathogenic | Likely Pathogenic | 0.46 | Likely Benign | 0.1 | 1.61 | Ambiguous | 1.04 | Ambiguous | 0.93 | Ambiguous | 0.730 | Likely Pathogenic | -2.53 | Deleterious | 0.976 | Probably Damaging | 0.675 | Possibly Damaging | 5.78 | Benign | 0.03 | Affected | 0.1740 | 0.3783 | 3 | 2 | 0.0 | -14.03 | ||||||||||||||||||||||||||||||
| c.1072T>C | F358L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 F358L missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, SIFT, and FATHMM, whereas those that agree on a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, and AlphaMissense‑Optimized. Uncertain or inconclusive results are reported for Rosetta, Foldetta, premPS, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also predicts pathogenic, while Foldetta’s stability prediction is unavailable due to inconclusiveness. Overall, the majority of available evidence points to a pathogenic impact. This conclusion does not contradict ClinVar status, as no ClinVar assertion exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.222385 | Structured | 0.407113 | Uncertain | 0.912 | 0.441 | 0.250 | -7.865 | In-Between | 0.964 | Likely Pathogenic | Likely Pathogenic | 0.18 | Likely Benign | 0.1 | 1.62 | Ambiguous | 0.90 | Ambiguous | 0.97 | Ambiguous | 0.290 | Likely Benign | -4.21 | Deleterious | 0.982 | Probably Damaging | 0.952 | Probably Damaging | 4.12 | Benign | 0.22 | Tolerated | 0.2555 | 0.3602 | 2 | 0 | 1.0 | -34.02 | ||||||||||||||||||||||||||||||
| c.1074C>A | F358L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 F358L missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, SIFT, and FATHMM, whereas those that agree on a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, and AlphaMissense‑Optimized. Uncertain or inconclusive results are reported for Rosetta, Foldetta, premPS, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also predicts pathogenic, while Foldetta’s stability prediction is unavailable due to inconclusiveness. Overall, the majority of available evidence points to a pathogenic impact. This conclusion does not contradict ClinVar status, as no ClinVar assertion exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.222385 | Structured | 0.407113 | Uncertain | 0.912 | 0.441 | 0.250 | -7.865 | In-Between | 0.964 | Likely Pathogenic | Likely Pathogenic | 0.18 | Likely Benign | 0.1 | 1.62 | Ambiguous | 0.90 | Ambiguous | 0.97 | Ambiguous | 0.215 | Likely Benign | -4.21 | Deleterious | 0.982 | Probably Damaging | 0.952 | Probably Damaging | 4.12 | Benign | 0.22 | Tolerated | 0.2555 | 0.3602 | 2 | 0 | 1.0 | -34.02 | ||||||||||||||||||||||||||||||
| c.1074C>G | F358L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 F358L missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, SIFT, and FATHMM, whereas those that agree on a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, and AlphaMissense‑Optimized. Uncertain or inconclusive results come from Rosetta, Foldetta, premPS, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also predicts pathogenic, while Foldetta’s stability prediction is unavailable. Overall, the majority of reliable tools predict a pathogenic impact for F358L. This conclusion does not contradict ClinVar status, which has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.222385 | Structured | 0.407113 | Uncertain | 0.912 | 0.441 | 0.250 | -7.865 | In-Between | 0.964 | Likely Pathogenic | Likely Pathogenic | 0.18 | Likely Benign | 0.1 | 1.62 | Ambiguous | 0.90 | Ambiguous | 0.97 | Ambiguous | 0.215 | Likely Benign | -4.21 | Deleterious | 0.982 | Probably Damaging | 0.952 | Probably Damaging | 4.12 | Benign | 0.22 | Tolerated | 0.2555 | 0.3602 | 2 | 0 | 1.0 | -34.02 | ||||||||||||||||||||||||||||||
| c.1109G>C | G370A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G370A is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Benign). Pathogenic predictions come from FoldX, Foldetta, and FATHMM, while Rosetta remains uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus also indicating a likely benign outcome, and Foldetta suggesting a pathogenic impact via combined FoldX‑MD and Rosetta stability analysis. Overall, the majority of evidence points to a benign effect, with only a minority of tools predicting pathogenicity. There is no ClinVar entry to contradict this assessment, so the variant is most likely benign based on current computational predictions. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.461924 | Structured | 0.434325 | Uncertain | 0.359 | 0.720 | 0.500 | -3.334 | Likely Benign | 0.080 | Likely Benign | Likely Benign | 2.44 | Destabilizing | 1.3 | 1.62 | Ambiguous | 2.03 | Destabilizing | -0.14 | Likely Benign | 0.304 | Likely Benign | 0.54 | Neutral | 0.000 | Benign | 0.000 | Benign | 1.33 | Pathogenic | 0.79 | Tolerated | 0.3883 | 0.5247 | 1 | 0 | 2.2 | 14.03 | |||||||||||||||||||||||||||||
| c.1307A>T | E436V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E436V variant is not reported in ClinVar and is absent from gnomAD. Prediction tools that indicate a benign effect include premPS and FATHMM, while the remaining evaluated algorithms (REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus) all predict a pathogenic or likely pathogenic outcome. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Based on the preponderance of pathogenic predictions and the high‑accuracy tools’ results, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.239899 | Structured | 0.321046 | Uncertain | 0.934 | 0.289 | 0.000 | -13.364 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 1.45 | Ambiguous | 0.0 | 1.62 | Ambiguous | 1.54 | Ambiguous | 0.30 | Likely Benign | 0.875 | Likely Pathogenic | -6.63 | Deleterious | 0.995 | Probably Damaging | 0.967 | Probably Damaging | 4.64 | Benign | 0.03 | Affected | 0.0727 | 0.5952 | -2 | -2 | 7.7 | -29.98 | |||||||||||||||||||||||||||||
| c.1463C>T | T488M 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant T488M is listed in ClinVar with an uncertain significance (ClinVar ID 2824521.0) and is present in gnomAD (ID 6‑33438495‑C‑T). Prediction tools that indicate a benign effect include premPS and FATHMM, whereas the majority of algorithms predict a pathogenic outcome: REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta) as inconclusive. No other tools provide definitive evidence. Based on the preponderance of pathogenic predictions, the variant is most likely pathogenic, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.206376 | Structured | 0.332663 | Uncertain | 0.928 | 0.233 | 0.125 | Uncertain | 1 | 6-33438495-C-T | 2 | 1.24e-6 | -12.459 | Likely Pathogenic | 0.973 | Likely Pathogenic | Likely Pathogenic | 0.66 | Ambiguous | 0.3 | 1.62 | Ambiguous | 1.14 | Ambiguous | 0.46 | Likely Benign | 0.746 | Likely Pathogenic | -5.70 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.21 | Benign | 0.00 | Affected | 3.37 | 35 | 0.1027 | 0.4857 | -1 | -1 | 2.6 | 30.09 | ||||||||||||||||||||||
| c.1565A>G | E522G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E522G missense variant is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Consensus from multiple in‑silico predictors indicates a pathogenic effect: SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the change as deleterious. No tool reports a benign outcome. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts a damaging effect, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is labeled Likely Pathogenic, while Foldetta’s stability analysis is inconclusive. FoldX, Rosetta, and premPS are uncertain and are treated as unavailable evidence. Overall, the computational evidence overwhelmingly points to a pathogenic impact, and this conclusion does not conflict with the absence of a ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.092881 | Structured | 0.046216 | Uncertain | 0.823 | 0.376 | 0.000 | -10.053 | Likely Pathogenic | 0.986 | Likely Pathogenic | Likely Pathogenic | 0.72 | Ambiguous | 0.0 | 1.62 | Ambiguous | 1.17 | Ambiguous | 0.58 | Ambiguous | 0.823 | Likely Pathogenic | -6.59 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | -1.34 | Pathogenic | 0.01 | Affected | 0.2672 | 0.3654 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.1961A>G | E654G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E654G missense variant is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on benign impact are premPS and FATHMM, while the majority (REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, and the SGM‑Consensus) predict pathogenicity. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Overall, the preponderance of evidence points to a pathogenic effect for E654G. This conclusion is consistent with the lack of ClinVar annotation, so there is no contradiction with existing clinical database status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.026892 | Structured | 0.303029 | Uncertain | 0.957 | 0.311 | 0.000 | -12.487 | Likely Pathogenic | 0.898 | Likely Pathogenic | Ambiguous | 1.29 | Ambiguous | 0.2 | 1.62 | Ambiguous | 1.46 | Ambiguous | 0.34 | Likely Benign | 0.547 | Likely Pathogenic | -6.73 | Deleterious | 0.999 | Probably Damaging | 0.935 | Probably Damaging | 3.26 | Benign | 0.01 | Affected | 0.2818 | 0.3109 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.716G>A | R239K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 R239K missense variant has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect are polyPhen‑2 HumVar and FATHMM, while the majority of other in silico predictors (SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default) indicate a pathogenic impact. High‑accuracy assessments show that AlphaMissense‑Optimized is uncertain, SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is uncertain. Because uncertain or unavailable results are not taken as evidence for or against pathogenicity, the overall evidence still leans toward a deleterious effect. Consequently, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.170161 | Structured | 0.336504 | Uncertain | 0.854 | 0.319 | 0.000 | -12.492 | Likely Pathogenic | 0.897 | Likely Pathogenic | Ambiguous | 1.93 | Ambiguous | 0.2 | 1.62 | Ambiguous | 1.78 | Ambiguous | 1.41 | Destabilizing | 0.719 | Likely Pathogenic | -2.52 | Deleterious | 0.882 | Possibly Damaging | 0.428 | Benign | 5.78 | Benign | 0.03 | Affected | 0.5222 | 0.4000 | Weaken | 3 | 2 | 0.6 | -28.01 | ||||||||||||||||||||||||||||
| c.726G>C | W242C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 W242C missense variant is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Prediction tools that agree on a pathogenic effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that are uncertain or inconclusive are FoldX, Rosetta, Foldetta, and premPS, and no tools predict a benign outcome. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta as uncertain. Based on the collective predictions, the variant is most likely pathogenic. This assessment does not contradict ClinVar status, as ClinVar has not yet classified the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.328603 | Structured | 0.352582 | Uncertain | 0.847 | 0.341 | 0.000 | -13.117 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 1.77 | Ambiguous | 0.4 | 1.62 | Ambiguous | 1.70 | Ambiguous | 0.63 | Ambiguous | 0.889 | Likely Pathogenic | -11.80 | Deleterious | 0.999 | Probably Damaging | 0.887 | Possibly Damaging | 1.52 | Pathogenic | 0.00 | Affected | 0.3626 | 0.1617 | -8 | -2 | 3.4 | -83.07 | |||||||||||||||||||||||||||||
| c.726G>T | W242C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant W242C is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Prediction tools that agree on a pathogenic effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that are uncertain or inconclusive are FoldX, Rosetta, Foldetta, and premPS, and no tools predict a benign outcome. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta as uncertain. Based on the collective predictions, the variant is most likely pathogenic. This assessment does not contradict ClinVar status, as ClinVar has not yet classified the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.328603 | Structured | 0.352582 | Uncertain | 0.847 | 0.341 | 0.000 | -13.117 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 1.77 | Ambiguous | 0.4 | 1.62 | Ambiguous | 1.70 | Ambiguous | 0.63 | Ambiguous | 0.889 | Likely Pathogenic | -11.80 | Deleterious | 0.999 | Probably Damaging | 0.887 | Possibly Damaging | 1.52 | Pathogenic | 0.00 | Affected | 0.3626 | 0.1617 | -8 | -2 | 3.4 | -83.07 | |||||||||||||||||||||||||||||
| c.790C>G | L264V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L264V is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect are REVEL and AlphaMissense‑Optimized; all other evaluated algorithms (SGM‑Consensus, FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default) predict a pathogenic or likely pathogenic outcome, while Rosetta remains uncertain. High‑accuracy assessments further support a pathogenic interpretation: AlphaMissense‑Optimized predicts benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) classifies the variant as Pathogenic. Overall, the preponderance of evidence points to a pathogenic effect for L264V, and this conclusion does not contradict the absence of a ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.185198 | Structured | 0.323473 | Uncertain | 0.939 | 0.264 | 0.000 | -10.621 | Likely Pathogenic | 0.630 | Likely Pathogenic | Likely Benign | 2.55 | Destabilizing | 0.1 | 1.62 | Ambiguous | 2.09 | Destabilizing | 1.24 | Destabilizing | 0.444 | Likely Benign | -2.76 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | 0.73 | Pathogenic | 0.01 | Affected | 0.1280 | 0.2289 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.1100T>A | L367Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L367Q is not reported in ClinVar and is absent from gnomAD. Prediction tools that indicate a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, AlphaMissense‑Optimized, and ESM1b. Tools that predict a pathogenic effect are SIFT and FATHMM. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as “Likely Benign.” High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus also as benign, while Foldetta (combining FoldX‑MD and Rosetta outputs) yields an uncertain result. Overall, the majority of evidence points to a benign impact. This conclusion is consistent with the lack of ClinVar annotation; there is no contradiction with existing clinical data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.370445 | Structured | 0.441805 | Uncertain | 0.790 | 0.657 | 0.250 | -3.432 | Likely Benign | 0.150 | Likely Benign | Likely Benign | 1.09 | Ambiguous | 0.3 | 1.63 | Ambiguous | 1.36 | Ambiguous | 0.31 | Likely Benign | 0.061 | Likely Benign | 0.38 | Neutral | 0.002 | Benign | 0.002 | Benign | 1.65 | Pathogenic | 0.02 | Affected | 0.1615 | 0.0973 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.1337A>G | E446G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E446G missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only FATHMM. The majority of tools predict a pathogenic impact: SGM‑Consensus, REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and Foldetta. Predictions that are uncertain or inconclusive are AlphaMissense‑Optimized, Rosetta, and premPS. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is uncertain, SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is Pathogenic. Overall, the preponderance of evidence indicates that the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.271506 | Structured | 0.276479 | Uncertain | 0.940 | 0.216 | 0.000 | -11.457 | Likely Pathogenic | 0.866 | Likely Pathogenic | Ambiguous | 2.62 | Destabilizing | 0.7 | 1.63 | Ambiguous | 2.13 | Destabilizing | 0.83 | Ambiguous | 0.510 | Likely Pathogenic | -6.42 | Deleterious | 1.000 | Probably Damaging | 0.997 | Probably Damaging | 3.24 | Benign | 0.00 | Affected | 0.2665 | 0.5202 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.1399G>A | D467N 2D  3DClick to see structure in 3D Viewer AISynGAP1 D467N is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a predominance of pathogenic calls: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus (majority vote of the four high‑accuracy predictors) all predict a deleterious effect. Benign predictions come from FoldX, premPS, and SIFT. Rosetta, Foldetta, and AlphaMissense‑Optimized are inconclusive. High‑accuracy methods specifically give an uncertain result for AlphaMissense‑Optimized, a pathogenic verdict for the SGM‑Consensus, and an uncertain outcome for Foldetta. Overall, the balance of evidence favors a pathogenic impact for D467N, and this assessment is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.268042 | Structured | 0.329932 | Uncertain | 0.940 | 0.246 | 0.000 | -11.881 | Likely Pathogenic | 0.913 | Likely Pathogenic | Ambiguous | 0.43 | Likely Benign | 0.1 | 1.63 | Ambiguous | 1.03 | Ambiguous | 0.38 | Likely Benign | 0.673 | Likely Pathogenic | -4.82 | Deleterious | 0.987 | Probably Damaging | 0.990 | Probably Damaging | -1.22 | Pathogenic | 0.06 | Tolerated | 0.0936 | 0.4879 | 2 | 1 | 0.0 | -0.98 | |||||||||||||||||||||||||||||
| c.1478C>G | A493G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A493G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus all predict pathogenicity, while only AlphaMissense‑Optimized predicts a benign outcome. Predictions from FoldX, Rosetta, and Foldetta are uncertain and therefore not considered evidence. High‑accuracy assessments show AlphaMissense‑Optimized as benign, whereas the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) remains pathogenic; Foldetta likewise yields an uncertain result. Overall, the preponderance of evidence points to a pathogenic effect for A493G, and this conclusion does not contradict the absence of a ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.102787 | Structured | 0.340081 | Uncertain | 0.966 | 0.182 | 0.000 | -11.379 | Likely Pathogenic | 0.571 | Likely Pathogenic | Likely Benign | 1.85 | Ambiguous | 0.0 | 1.63 | Ambiguous | 1.74 | Ambiguous | 1.40 | Destabilizing | 0.764 | Likely Pathogenic | -3.54 | Deleterious | 0.999 | Probably Damaging | 0.995 | Probably Damaging | -1.40 | Pathogenic | 0.02 | Affected | 0.1673 | 0.2228 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.1525G>T | A509S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A509S is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include PROVEAN, polyPhen‑2 HumDiv, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are REVEL, polyPhen‑2 HumVar, SIFT, ESM1b, and FATHMM. Four tools give uncertain or inconclusive results (FoldX, Rosetta, Foldetta, premPS). High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is a 2‑vs‑2 tie and thus inconclusive, and Foldetta is also inconclusive. Overall, the majority of standard predictors lean toward pathogenicity, while the most reliable single‑tool prediction (AlphaMissense‑Optimized) suggests benign. Given the balance of evidence, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.025762 | Structured | 0.250110 | Uncertain | 0.923 | 0.256 | 0.000 | -9.997 | Likely Pathogenic | 0.171 | Likely Benign | Likely Benign | 0.83 | Ambiguous | 0.1 | 1.63 | Ambiguous | 1.23 | Ambiguous | 0.59 | Ambiguous | 0.621 | Likely Pathogenic | -2.18 | Neutral | 0.119 | Benign | 0.468 | Possibly Damaging | -1.35 | Pathogenic | 0.01 | Affected | 0.2410 | 0.4384 | 1 | 1 | -2.6 | 16.00 | ||||||||||||||||||||||||||||||
| c.1537T>C | F513L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F513L is not reported in ClinVar and is absent from gnomAD. Prediction tools cluster into two groups: benign predictions are provided only by SIFT, whereas the remaining tools—REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Pathogenic)—all predict a deleterious effect. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta, which integrates FoldX‑MD and Rosetta outputs, is uncertain. No evidence from FoldX or Rosetta is considered decisive. Overall, the preponderance of predictions indicates that the variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.102787 | Structured | 0.250651 | Uncertain | 0.949 | 0.269 | 0.000 | -10.370 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 1.36 | Ambiguous | 0.2 | 1.63 | Ambiguous | 1.50 | Ambiguous | 1.14 | Destabilizing | 0.674 | Likely Pathogenic | -5.63 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | -1.07 | Pathogenic | 0.19 | Tolerated | 0.1645 | 0.2447 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1539C>A | F513L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F513L is not reported in ClinVar and is absent from gnomAD. Prediction tools cluster into two groups: benign predictions are provided only by SIFT, whereas the remaining tools—REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Pathogenic)—all predict a deleterious effect. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta, which integrates FoldX‑MD and Rosetta outputs, is uncertain. No evidence from FoldX or Rosetta is considered decisive. Overall, the preponderance of predictions indicates that the variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.102787 | Structured | 0.250651 | Uncertain | 0.949 | 0.269 | 0.000 | -10.370 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 1.36 | Ambiguous | 0.2 | 1.63 | Ambiguous | 1.50 | Ambiguous | 1.14 | Destabilizing | 0.537 | Likely Pathogenic | -5.63 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | -1.07 | Pathogenic | 0.19 | Tolerated | 0.1645 | 0.2447 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1539C>G | F513L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 F513L missense variant has no ClinVar entry and is absent from gnomAD. Prediction tools cluster into two groups: benign predictions are provided only by SIFT, whereas the remaining tools—REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus—consistently predict pathogenicity. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized scores the variant as pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) labels it likely pathogenic, and the Foldetta stability analysis is inconclusive and therefore not considered evidence. FoldX and Rosetta predictions are uncertain and treated as unavailable. Overall, the preponderance of evidence indicates that F513L is most likely pathogenic, and this conclusion does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.102787 | Structured | 0.250651 | Uncertain | 0.949 | 0.269 | 0.000 | -10.370 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 1.36 | Ambiguous | 0.2 | 1.63 | Ambiguous | 1.50 | Ambiguous | 1.14 | Destabilizing | 0.537 | Likely Pathogenic | -5.63 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | -1.07 | Pathogenic | 0.19 | Tolerated | 0.1645 | 0.2447 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1709C>T | A570V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 A570V missense variant is catalogued in gnomAD (ID 6‑33440761‑C‑T) but has no ClinVar entry. Functional prediction tools cluster into two groups: benign predictions come from premPS and SIFT, while pathogenic predictions are made by REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, and AlphaMissense‑Default. Four tools report uncertainty: FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Overall, the majority of evidence points to a pathogenic effect. This conclusion is not contradicted by ClinVar status, which currently contains no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.046336 | Structured | 0.054494 | Uncertain | 0.932 | 0.263 | 0.000 | 6-33440761-C-T | 1 | 6.22e-7 | -13.083 | Likely Pathogenic | 0.882 | Likely Pathogenic | Ambiguous | 0.88 | Ambiguous | 0.3 | 1.63 | Ambiguous | 1.26 | Ambiguous | 0.46 | Likely Benign | 0.669 | Likely Pathogenic | -3.75 | Deleterious | 0.999 | Probably Damaging | 0.988 | Probably Damaging | -1.30 | Pathogenic | 0.06 | Tolerated | 3.37 | 35 | 0.1050 | 0.3173 | 0 | 0 | 2.4 | 28.05 | ||||||||||||||||||||||||
| c.1801G>T | A601S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 A601S missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that agree on a pathogenic effect are REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and ESM1b. The remaining tools—FoldX, Rosetta, Foldetta, and premPS—return uncertain or inconclusive results and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized predicts benign, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is a 2‑vs‑2 tie and thus inconclusive. Foldetta, which integrates FoldX‑MD and Rosetta outputs, also yields an uncertain result. Overall, more tools (six) predict pathogenicity than benign (three), and no ClinVar evidence contradicts this assessment. Therefore, the variant is most likely pathogenic based on the available predictions. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.008895 | Structured | 0.174517 | Uncertain | 0.955 | 0.156 | 0.000 | -11.248 | Likely Pathogenic | 0.136 | Likely Benign | Likely Benign | 0.79 | Ambiguous | 0.1 | 1.63 | Ambiguous | 1.21 | Ambiguous | 0.79 | Ambiguous | 0.541 | Likely Pathogenic | -2.99 | Deleterious | 0.983 | Probably Damaging | 0.993 | Probably Damaging | 2.56 | Benign | 0.01 | Affected | 0.2732 | 0.4730 | 1 | 1 | -2.6 | 16.00 | ||||||||||||||||||||||||||||||
| c.2018T>C | L673P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L673P is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. All other evaluated tools—SGM‑Consensus, FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default—predict a pathogenic impact. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized indicates benign, but the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) resolves as Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts Pathogenic. Taken together, the overwhelming majority of predictions, including the high‑accuracy tools, classify the variant as pathogenic. This conclusion is not contradicted by ClinVar, which contains no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.060549 | Structured | 0.104692 | Uncertain | 0.545 | 0.369 | 0.000 | -12.160 | Likely Pathogenic | 0.622 | Likely Pathogenic | Likely Benign | 2.41 | Destabilizing | 0.3 | 1.63 | Ambiguous | 2.02 | Destabilizing | 1.17 | Destabilizing | 0.207 | Likely Benign | -4.10 | Deleterious | 1.000 | Probably Damaging | 0.978 | Probably Damaging | 3.37 | Benign | 0.02 | Affected | 0.3552 | 0.1474 | -3 | -3 | -5.4 | -16.04 | |||||||||||||||||||||||||||||
| c.2146C>G | R716G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R716G is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized, whereas a majority of tools predict a pathogenic impact: PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, and AlphaMissense‑Default. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is classified as Likely Pathogenic. High‑accuracy assessments show AlphaMissense‑Optimized as Benign, SGM‑Consensus as Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as Uncertain. Other stability predictors (FoldX, Rosetta, premPS) are also Uncertain. Overall, the balance of evidence from the majority of tools and the SGM‑Consensus indicates a pathogenic effect, and this conclusion does not contradict any ClinVar annotation because no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.247041 | Structured | 0.419135 | Uncertain | 0.962 | 0.379 | 0.000 | -9.927 | Likely Pathogenic | 0.728 | Likely Pathogenic | Likely Benign | 1.32 | Ambiguous | 0.1 | 1.63 | Ambiguous | 1.48 | Ambiguous | 0.72 | Ambiguous | 0.359 | Likely Benign | -5.70 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.36 | Benign | 0.01 | Affected | 0.3437 | 0.2466 | -3 | -2 | 4.1 | -99.14 | |||||||||||||||||||||||||||||
| c.788T>G | V263G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 V263G missense change is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are FATHMM and AlphaMissense‑Optimized; those that agree on a pathogenic effect are REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. Rosetta and Foldetta give uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized predicts benign, while the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts pathogenic. Foldetta’s stability prediction is uncertain. Overall, the preponderance of evidence points to a pathogenic impact for V263G, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.268042 | Structured | 0.356141 | Uncertain | 0.918 | 0.257 | 0.000 | -10.388 | Likely Pathogenic | 0.669 | Likely Pathogenic | Likely Benign | 2.27 | Destabilizing | 0.2 | 1.63 | Ambiguous | 1.95 | Ambiguous | 1.88 | Destabilizing | 0.820 | Likely Pathogenic | -4.59 | Deleterious | 0.991 | Probably Damaging | 0.999 | Probably Damaging | 6.07 | Benign | 0.01 | Affected | 0.1790 | 0.1868 | -1 | -3 | -4.6 | -42.08 | ||||||||||||||||||||||||||||||
| c.856C>G | L286V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L286V is not reported in ClinVar (ClinVar status: none) and is absent from gnomAD (gnomAD ID: none). Prediction tools that agree on a benign effect include only AlphaMissense‑Optimized, whereas the remaining tools—SGM‑Consensus, REVEL, FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and FATHMM—consistently predict a pathogenic impact; AlphaMissense‑Default and Rosetta are uncertain. High‑accuracy methods give a mixed picture: AlphaMissense‑Optimized indicates benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) yields pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenic. Overall, the preponderance of evidence points to a pathogenic effect. This conclusion does not contradict ClinVar status, as no ClinVar assertion exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.122885 | Structured | 0.385647 | Uncertain | 0.932 | 0.260 | 0.000 | -9.986 | Likely Pathogenic | 0.500 | Ambiguous | Likely Benign | 2.44 | Destabilizing | 0.5 | 1.63 | Ambiguous | 2.04 | Destabilizing | 1.26 | Destabilizing | 0.676 | Likely Pathogenic | -2.76 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | 1.71 | Pathogenic | 0.01 | Affected | 0.1492 | 0.3007 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.1423C>G | R475G 2D  AIThe SynGAP1 missense variant R475G is not reported in ClinVar and is absent from gnomAD. Prediction tools that assess pathogenicity largely agree on a deleterious effect: SGM‑Consensus, REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default all predict pathogenicity. Only Rosetta and AlphaMissense‑Optimized return uncertain results, and no tool predicts a benign outcome. High‑accuracy methods provide a consistent view: AlphaMissense‑Optimized is uncertain, SGM‑Consensus indicates likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenicity. Taken together, the overwhelming majority of evidence supports a pathogenic classification for R475G, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.175930 | Structured | 0.382696 | Uncertain | 0.852 | 0.261 | 0.000 | -14.466 | Likely Pathogenic | 0.939 | Likely Pathogenic | Ambiguous | 2.39 | Destabilizing | 1.0 | 1.64 | Ambiguous | 2.02 | Destabilizing | 1.11 | Destabilizing | 0.697 | Likely Pathogenic | -6.53 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.41 | Pathogenic | 0.00 | Affected | 0.2779 | 0.2310 | -3 | -2 | 4.1 | -99.14 | |||||||||||||||||||||||||||||
| c.1969T>A | W657R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant W657R is not reported in ClinVar and has no entries in gnomAD. Functional prediction tools show a split: benign calls come from REVEL, SIFT, and FATHMM, whereas pathogenic calls are made by premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus score, which is labeled Likely Pathogenic. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates Likely Pathogenic, and the Foldetta stability analysis is inconclusive. Overall, the majority of evidence points to a pathogenic impact for W657R, and this conclusion does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.030611 | Structured | 0.208729 | Uncertain | 0.941 | 0.245 | 0.000 | -13.391 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 1.56 | Ambiguous | 0.2 | 1.64 | Ambiguous | 1.60 | Ambiguous | 1.29 | Destabilizing | 0.461 | Likely Benign | -11.96 | Deleterious | 0.999 | Probably Damaging | 0.964 | Probably Damaging | 3.48 | Benign | 0.07 | Tolerated | 0.4684 | 0.0000 | 2 | -3 | -3.6 | -30.03 | |||||||||||||||||||||||||||||
| c.1969T>C | W657R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant W657R is not reported in ClinVar and has no entry in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, and FATHMM, while pathogenic predictions are made by premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is classified as Likely Pathogenic. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority of the four high‑accuracy tools) is pathogenic, and Foldetta, which integrates FoldX‑MD and Rosetta outputs, is inconclusive. FoldX and Rosetta individually report uncertain effects on protein stability. Overall, the preponderance of evidence from both general and high‑accuracy predictors indicates that the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.030611 | Structured | 0.208729 | Uncertain | 0.941 | 0.245 | 0.000 | -13.391 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 1.56 | Ambiguous | 0.2 | 1.64 | Ambiguous | 1.60 | Ambiguous | 1.29 | Destabilizing | 0.461 | Likely Benign | -11.96 | Deleterious | 0.999 | Probably Damaging | 0.964 | Probably Damaging | 3.48 | Benign | 0.07 | Tolerated | 0.4684 | 0.0000 | 2 | -3 | -3.6 | -30.03 | |||||||||||||||||||||||||||||
| c.1494G>A | M498I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M498I has no ClinVar record (ClinVar ID None) and is not reported in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are FoldX, SIFT, FATHMM, and AlphaMissense‑Default. Two tools, Rosetta and premPS, give uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, Foldetta as pathogenic, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive due to a 2‑to‑2 split. Overall, the majority of predictions lean toward a benign impact, and this is not contradicted by ClinVar status, which lacks an entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.092881 | Structured | 0.399612 | Uncertain | 0.932 | 0.158 | 0.000 | -4.796 | Likely Benign | 0.760 | Likely Pathogenic | Likely Benign | 2.56 | Destabilizing | 0.3 | 1.65 | Ambiguous | 2.11 | Destabilizing | 0.88 | Ambiguous | 0.418 | Likely Benign | -1.09 | Neutral | 0.018 | Benign | 0.012 | Benign | -1.28 | Pathogenic | 0.04 | Affected | 0.1184 | 0.2537 | 2 | 1 | 2.6 | -18.03 | ||||||||||||||||||||||||||||||
| c.1494G>C | M498I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M498I has no ClinVar record (ClinVar ID None) and is not reported in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are FoldX, SIFT, FATHMM, and AlphaMissense‑Default. Two tools, Rosetta and premPS, give uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, Foldetta as pathogenic, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive due to a 2‑to‑2 split. Overall, the majority of predictions lean toward a benign impact, and this is not contradicted by ClinVar status, which lacks an entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.092881 | Structured | 0.399612 | Uncertain | 0.932 | 0.158 | 0.000 | -4.796 | Likely Benign | 0.760 | Likely Pathogenic | Likely Benign | 2.56 | Destabilizing | 0.3 | 1.65 | Ambiguous | 2.11 | Destabilizing | 0.88 | Ambiguous | 0.418 | Likely Benign | -1.09 | Neutral | 0.018 | Benign | 0.012 | Benign | -1.28 | Pathogenic | 0.04 | Affected | 0.1184 | 0.2537 | 2 | 1 | 2.6 | -18.03 | ||||||||||||||||||||||||||||||
| c.1494G>T | M498I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M498I has no ClinVar record (ClinVar ID None) and is not reported in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are FoldX, SIFT, FATHMM, and AlphaMissense‑Default. Two tools, Rosetta and premPS, give uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, Foldetta as pathogenic, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive due to a 2‑to‑2 split. Overall, the majority of predictions lean toward a benign impact, and this is not contradicted by ClinVar status, which lacks an entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.092881 | Structured | 0.399612 | Uncertain | 0.932 | 0.158 | 0.000 | -4.796 | Likely Benign | 0.760 | Likely Pathogenic | Likely Benign | 2.56 | Destabilizing | 0.3 | 1.65 | Ambiguous | 2.11 | Destabilizing | 0.88 | Ambiguous | 0.414 | Likely Benign | -1.09 | Neutral | 0.018 | Benign | 0.012 | Benign | -1.28 | Pathogenic | 0.04 | Affected | 0.1184 | 0.2537 | 2 | 1 | 2.6 | -18.03 | ||||||||||||||||||||||||||||||
| c.691T>A | F231I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F231I is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include polyPhen‑2 HumVar and FATHMM, whereas the majority of tools predict a pathogenic impact: REVEL, SIFT, PROVEAN, polyPhen‑2 HumDiv, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Pathogenic). High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, while Foldetta’s stability prediction is uncertain and thus not considered evidence. No other tools provide definitive pathogenic or benign conclusions. Based on the preponderance of pathogenic predictions and the lack of contrary evidence, the variant is most likely pathogenic; this assessment does not contradict any ClinVar status, as none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.366687 | Structured | 0.306467 | Uncertain | 0.895 | 0.300 | 0.000 | -13.827 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 1.16 | Ambiguous | 0.4 | 1.65 | Ambiguous | 1.41 | Ambiguous | 0.94 | Ambiguous | 0.894 | Likely Pathogenic | -5.01 | Deleterious | 0.759 | Possibly Damaging | 0.328 | Benign | 5.76 | Benign | 0.00 | Affected | 0.2122 | 0.2813 | 1 | 0 | 1.7 | -34.02 | |||||||||||||||||||||||||||||
| c.914C>A | T305N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant T305N is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the substitution as tolerated or benign, and the SGM‑Consensus score is “Likely Benign.” Only FATHMM predicts a pathogenic outcome. Stability‑based methods (FoldX, Rosetta, and the combined Foldetta) return uncertain or inconclusive results. High‑accuracy assessments reinforce the benign consensus: AlphaMissense‑Optimized is benign, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is also benign, and Foldetta remains uncertain. Overall, the majority of evidence supports a benign impact for T305N, and this conclusion is consistent with the absence of a ClinVar pathogenic classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.359901 | Structured | 0.299706 | Uncertain | 0.872 | 0.274 | 0.125 | -3.261 | Likely Benign | 0.158 | Likely Benign | Likely Benign | 1.18 | Ambiguous | 0.2 | 1.65 | Ambiguous | 1.42 | Ambiguous | 0.09 | Likely Benign | 0.098 | Likely Benign | 0.71 | Neutral | 0.046 | Benign | 0.040 | Benign | 2.34 | Pathogenic | 0.67 | Tolerated | 0.1327 | 0.4352 | 0 | 0 | -2.8 | 13.00 | |||||||||||||||||||||||||||||
| c.929A>C | E310A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E310A missense variant is not reported in ClinVar or gnomAD. Prediction tools largely converge on a pathogenic effect: SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all indicate pathogenicity, while premPS is the sole benign predictor. Uncertain calls come from FoldX, Rosetta, and Foldetta. High‑accuracy methods reinforce this view: AlphaMissense‑Optimized predicts pathogenic, SGM‑Consensus labels it likely pathogenic, and Foldetta remains inconclusive. With the overwhelming majority of evidence pointing to deleterious impact and no ClinVar annotation to contradict, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.222385 | Structured | 0.346136 | Uncertain | 0.914 | 0.337 | 0.125 | -13.878 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 1.65 | Ambiguous | 0.6 | 1.65 | Ambiguous | 1.65 | Ambiguous | 0.50 | Likely Benign | 0.850 | Likely Pathogenic | -5.52 | Deleterious | 0.999 | Probably Damaging | 0.995 | Probably Damaging | 1.16 | Pathogenic | 0.01 | Affected | 0.3980 | 0.8097 | 0 | -1 | 5.3 | -58.04 | |||||||||||||||||||||||||||||
| c.1034T>G | L345R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L345R is not reported in ClinVar (ClinVar ID None) and has no entries in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are REVEL and AlphaMissense‑Optimized, whereas the majority of tools predict pathogenicity: SGM‑Consensus, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. High‑accuracy assessments further support this: AlphaMissense‑Optimized classifies the variant as benign, while the SGM Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) yields a pathogenic prediction. Foldetta, which integrates FoldX‑MD and Rosetta outputs, is inconclusive. Consequently, the variant is most likely pathogenic based on the collective predictions, and this assessment does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.260850 | Structured | 0.354989 | Uncertain | 0.936 | 0.478 | 0.125 | -10.325 | Likely Pathogenic | 0.635 | Likely Pathogenic | Likely Benign | 0.96 | Ambiguous | 0.1 | 1.66 | Ambiguous | 1.31 | Ambiguous | 1.28 | Destabilizing | 0.247 | Likely Benign | -3.98 | Deleterious | 0.993 | Probably Damaging | 0.796 | Possibly Damaging | 1.81 | Pathogenic | 0.04 | Affected | 0.1140 | 0.0685 | -3 | -2 | -8.3 | 43.03 | |||||||||||||||||||||||||||||
| c.1072T>A | F358I 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant F358I is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions from REVEL, SIFT, and FATHMM; pathogenic predictions from PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, and ESM1b. Five tools (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Optimized) give uncertain or inconclusive results. High‑accuracy assessments further clarify the picture: AlphaMissense‑Optimized remains uncertain; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, indicates a likely pathogenic effect; Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, also reports an uncertain impact. Overall, the preponderance of evidence points to a pathogenic effect for F358I, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.222385 | Structured | 0.407113 | Uncertain | 0.912 | 0.441 | 0.250 | -10.636 | Likely Pathogenic | 0.884 | Likely Pathogenic | Ambiguous | 0.93 | Ambiguous | 0.2 | 1.66 | Ambiguous | 1.30 | Ambiguous | 0.95 | Ambiguous | 0.393 | Likely Benign | -4.45 | Deleterious | 0.993 | Probably Damaging | 0.977 | Probably Damaging | 4.07 | Benign | 0.13 | Tolerated | 0.2331 | 0.2821 | 1 | 0 | 1.7 | -34.02 | |||||||||||||||||||||||||||||
| c.1150G>A | G384S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G384S (gnomAD ID 6-33438055‑G‑A) is listed in ClinVar with an uncertain significance. Functional prediction tools cluster into two groups: benign predictions from REVEL, premPS, PROVEAN, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized; pathogenic predictions from polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and FATHMM. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a likely benign outcome. High‑accuracy assessments further support benignity: AlphaMissense‑Optimized predicts benign, the SGM‑Consensus (majority vote) is likely benign, and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is inconclusive. No evidence from FoldX or Rosetta alone is available. Overall, the preponderance of evidence points to a benign effect, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.728858 | Disordered | 0.427831 | Uncertain | 0.323 | 0.934 | 0.750 | Uncertain | 1 | 6-33438055-G-A | 1 | 6.22e-7 | -5.243 | Likely Benign | 0.090 | Likely Benign | Likely Benign | 1.92 | Ambiguous | 0.2 | 1.66 | Ambiguous | 1.79 | Ambiguous | 0.19 | Likely Benign | 0.315 | Likely Benign | -0.67 | Neutral | 0.980 | Probably Damaging | 0.968 | Probably Damaging | 1.33 | Pathogenic | 0.04 | Affected | 4.32 | 2 | 0.2905 | 0.4924 | 1 | 0 | -0.4 | 30.03 | 202.4 | -49.8 | 0.5 | 1.0 | -0.2 | 0.0 | Uncertain | Gly384 is located in the Gly-rich Ω loop (res. Pro364-Pro398) between two anti-parallel β sheet strands (res. Thr359-Pro364, res. Ala399-Ile411). Because the Ω loop is assumed to directly interact with the membrane, it moves arbitrarily throughout the WT solvent simulations. The Ω loop potentially plays a crucial role in the SynGAP-membrane complex association, stability, and dynamics. However, this aspect cannot be fully addressed through solvent simulations alone.Ω loops are known to play major roles in protein functions that require flexibility, and so they are rich in glycines, prolines, and, to a lesser extent, small hydrophilic residues to ensure maximum flexibility. Thus, the variant’s Ser384 is potentially tolerated in the Ω loop, although the hydroxyl group of Ser384 forms various hydrogen bonds with several other loop residues in the variant simulations. However, since the effects on Gly-rich Ω loop dynamics can only be studied through the SynGAP-membrane complex, no definite conclusions can be drawn. | ||||||||||||||
| c.1643A>G | E548G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E548G missense variant is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include SIFT and FATHMM, while the majority of other in silico predictors (REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) indicate a pathogenic impact; SGM‑Consensus also classifies the variant as likely pathogenic. Uncertain results are reported by FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized predicting pathogenicity, SGM‑Consensus confirming a likely pathogenic status, and Foldetta yielding an inconclusive stability change. Overall, the consensus of the available predictions points to a pathogenic effect for E548G, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.054297 | Structured | 0.008632 | Uncertain | 0.965 | 0.288 | 0.000 | -12.010 | Likely Pathogenic | 0.990 | Likely Pathogenic | Likely Pathogenic | 0.97 | Ambiguous | 0.1 | 1.66 | Ambiguous | 1.32 | Ambiguous | 0.59 | Ambiguous | 0.521 | Likely Pathogenic | -6.73 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.32 | Benign | 0.06 | Tolerated | 0.2638 | 0.3654 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.1936C>A | L646M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L646M is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool predicts a pathogenic outcome. Predictions that are inconclusive are FoldX, Rosetta, and Foldetta. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely benign, while Foldetta’s stability analysis remains uncertain. Overall, the evidence overwhelmingly supports a benign classification for this variant, and this conclusion does not contradict the ClinVar status, which currently has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.048328 | Structured | 0.303751 | Uncertain | 0.941 | 0.344 | 0.000 | -1.911 | Likely Benign | 0.152 | Likely Benign | Likely Benign | 0.70 | Ambiguous | 0.1 | 1.66 | Ambiguous | 1.18 | Ambiguous | -1.10 | Stabilizing | 0.106 | Likely Benign | 1.86 | Neutral | 0.211 | Benign | 0.055 | Benign | 3.57 | Benign | 1.00 | Tolerated | 0.2041 | 0.3427 | 4 | 2 | -1.9 | 18.03 | |||||||||||||||||||||||||||||
| c.807A>G | I269M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I269M is not reported in ClinVar and is absent from gnomAD. Prediction tools that classify it as benign include PROVEAN and AlphaMissense‑Optimized, whereas REVEL, polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, and AlphaMissense‑Default predict it to be pathogenic. Five tools (FoldX, Rosetta, Foldetta, premPS, and ESM1b) give uncertain results. High‑accuracy methods give mixed evidence: AlphaMissense‑Optimized reports a benign effect, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts pathogenicity, and Foldetta remains uncertain. Overall, the majority of predictions support a pathogenic impact, and this assessment does not contradict the absence of a ClinVar entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.216401 | Structured | 0.343787 | Uncertain | 0.937 | 0.244 | 0.125 | -7.863 | In-Between | 0.715 | Likely Pathogenic | Likely Benign | 0.91 | Ambiguous | 0.1 | 1.66 | Ambiguous | 1.29 | Ambiguous | 0.94 | Ambiguous | 0.507 | Likely Pathogenic | -2.19 | Neutral | 0.999 | Probably Damaging | 0.998 | Probably Damaging | 1.75 | Pathogenic | 0.05 | Affected | 0.0580 | 0.2283 | 2 | 1 | -2.6 | 18.03 | ||||||||||||||||||||||||||||||
| c.1386G>C | K462N 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant K462N is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign predictions come from REVEL, SIFT, and FATHMM, whereas pathogenic predictions are returned by PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as Likely Pathogenic. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts Pathogenic, and the SGM Consensus also indicates Likely Pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta outputs, yields an uncertain result, providing no definitive evidence. Overall, the majority of high‑confidence predictors lean toward pathogenicity, contradicting the absence of a ClinVar classification. Thus, the variant is most likely pathogenic based on current predictions. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.264545 | Structured | 0.297737 | Uncertain | 0.921 | 0.159 | 0.125 | -12.823 | Likely Pathogenic | 0.994 | Likely Pathogenic | Likely Pathogenic | 0.69 | Ambiguous | 0.1 | 1.67 | Ambiguous | 1.18 | Ambiguous | 0.90 | Ambiguous | 0.304 | Likely Benign | -4.83 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 3.42 | Benign | 0.07 | Tolerated | 3.37 | 34 | 0.3967 | 0.1242 | 0 | 1 | 0.4 | -14.07 | |||||||||||||||||||||||||||
| c.1386G>T | K462N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K462N is reported in gnomAD (ID 6‑33438291‑G‑T) but has no ClinVar entry. Functional prediction tools cluster into two groups: benign predictions from REVEL, SIFT, and FATHMM; pathogenic predictions from PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Stability‑based methods (FoldX, Rosetta, premPS) and Foldetta are inconclusive. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, SGM Consensus as likely pathogenic, while Foldetta remains uncertain. Overall, the preponderance of evidence points to a pathogenic effect for K462N. This conclusion is not contradicted by ClinVar, which contains no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.264545 | Structured | 0.297737 | Uncertain | 0.921 | 0.159 | 0.125 | 6-33438291-G-T | 1 | 6.20e-7 | -12.823 | Likely Pathogenic | 0.994 | Likely Pathogenic | Likely Pathogenic | 0.69 | Ambiguous | 0.1 | 1.67 | Ambiguous | 1.18 | Ambiguous | 0.90 | Ambiguous | 0.303 | Likely Benign | -4.83 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 3.42 | Benign | 0.07 | Tolerated | 3.37 | 34 | 0.3967 | 0.1242 | 0 | 1 | 0.4 | -14.07 | ||||||||||||||||||||||||
| c.1708G>A | A570T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A570T is not reported in ClinVar and is absent from gnomAD. Prediction tools that provide a definitive call all indicate a deleterious effect: SGM‑Consensus (Likely Pathogenic), REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default all predict pathogenicity. No tool reports a benign outcome; the remaining predictions (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Optimized) are uncertain and therefore do not influence the overall assessment. High‑accuracy methods specifically show SGM‑Consensus as Likely Pathogenic, AlphaMissense‑Optimized as uncertain, and Foldetta as uncertain. Taken together, the majority of conclusive predictions support a pathogenic effect. Consequently, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status, as no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.046336 | Structured | 0.054494 | Uncertain | 0.932 | 0.263 | 0.000 | -11.390 | Likely Pathogenic | 0.801 | Likely Pathogenic | Ambiguous | 1.45 | Ambiguous | 0.3 | 1.67 | Ambiguous | 1.56 | Ambiguous | 0.86 | Ambiguous | 0.568 | Likely Pathogenic | -3.28 | Deleterious | 0.998 | Probably Damaging | 0.993 | Probably Damaging | -1.26 | Pathogenic | 0.05 | Affected | 0.1345 | 0.3874 | 1 | 0 | -2.5 | 30.03 | |||||||||||||||||||||||||||||
| c.1780T>A | F594I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F594I is not reported in ClinVar and has no entries in gnomAD. In silico assessment shows that all evaluated pathogenicity predictors—REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default—classify the substitution as pathogenic, while FoldX, Rosetta, and Foldetta provide inconclusive results. High‑accuracy tools further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates likely pathogenic, and Foldetta remains uncertain. Consequently, the overwhelming majority of predictions point to a pathogenic impact. The variant is therefore most likely pathogenic, and this assessment does not contradict any ClinVar annotation, as none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.009187 | Structured | 0.120166 | Uncertain | 0.946 | 0.147 | 0.000 | -11.271 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 1.93 | Ambiguous | 0.1 | 1.67 | Ambiguous | 1.80 | Ambiguous | 1.61 | Destabilizing | 0.935 | Likely Pathogenic | -5.97 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | -1.91 | Pathogenic | 0.02 | Affected | 0.1694 | 0.1925 | 1 | 0 | 1.7 | -34.02 | |||||||||||||||||||||||||||||
| c.743G>A | R248Q 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant R248Q is catalogued in gnomAD (ID 6‑33435594‑G‑A) but has no ClinVar entry. Functional prediction tools largely agree on a deleterious effect: benign predictions come from FoldX and FATHMM, whereas pathogenic predictions are made by REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Pathogenic). Uncertain results are reported by Rosetta and Foldetta. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts Pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta remains Uncertain. Overall, the preponderance of evidence points to a pathogenic effect for R248Q, and this conclusion is not contradicted by ClinVar, which contains no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.513880 | Disordered | 0.267126 | Uncertain | 0.781 | 0.346 | 0.250 | 6-33435594-G-A | 2 | 1.24e-6 | -10.573 | Likely Pathogenic | 0.979 | Likely Pathogenic | Likely Pathogenic | 0.45 | Likely Benign | 0.2 | 1.67 | Ambiguous | 1.06 | Ambiguous | 1.05 | Destabilizing | 0.739 | Likely Pathogenic | -3.34 | Deleterious | 0.999 | Probably Damaging | 0.715 | Possibly Damaging | 5.74 | Benign | 0.01 | Affected | 3.41 | 14 | 0.2572 | 0.2549 | 1 | 1 | 1.0 | -28.06 | ||||||||||||||||||||||||
| c.959T>A | V320D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V320D is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions from REVEL and SIFT; pathogenic predictions from premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Pathogenic). Stability‑based methods FoldX and Rosetta returned uncertain results, and Foldetta, which integrates FoldX‑MD and Rosetta outputs, also yielded an uncertain prediction. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points toward a pathogenic effect, which is consistent with the lack of ClinVar annotation and gnomAD absence. Thus, the variant is most likely pathogenic, and this prediction does not contradict ClinVar status because ClinVar has no entry for it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.185198 | Structured | 0.419626 | Uncertain | 0.905 | 0.266 | 0.125 | -11.269 | Likely Pathogenic | 0.996 | Likely Pathogenic | Likely Pathogenic | 1.99 | Ambiguous | 1.0 | 1.67 | Ambiguous | 1.83 | Ambiguous | 1.50 | Destabilizing | 0.405 | Likely Benign | -5.58 | Deleterious | 0.999 | Probably Damaging | 0.972 | Probably Damaging | 1.93 | Pathogenic | 0.07 | Tolerated | 0.1101 | 0.0482 | -2 | -3 | -7.7 | 15.96 | |||||||||||||||||||||||||||||
| c.978T>A | H326Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant H326Q is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, FoldX, and SIFT, whereas a majority of tools predict a pathogenic outcome: premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Pathogenic). Two tools, Foldetta (combining FoldX‑MD and Rosetta) and Rosetta alone, return uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. Overall, the preponderance of evidence points to a pathogenic effect for H326Q. This conclusion is not contradicted by ClinVar status, which currently contains no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.342579 | Structured | 0.418150 | Uncertain | 0.944 | 0.455 | 0.000 | -8.688 | Likely Pathogenic | 0.976 | Likely Pathogenic | Likely Pathogenic | 0.46 | Likely Benign | 0.1 | 1.67 | Ambiguous | 1.07 | Ambiguous | 1.00 | Destabilizing | 0.444 | Likely Benign | -6.89 | Deleterious | 0.999 | Probably Damaging | 0.996 | Probably Damaging | 2.07 | Pathogenic | 0.10 | Tolerated | 0.1485 | 0.3500 | 3 | 0 | -0.3 | -9.01 | |||||||||||||||||||||||||||||
| c.978T>G | H326Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant H326Q is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, FoldX, and SIFT, whereas a majority of tools predict a pathogenic outcome: premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Pathogenic). Two tools, Foldetta (combining FoldX‑MD and Rosetta) and Rosetta alone, return uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus also indicates likely pathogenic, while Foldetta remains uncertain. Overall, the preponderance of evidence from multiple independent predictors points to a pathogenic effect for H326Q. This conclusion is not contradicted by ClinVar status, which currently contains no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.342579 | Structured | 0.418150 | Uncertain | 0.944 | 0.455 | 0.000 | -8.688 | Likely Pathogenic | 0.976 | Likely Pathogenic | Likely Pathogenic | 0.46 | Likely Benign | 0.1 | 1.67 | Ambiguous | 1.07 | Ambiguous | 1.00 | Destabilizing | 0.444 | Likely Benign | -6.89 | Deleterious | 0.999 | Probably Damaging | 0.996 | Probably Damaging | 2.07 | Pathogenic | 0.10 | Tolerated | 0.1485 | 0.3500 | 3 | 0 | -0.3 | -9.01 | |||||||||||||||||||||||||||||
| c.1045C>G | P349A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P349A is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions include REVEL, AlphaMissense‑Default, and AlphaMissense‑Optimized; pathogenic predictions include PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and FATHMM, with the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) labeling it likely pathogenic. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. No other stability or pathogenicity scores are available. Overall, the majority of evidence points to a pathogenic effect, and this assessment does not contradict any ClinVar annotation because the variant is not yet catalogued there. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.167087 | Structured | 0.348607 | Uncertain | 0.947 | 0.396 | 0.000 | -8.663 | Likely Pathogenic | 0.202 | Likely Benign | Likely Benign | 1.37 | Ambiguous | 0.0 | 1.68 | Ambiguous | 1.53 | Ambiguous | 0.76 | Ambiguous | 0.257 | Likely Benign | -6.01 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | 1.54 | Pathogenic | 0.01 | Affected | 0.3789 | 0.5505 | 1 | -1 | 3.4 | -26.04 | |||||||||||||||||||||||||||||
| c.1072T>G | F358V 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant F358V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions from REVEL, SIFT, and FATHMM; pathogenic predictions from PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, and ESM1b. Five tools (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Optimized) give uncertain or inconclusive results. High‑accuracy assessments further clarify the picture: AlphaMissense‑Optimized remains uncertain; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is classified as Likely Pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is also uncertain. Overall, the preponderance of evidence points to a pathogenic effect for F358V, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.222385 | Structured | 0.407113 | Uncertain | 0.912 | 0.441 | 0.250 | -9.021 | Likely Pathogenic | 0.847 | Likely Pathogenic | Ambiguous | 1.42 | Ambiguous | 0.2 | 1.68 | Ambiguous | 1.55 | Ambiguous | 0.93 | Ambiguous | 0.408 | Likely Benign | -5.32 | Deleterious | 0.993 | Probably Damaging | 0.968 | Probably Damaging | 4.09 | Benign | 0.18 | Tolerated | 0.2222 | 0.2978 | -1 | -1 | 1.4 | -48.04 | |||||||||||||||||||||||||||||
| c.1160G>C | G387A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G387A is reported in gnomAD (variant ID 6‑33438065‑G‑C) but has no ClinVar entry. Functional prediction tools largely agree on a benign effect: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the change as benign. Only FATHMM predicts it as pathogenic. High‑accuracy assessments reinforce the benign consensus: AlphaMissense‑Optimized is benign; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is “Likely Benign”; and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is inconclusive (treated as unavailable). No other tools provide decisive evidence. Based on the collective predictions, the variant is most likely benign, and this assessment does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.642678 | Disordered | 0.422910 | Uncertain | 0.293 | 0.861 | 0.750 | 6-33438065-G-C | 3 | 1.87e-6 | -6.313 | Likely Benign | 0.104 | Likely Benign | Likely Benign | 1.95 | Ambiguous | 0.1 | 1.68 | Ambiguous | 1.82 | Ambiguous | -0.02 | Likely Benign | 0.300 | Likely Benign | -0.29 | Neutral | 0.007 | Benign | 0.010 | Benign | 1.33 | Pathogenic | 0.06 | Tolerated | 4.32 | 3 | 0.4149 | 0.4576 | 0 | 1 | 2.2 | 14.03 | ||||||||||||||||||||||||
| c.1219C>G | Q407E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant Q407E is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized, whereas premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and ESM1b all predict a pathogenic outcome. AlphaMissense‑Default, FoldX, Rosetta, and Foldetta are uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the majority of tools and the SGM Consensus support a pathogenic classification, while a minority predict benign. No ClinVar entry exists to contradict these predictions. Thus, the variant is most likely pathogenic based on the available computational evidence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.109221 | Structured | 0.382522 | Uncertain | 0.916 | 0.271 | 0.000 | -12.631 | Likely Pathogenic | 0.466 | Ambiguous | Likely Benign | 0.50 | Ambiguous | 0.1 | 1.68 | Ambiguous | 1.09 | Ambiguous | 1.30 | Destabilizing | 0.243 | Likely Benign | -2.66 | Deleterious | 0.989 | Probably Damaging | 0.930 | Probably Damaging | 3.96 | Benign | 0.03 | Affected | 0.1199 | 0.2000 | 2 | 2 | 0.0 | 0.98 | ||||||||||||||||||||||||||||||
| c.1285C>G | R429G 2D  3DClick to see structure in 3D Viewer AISynGAP1 R429G is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized; pathogenic predictions come from premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized predicts benign, while the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive due to a 2‑vs‑2 split, and Foldetta’s stability analysis is uncertain. No evidence from FoldX or Rosetta is available. Overall, the predictions are evenly split, with no clear consensus. The variant is most likely of uncertain significance; it is not contradicted by ClinVar status, which has no entry for this change. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.074921 | Structured | 0.390504 | Uncertain | 0.959 | 0.290 | 0.000 | -9.157 | Likely Pathogenic | 0.333 | Likely Benign | Likely Benign | 1.58 | Ambiguous | 0.0 | 1.68 | Ambiguous | 1.63 | Ambiguous | 1.21 | Destabilizing | 0.257 | Likely Benign | -3.37 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.43 | Benign | 0.34 | Tolerated | 0.2888 | 0.2717 | -3 | -2 | 4.1 | -99.14 | ||||||||||||||||||||||||||||||
| c.1708G>T | A570S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A570S is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict a pathogenic outcome are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. The remaining tools (FoldX, Rosetta, Foldetta, premPS, and ESM1b) yield uncertain or inconclusive results and are treated as unavailable evidence. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also favors benign, while Foldetta remains uncertain. Overall, the majority of reliable predictions indicate a benign effect. This conclusion does not contradict ClinVar status, as no ClinVar assertion exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.046336 | Structured | 0.054494 | Uncertain | 0.932 | 0.263 | 0.000 | -7.893 | In-Between | 0.194 | Likely Benign | Likely Benign | 0.77 | Ambiguous | 0.1 | 1.68 | Ambiguous | 1.23 | Ambiguous | 0.51 | Ambiguous | 0.399 | Likely Benign | -2.26 | Neutral | 0.983 | Probably Damaging | 0.993 | Probably Damaging | -1.19 | Pathogenic | 0.17 | Tolerated | 0.2091 | 0.3256 | 1 | 1 | -2.6 | 16.00 | ||||||||||||||||||||||||||||||
| c.1933T>A | F645I 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant F645I is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show mixed results: benign predictions come from REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM, while pathogenic predictions are given by premPS, PROVEAN, SIFT, ESM1b, and AlphaMissense‑Default. Four tools (FoldX, Rosetta, Foldetta, AlphaMissense‑Optimized) report uncertain or inconclusive outcomes. High‑accuracy assessments indicate that AlphaMissense‑Optimized is uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic, and Foldetta is uncertain. Overall, the majority of evaluated predictors (five pathogenic vs four benign) lean toward a pathogenic effect. Therefore, the variant is most likely pathogenic, and this assessment does not contradict ClinVar status, which currently has no entry for F645I. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.046336 | Structured | 0.276445 | Uncertain | 0.921 | 0.325 | 0.000 | -13.055 | Likely Pathogenic | 0.878 | Likely Pathogenic | Ambiguous | 1.12 | Ambiguous | 0.2 | 1.68 | Ambiguous | 1.40 | Ambiguous | 1.01 | Destabilizing | 0.299 | Likely Benign | -4.24 | Deleterious | 0.190 | Benign | 0.019 | Benign | 3.44 | Benign | 0.04 | Affected | 0.1920 | 0.2891 | 1 | 0 | 1.7 | -34.02 | |||||||||||||||||||||||||||||
| c.2001C>G | I667M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I667M is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. Tools that agree on a pathogenic effect include SGM‑Consensus, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. Predictions from FoldX, Rosetta, Foldetta, and premPS are uncertain and are treated as unavailable. High‑accuracy methods give AlphaMissense‑Optimized a benign prediction, SGM‑Consensus a pathogenic prediction (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN), and Foldetta an uncertain result. Overall, the majority of evidence—including the high‑accuracy consensus—points to a pathogenic impact. This conclusion does not contradict ClinVar status, as no ClinVar assertion exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.142424 | Structured | 0.083597 | Uncertain | 0.927 | 0.379 | 0.000 | -10.679 | Likely Pathogenic | 0.579 | Likely Pathogenic | Likely Benign | 1.29 | Ambiguous | 0.3 | 1.68 | Ambiguous | 1.49 | Ambiguous | 0.92 | Ambiguous | 0.360 | Likely Benign | -2.90 | Deleterious | 1.000 | Probably Damaging | 0.993 | Probably Damaging | 2.98 | Benign | 0.00 | Affected | 0.0746 | 0.2612 | 2 | 1 | -2.6 | 18.03 | |||||||||||||||||||||||||||||
| c.2132T>A | L711Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L711Q is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect are REVEL and FATHMM. Tools that predict a pathogenic effect include FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, SGM Consensus, and Foldetta; Rosetta is uncertain. High‑accuracy methods give a consistent pathogenic signal: AlphaMissense‑Optimized predicts Pathogenic, SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts Pathogenic. No predictions are missing or inconclusive. Based on the preponderance of pathogenic predictions, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.308712 | Structured | 0.377436 | Uncertain | 0.950 | 0.364 | 0.000 | -11.792 | Likely Pathogenic | 0.994 | Likely Pathogenic | Likely Pathogenic | 2.93 | Destabilizing | 0.0 | 1.68 | Ambiguous | 2.31 | Destabilizing | 1.63 | Destabilizing | 0.388 | Likely Benign | -5.67 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.31 | Benign | 0.00 | Affected | 0.1041 | 0.0488 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.1162G>A | G388S 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant G388S is catalogued in gnomAD (ID 6‑33438067‑G‑A) but has no ClinVar entry. Functional prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that converge on a pathogenic interpretation are FoldX, polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM. The high‑accuracy AlphaMissense‑Optimized score is benign, while the SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also reports a likely benign outcome. In contrast, the protein‑folding stability predictor Foldetta, which integrates FoldX‑MD and Rosetta outputs, indicates a pathogenic effect. No prediction is available from Rosetta alone, and the SGM‑Consensus result aligns with the benign consensus. Overall, the majority of evidence points to a benign impact, and this assessment does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.736850 | Disordered | 0.420316 | Uncertain | 0.319 | 0.827 | 0.750 | 6-33438067-G-A | 1 | 6.21e-7 | -5.036 | Likely Benign | 0.089 | Likely Benign | Likely Benign | 3.42 | Destabilizing | 3.5 | 1.69 | Ambiguous | 2.56 | Destabilizing | 0.18 | Likely Benign | 0.446 | Likely Benign | -0.52 | Neutral | 0.980 | Probably Damaging | 0.968 | Probably Damaging | 1.33 | Pathogenic | 0.05 | Affected | 4.32 | 3 | 0.2901 | 0.4985 | 0 | 1 | -0.4 | 30.03 | ||||||||||||||||||||||||
| c.1873C>G | L625V 2D  AISynGAP1 missense variant L625V is listed in ClinVar with an uncertain significance (ClinVar ID 3392716.0) and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL and FATHMM, while pathogenic predictions are made by premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, and AlphaMissense‑Default. Four tools (FoldX, Rosetta, Foldetta, AlphaMissense‑Optimized) give inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points toward a pathogenic effect, which does not contradict the ClinVar uncertain status but suggests a higher likelihood of pathogenicity. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.229226 | Structured | 0.045896 | Uncertain | 0.966 | 0.215 | 0.000 | Uncertain | 1 | -11.319 | Likely Pathogenic | 0.833 | Likely Pathogenic | Ambiguous | 1.80 | Ambiguous | 0.7 | 1.69 | Ambiguous | 1.75 | Ambiguous | 1.42 | Destabilizing | 0.480 | Likely Benign | -2.96 | Deleterious | 0.998 | Probably Damaging | 0.992 | Probably Damaging | 3.07 | Benign | 0.01 | Affected | 0.1306 | 0.3427 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||
| c.1971G>C | W657C 2D  AISynGAP1 missense variant W657C is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools that classify the variant as benign include REVEL and FATHMM. Those that predict a deleterious effect are FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized; Rosetta reports an uncertain outcome. High‑accuracy assessments further support a damaging interpretation: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. Overall, the preponderance of evidence indicates that W657C is most likely pathogenic, which does not contradict the current ClinVar status of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.030611 | Structured | 0.208729 | Uncertain | 0.941 | 0.245 | 0.000 | Uncertain | 1 | -12.035 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 2.74 | Destabilizing | 0.3 | 1.69 | Ambiguous | 2.22 | Destabilizing | 1.30 | Destabilizing | 0.463 | Likely Benign | -11.06 | Deleterious | 1.000 | Probably Damaging | 0.982 | Probably Damaging | 3.43 | Benign | 0.03 | Affected | 0.3834 | 0.0766 | -8 | -2 | 3.4 | -83.07 | |||||||||||||||||||||||||||
| c.1971G>T | W657C 2D  AIThe SynGAP1 W657C missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are REVEL and FATHMM, whereas the majority of tools (SGM‑Consensus, FoldX, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic impact; Foldetta also supports pathogenicity, while Rosetta remains uncertain. High‑accuracy assessments further reinforce a deleterious outcome: AlphaMissense‑Optimized classifies the variant as pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates likely pathogenic; and Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, predicts a pathogenic effect. Overall, the preponderance of evidence points to the variant being most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.030611 | Structured | 0.208729 | Uncertain | 0.941 | 0.245 | 0.000 | -12.035 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 2.74 | Destabilizing | 0.3 | 1.69 | Ambiguous | 2.22 | Destabilizing | 1.30 | Destabilizing | 0.463 | Likely Benign | -11.06 | Deleterious | 1.000 | Probably Damaging | 0.982 | Probably Damaging | 3.43 | Benign | 0.03 | Affected | 0.3834 | 0.0766 | -8 | -2 | 3.4 | -83.07 | |||||||||||||||||||||||||||||
| c.2005A>C | N669H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N669H has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and ESM1b. Remaining tools (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Default) are uncertain and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points toward a pathogenic impact, and this conclusion is not contradicted by any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.142424 | Structured | 0.086615 | Uncertain | 0.872 | 0.380 | 0.000 | -10.364 | Likely Pathogenic | 0.421 | Ambiguous | Likely Benign | 1.26 | Ambiguous | 0.2 | 1.69 | Ambiguous | 1.48 | Ambiguous | 0.80 | Ambiguous | 0.432 | Likely Benign | -4.49 | Deleterious | 0.999 | Probably Damaging | 0.993 | Probably Damaging | 3.35 | Benign | 0.01 | Affected | 0.1732 | 0.4839 | 2 | 1 | 0.3 | 23.04 | ||||||||||||||||||||||||||||||
| c.952C>G | P318A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P318A missense variant has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a pathogenic effect include SGM‑Consensus (Likely Pathogenic), REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. No tool predicts a benign outcome. Uncertain or inconclusive results come from FoldX, Rosetta, Foldetta, premPS, and AlphaMissense‑Optimized. High‑accuracy assessments show that the SGM Consensus (derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts pathogenicity, while AlphaMissense‑Optimized remains uncertain and Foldetta is also uncertain. Taken together, the overwhelming majority of evidence points to a pathogenic effect. Therefore, the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which is currently unreported. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.111485 | Structured | 0.400936 | Uncertain | 0.858 | 0.234 | 0.000 | -9.642 | Likely Pathogenic | 0.872 | Likely Pathogenic | Ambiguous | 1.90 | Ambiguous | 0.2 | 1.69 | Ambiguous | 1.80 | Ambiguous | 0.94 | Ambiguous | 0.546 | Likely Pathogenic | -7.12 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 1.91 | Pathogenic | 0.04 | Affected | 0.3760 | 0.5585 | 1 | -1 | 3.4 | -26.04 | |||||||||||||||||||||||||||||
| c.1465C>G | L489V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L489V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: pathogenic calls come from SGM‑Consensus, REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and FATHMM, while only AlphaMissense‑Optimized predicts a benign outcome. Uncertain results are reported by Rosetta, Foldetta, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as inconclusive. Overall, the preponderance of evidence points to a pathogenic effect for L489V. This conclusion is consistent with the lack of ClinVar annotation and does not contradict any existing database record. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.191378 | Structured | 0.326126 | Uncertain | 0.949 | 0.234 | 0.125 | -9.345 | Likely Pathogenic | 0.467 | Ambiguous | Likely Benign | 2.09 | Destabilizing | 0.0 | 1.70 | Ambiguous | 1.90 | Ambiguous | 1.25 | Destabilizing | 0.586 | Likely Pathogenic | -2.55 | Deleterious | 0.998 | Probably Damaging | 0.992 | Probably Damaging | -1.34 | Pathogenic | 0.01 | Affected | 0.1665 | 0.4108 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.1468G>A | A490T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A490T is listed in gnomAD (variant ID 6‑33438500‑G‑A) but has no ClinVar entry. Prediction tools that agree on a pathogenic effect include REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. Tools that are inconclusive or uncertain are FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. No tool predicts a benign outcome. High‑accuracy assessments show that the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts pathogenicity, while AlphaMissense‑Optimized remains uncertain and Foldetta is also uncertain. Based on the preponderance of pathogenic predictions and the lack of any benign calls, the variant is most likely pathogenic. This conclusion is not contradicted by ClinVar, as no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.120615 | Structured | 0.322979 | Uncertain | 0.938 | 0.210 | 0.125 | 6-33438500-G-A | 1 | 6.20e-7 | -10.266 | Likely Pathogenic | 0.892 | Likely Pathogenic | Ambiguous | 0.80 | Ambiguous | 0.2 | 1.70 | Ambiguous | 1.25 | Ambiguous | 1.00 | Destabilizing | 0.821 | Likely Pathogenic | -3.87 | Deleterious | 0.998 | Probably Damaging | 0.993 | Probably Damaging | -1.34 | Pathogenic | 0.03 | Affected | 3.37 | 35 | 0.1065 | 0.4495 | 0 | 1 | -2.5 | 30.03 | ||||||||||||||||||||||||
| c.1502T>A | I501N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I501N is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect are REVEL and FATHMM, while a majority of tools (SGM‑Consensus, FoldX, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) predict a pathogenic impact. Uncertain results come from AlphaMissense‑Optimized, Foldetta, and Rosetta. High‑accuracy assessments show AlphaMissense‑Optimized as inconclusive, whereas the SGM Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates a likely pathogenic outcome; Foldetta likewise remains inconclusive. Overall, the balance of evidence favors a pathogenic classification, and this conclusion does not contradict any existing ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.079919 | Structured | 0.366596 | Uncertain | 0.886 | 0.153 | 0.000 | -11.105 | Likely Pathogenic | 0.854 | Likely Pathogenic | Ambiguous | 2.22 | Destabilizing | 0.0 | 1.70 | Ambiguous | 1.96 | Ambiguous | 1.90 | Destabilizing | 0.445 | Likely Benign | -6.13 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.37 | Benign | 0.01 | Affected | 0.0825 | 0.0270 | -2 | -3 | -8.0 | 0.94 | |||||||||||||||||||||||||||||
| c.1553A>G | Y518C 2D  AIThe SynGAP1 missense variant Y518C is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into three groups: benign predictions from REVEL, SIFT, and FATHMM; pathogenic predictions from SGM‑Consensus, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and AlphaMissense‑Default; and uncertain results from Rosetta, premPS, and AlphaMissense‑Optimized. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is inconclusive, SGM‑Consensus indicates likely pathogenic, and Foldetta predicts pathogenic folding instability. Overall, the majority of evidence points to a pathogenic impact, and this conclusion does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.139895 | Structured | 0.126970 | Uncertain | 0.897 | 0.321 | 0.000 | -9.813 | Likely Pathogenic | 0.794 | Likely Pathogenic | Ambiguous | 2.99 | Destabilizing | 0.9 | 1.70 | Ambiguous | 2.35 | Destabilizing | 0.69 | Ambiguous | 0.415 | Likely Benign | -8.35 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.45 | Benign | 0.16 | Tolerated | 0.2905 | 0.1128 | 0 | -2 | 3.8 | -60.04 | |||||||||||||||||||||||||||||
| c.1664T>A | V555D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V555D is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect are none, while a majority of algorithms predict a pathogenic impact: REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. FoldX, Rosetta, and Foldetta provide uncertain results. High‑accuracy methods reinforce the pathogenic prediction: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic, and Foldetta remains uncertain. Overall, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.013265 | Structured | 0.008218 | Uncertain | 0.943 | 0.225 | 0.000 | -16.413 | Likely Pathogenic | 0.994 | Likely Pathogenic | Likely Pathogenic | 1.84 | Ambiguous | 0.1 | 1.70 | Ambiguous | 1.77 | Ambiguous | 1.25 | Destabilizing | 0.896 | Likely Pathogenic | -5.71 | Deleterious | 1.000 | Probably Damaging | 0.987 | Probably Damaging | -1.39 | Pathogenic | 0.00 | Affected | 0.1411 | 0.0541 | -2 | -3 | -7.7 | 15.96 | |||||||||||||||||||||||||||||
| c.1741C>G | R581G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R581G is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that assess pathogenicity all converge on a deleterious effect: REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict pathogenicity. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also indicates “Likely Pathogenic.” High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized is pathogenic; the SGM‑Consensus is likely pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, predicts pathogenicity. Uncertain results from Rosetta and premPS are treated as unavailable and do not alter the overall conclusion. **Thus, the variant is most likely pathogenic based on the available predictions, and this assessment does not contradict any ClinVar status (none reported).** Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.104810 | Structured | 0.029544 | Uncertain | 0.829 | 0.236 | 0.000 | -12.097 | Likely Pathogenic | 0.985 | Likely Pathogenic | Likely Pathogenic | 2.48 | Destabilizing | 0.2 | 1.70 | Ambiguous | 2.09 | Destabilizing | 0.90 | Ambiguous | 0.581 | Likely Pathogenic | -5.93 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.30 | Pathogenic | 0.04 | Affected | 0.3102 | 0.2566 | -3 | -2 | 4.1 | -99.14 | |||||||||||||||||||||||||||||
| c.1751T>C | I584T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I584T is not reported in ClinVar and is absent from gnomAD. Consensus from most in silico predictors indicates a pathogenic effect: REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus score (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) all classify it as likely pathogenic. Only AlphaMissense‑Optimized predicts a benign outcome, while Rosetta and Foldetta are uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely pathogenic, and Foldetta as uncertain. With the overwhelming majority of tools supporting pathogenicity and no ClinVar entry to contradict this, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.059222 | Structured | 0.046673 | Uncertain | 0.846 | 0.244 | 0.000 | -10.413 | Likely Pathogenic | 0.765 | Likely Pathogenic | Likely Benign | 2.05 | Destabilizing | 0.1 | 1.70 | Ambiguous | 1.88 | Ambiguous | 1.66 | Destabilizing | 0.748 | Likely Pathogenic | -4.63 | Deleterious | 0.999 | Probably Damaging | 0.993 | Probably Damaging | -1.11 | Pathogenic | 0.02 | Affected | 0.0911 | 0.0608 | 0 | -1 | -5.2 | -12.05 | |||||||||||||||||||||||||||||
| c.1933T>G | F645V 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant F645V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. Tools that agree on a pathogenic effect are premPS, PROVEAN, SIFT, ESM1b, and AlphaMissense‑Default. The high‑accuracy assessments give an uncertain result from AlphaMissense‑Optimized, a pathogenic outcome from the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN), and an uncertain outcome from Foldetta (combining FoldX‑MD and Rosetta). No definitive folding‑stability change is reported. Overall, the majority of evidence points to a pathogenic impact for F645V, and this conclusion does not contradict any ClinVar annotation because the variant is not yet catalogued there. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.046336 | Structured | 0.276445 | Uncertain | 0.921 | 0.325 | 0.000 | -12.175 | Likely Pathogenic | 0.810 | Likely Pathogenic | Ambiguous | 1.98 | Ambiguous | 0.1 | 1.70 | Ambiguous | 1.84 | Ambiguous | 1.02 | Destabilizing | 0.316 | Likely Benign | -4.41 | Deleterious | 0.036 | Benign | 0.018 | Benign | 3.40 | Benign | 0.02 | Affected | 0.2163 | 0.2919 | -1 | -1 | 1.4 | -48.04 | |||||||||||||||||||||||||||||
| c.1957C>A | L653M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L653M has no ClinVar record and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, FATHMM, and AlphaMissense‑Optimized, while those that predict a pathogenic outcome are polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. Tools with uncertain or missing results (FoldX, Rosetta, Foldetta, premPS) are not considered evidence for either side. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive (two pathogenic vs. two benign votes), and Foldetta is also inconclusive. Overall, the majority of standard predictors lean toward pathogenicity, but the most reliable high‑accuracy tool indicates a benign effect, leaving the variant’s impact uncertain. No ClinVar entry exists, so there is no contradiction with clinical annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.049374 | Structured | 0.335213 | Uncertain | 0.963 | 0.332 | 0.000 | -8.838 | Likely Pathogenic | 0.713 | Likely Pathogenic | Likely Benign | 0.99 | Ambiguous | 0.2 | 1.70 | Ambiguous | 1.35 | Ambiguous | 0.94 | Ambiguous | 0.259 | Likely Benign | -1.76 | Neutral | 1.000 | Probably Damaging | 0.991 | Probably Damaging | 3.13 | Benign | 0.01 | Affected | 0.0935 | 0.3227 | 4 | 2 | -1.9 | 18.03 | ||||||||||||||||||||||||||||||
| c.880A>T | T294S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 T294S missense variant is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). In silico predictors that classify the variant as benign are absent; all available tools that provide a definitive call predict a deleterious effect. Pathogenic predictions come from REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, and AlphaMissense‑Default. Uncertain or inconclusive results are reported for FoldX, Rosetta, Foldetta, ESM1b, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as Uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta as Uncertain. Overall, the preponderance of pathogenic calls and the SGM Consensus support a pathogenic classification. Thus, the variant is most likely pathogenic, and this conclusion does not contradict the current ClinVar status, which contains no opposing evidence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.328603 | Structured | 0.316932 | Uncertain | 0.919 | 0.267 | 0.125 | -7.785 | In-Between | 0.954 | Likely Pathogenic | Ambiguous | 1.12 | Ambiguous | 0.2 | 1.70 | Ambiguous | 1.41 | Ambiguous | 1.19 | Destabilizing | 0.613 | Likely Pathogenic | -3.68 | Deleterious | 0.999 | Probably Damaging | 0.992 | Probably Damaging | 0.08 | Pathogenic | 0.03 | Affected | 0.2983 | 0.3396 | 1 | 1 | -0.1 | -14.03 | |||||||||||||||||||||||||||||
| c.881C>G | T294S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 T294S missense variant is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). In silico predictors that classify the variant as benign are absent; all available tools that provide a definitive call predict a deleterious effect. Pathogenic predictions come from REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, and AlphaMissense‑Default. Uncertain or inconclusive results are reported for FoldX, Rosetta, Foldetta, ESM1b, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as Uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta as Uncertain. Overall, the preponderance of pathogenic calls and the SGM Consensus support a pathogenic classification. Thus, the variant is most likely pathogenic, and this conclusion does not contradict the current ClinVar status, which contains no opposing evidence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.328603 | Structured | 0.316932 | Uncertain | 0.919 | 0.267 | 0.125 | -7.785 | In-Between | 0.954 | Likely Pathogenic | Ambiguous | 1.12 | Ambiguous | 0.2 | 1.70 | Ambiguous | 1.41 | Ambiguous | 1.19 | Destabilizing | 0.583 | Likely Pathogenic | -3.68 | Deleterious | 0.999 | Probably Damaging | 0.992 | Probably Damaging | 0.08 | Pathogenic | 0.03 | Affected | 0.2983 | 0.3396 | 1 | 1 | -0.1 | -14.03 | |||||||||||||||||||||||||||||
| c.1103C>A | P368Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P368Q missense variant has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, ESM1b, and AlphaMissense‑Optimized. Those that agree on a pathogenic effect are PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM. Remaining tools (AlphaMissense‑Default, FoldX, Rosetta, Foldetta, premPS) give uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the majority of definitive predictions (five pathogenic vs. three benign) indicate that the variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.363090 | Structured | 0.439989 | Uncertain | 0.580 | 0.677 | 0.250 | -6.019 | Likely Benign | 0.413 | Ambiguous | Likely Benign | 1.56 | Ambiguous | 0.8 | 1.71 | Ambiguous | 1.64 | Ambiguous | 0.71 | Ambiguous | 0.205 | Likely Benign | -5.10 | Deleterious | 0.991 | Probably Damaging | 0.881 | Possibly Damaging | 1.71 | Pathogenic | 0.04 | Affected | 0.1694 | 0.5283 | 0 | -1 | -1.9 | 31.01 | ||||||||||||||||||||||||||||||
| c.1808T>C | M603T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M603T is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that provide a clear verdict all classify the substitution as pathogenic: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). No tool in the dataset reports a benign outcome. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus indicates likely pathogenic, while the Foldetta stability analysis is inconclusive and therefore not considered evidence. No contradictory evidence is present in ClinVar. Consequently, the variant is most likely pathogenic based on the available predictions, with no conflict with ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.011342 | Structured | 0.197847 | Uncertain | 0.942 | 0.176 | 0.000 | -13.152 | Likely Pathogenic | 0.976 | Likely Pathogenic | Likely Pathogenic | 1.71 | Ambiguous | 0.2 | 1.71 | Ambiguous | 1.71 | Ambiguous | 0.66 | Ambiguous | 0.903 | Likely Pathogenic | -5.48 | Deleterious | 0.996 | Probably Damaging | 0.985 | Probably Damaging | -1.29 | Pathogenic | 0.00 | Affected | 0.1883 | 0.1495 | -1 | -1 | -2.6 | -30.09 | |||||||||||||||||||||||||||||
| c.1897C>G | L633V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L633V is not reported in ClinVar and is present in the gnomAD database (ID 6‑33440949‑C‑G). Prediction tools that indicate a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect comprise FoldX, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, SGM‑Consensus, and Foldetta; the Rosetta score is uncertain and therefore not considered. High‑accuracy methods give a pathogenic consensus: AlphaMissense‑Optimized predicts benign, but the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) and Foldetta both predict pathogenic. Overall, the majority of evidence supports a pathogenic impact for L633V, and this assessment does not contradict the ClinVar status, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.045352 | Structured | 0.045407 | Uncertain | 0.952 | 0.252 | 0.000 | 6-33440949-C-G | 1 | 6.20e-7 | -9.992 | Likely Pathogenic | 0.760 | Likely Pathogenic | Likely Benign | 2.32 | Destabilizing | 0.2 | 1.71 | Ambiguous | 2.02 | Destabilizing | 1.32 | Destabilizing | 0.327 | Likely Benign | -2.99 | Deleterious | 0.996 | Probably Damaging | 0.992 | Probably Damaging | 2.86 | Benign | 0.03 | Affected | 3.37 | 34 | 0.1517 | 0.2766 | 1 | 2 | 0.4 | -14.03 | ||||||||||||||||||||||||
| c.641T>A | L214Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense change at residue 214 (Leu→Gln) is not reported in ClinVar and is absent from gnomAD. Across the available in‑silico predictors, the majority (SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic effect, while only FATHMM predicts a benign outcome. Three tools (FoldX, Rosetta, Foldetta) return uncertain results. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta is uncertain. Taken together, the overwhelming consensus of pathogenic predictions indicates that the variant is most likely pathogenic, with no ClinVar evidence contradicting this assessment. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.155435 | Structured | 0.372604 | Uncertain | 0.818 | 0.302 | 0.125 | -10.119 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 1.60 | Ambiguous | 0.6 | 1.71 | Ambiguous | 1.66 | Ambiguous | 1.73 | Destabilizing | 0.912 | Likely Pathogenic | -5.12 | Deleterious | 0.997 | Probably Damaging | 0.936 | Probably Damaging | 5.74 | Benign | 0.00 | Affected | 0.1140 | 0.0930 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.952C>T | P318S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P318S is present in gnomAD (variant ID 6‑33437857‑C‑T) but has no ClinVar entry. Functional prediction tools uniformly indicate a deleterious effect. Pathogenic predictions come from SGM‑Consensus, REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Uncertain predictions come from Rosetta and Foldetta. No tool predicts a benign outcome. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized is pathogenic, SGM‑Consensus is likely pathogenic, and Foldetta remains uncertain. Taken together, the overwhelming majority of evidence supports a pathogenic classification, and this conclusion is consistent with the absence of a ClinVar record rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.111485 | Structured | 0.400936 | Uncertain | 0.858 | 0.234 | 0.000 | 6-33437857-C-T | 1 | 6.19e-7 | -9.954 | Likely Pathogenic | 0.956 | Likely Pathogenic | Likely Pathogenic | 2.22 | Destabilizing | 0.1 | 1.71 | Ambiguous | 1.97 | Ambiguous | 1.00 | Destabilizing | 0.626 | Likely Pathogenic | -7.05 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.87 | Pathogenic | 0.03 | Affected | 3.38 | 23 | 0.3692 | 0.5653 | -1 | 1 | 0.8 | -10.04 | ||||||||||||||||||||||||
| c.1079A>C | E360A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E360A is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only premPS. All other evaluated tools—SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default—predict a pathogenic impact. High‑accuracy methods give the following results: AlphaMissense‑Optimized is uncertain, SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates a likely pathogenic effect, and Foldetta (combining FoldX‑MD and Rosetta outputs) is uncertain. Because the majority of consensus tools predict pathogenicity and no ClinVar entry contradicts this, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.250310 | Structured | 0.421183 | Uncertain | 0.955 | 0.498 | 0.250 | -13.229 | Likely Pathogenic | 0.939 | Likely Pathogenic | Ambiguous | 1.37 | Ambiguous | 0.1 | 1.72 | Ambiguous | 1.55 | Ambiguous | 0.39 | Likely Benign | 0.545 | Likely Pathogenic | -5.52 | Deleterious | 0.997 | Probably Damaging | 0.980 | Probably Damaging | 1.63 | Pathogenic | 0.01 | Affected | 0.4295 | 0.8243 | 0 | -1 | 5.3 | -58.04 | |||||||||||||||||||||||||||||
| c.1099C>G | L367V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L367V is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). Only FATHMM predicts a pathogenic outcome. Stability‑based methods (FoldX, Rosetta, Foldetta) are inconclusive, so they provide no evidence for or against pathogenicity. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus as likely benign, and Foldetta as uncertain. Overall, the preponderance of evidence points to a benign effect for L367V, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.370445 | Structured | 0.441805 | Uncertain | 0.790 | 0.657 | 0.250 | -2.383 | Likely Benign | 0.066 | Likely Benign | Likely Benign | 1.48 | Ambiguous | 0.2 | 1.72 | Ambiguous | 1.60 | Ambiguous | 0.15 | Likely Benign | 0.040 | Likely Benign | 0.19 | Neutral | 0.410 | Benign | 0.104 | Benign | 1.67 | Pathogenic | 0.13 | Tolerated | 0.2321 | 0.3285 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.1108G>C | G370R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G370R is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and AlphaMissense‑Optimized. Those that agree on a pathogenic effect are SGM‑Consensus, FoldX, ESM1b, FATHMM, AlphaMissense‑Default, and Foldetta; Rosetta is uncertain. High‑accuracy assessments show AlphaMissense‑Optimized predicts benign, while the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenic. Overall, the balance of evidence leans toward pathogenicity, with two of the three high‑accuracy tools supporting this view. The variant is most likely pathogenic, and this assessment does not contradict any ClinVar status, as no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.461924 | Structured | 0.434325 | Uncertain | 0.359 | 0.720 | 0.500 | -8.375 | Likely Pathogenic | 0.731 | Likely Pathogenic | Likely Benign | 3.62 | Destabilizing | 3.7 | 1.72 | Ambiguous | 2.67 | Destabilizing | 0.22 | Likely Benign | 0.373 | Likely Benign | -0.80 | Neutral | 0.016 | Benign | 0.002 | Benign | 1.32 | Pathogenic | 0.55 | Tolerated | 0.0978 | 0.4313 | -3 | -2 | -4.1 | 99.14 | |||||||||||||||||||||||||||||
| c.1237C>T | P413S 2D  3DClick to see structure in 3D Viewer AISynGAP1 P413S is not reported in ClinVar and is absent from gnomAD. Prediction tools that classify the variant as benign include only FATHMM. All other evaluated predictors—SGM‑Consensus, REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and Foldetta—indicate a pathogenic effect. High‑accuracy methods give a consistent pathogenic verdict: AlphaMissense‑Optimized predicts pathogenicity, SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also predicts pathogenic, and Foldetta (which integrates FoldX‑MD and Rosetta outputs) predicts pathogenic; FoldX alone supports pathogenicity while Rosetta remains uncertain. Taken together, the overwhelming majority of predictions support a pathogenic classification, and this is not contradicted by the absence of a ClinVar assertion. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.113710 | Structured | 0.332472 | Uncertain | 0.927 | 0.201 | 0.000 | -12.586 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 3.06 | Destabilizing | 0.1 | 1.72 | Ambiguous | 2.39 | Destabilizing | 0.93 | Ambiguous | 0.509 | Likely Pathogenic | -7.37 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 3.19 | Benign | 0.04 | Affected | 0.3454 | 0.3819 | 1 | -1 | 0.8 | -10.04 | |||||||||||||||||||||||||||||
| c.714A>C | E238D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E238D missense variant has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include only FATHMM, whereas the remaining tools (REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, AlphaMissense‑Default, AlphaMissense‑Optimized) uniformly predict a pathogenic impact; the remaining methods (FoldX, Rosetta, Foldetta, premPS, ESM1b) are uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points to a pathogenic effect. The variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar status because no ClinVar record exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.194234 | Structured | 0.332638 | Uncertain | 0.796 | 0.326 | 0.000 | -7.861 | In-Between | 0.995 | Likely Pathogenic | Likely Pathogenic | 1.25 | Ambiguous | 0.4 | 1.72 | Ambiguous | 1.49 | Ambiguous | 0.78 | Ambiguous | 0.691 | Likely Pathogenic | -2.72 | Deleterious | 0.868 | Possibly Damaging | 0.504 | Possibly Damaging | 5.57 | Benign | 0.05 | Affected | 0.1975 | 0.3619 | 3 | 2 | 0.0 | -14.03 | ||||||||||||||||||||||||||||||
| c.714A>T | E238D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E238D missense variant has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include only FATHMM, whereas the remaining tools (REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, AlphaMissense‑Default, AlphaMissense‑Optimized) uniformly predict a pathogenic impact; the remaining methods (FoldX, Rosetta, Foldetta, premPS, ESM1b) are uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points to a pathogenic effect. The variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar status because no ClinVar record exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.194234 | Structured | 0.332638 | Uncertain | 0.796 | 0.326 | 0.000 | -7.861 | In-Between | 0.995 | Likely Pathogenic | Likely Pathogenic | 1.25 | Ambiguous | 0.4 | 1.72 | Ambiguous | 1.49 | Ambiguous | 0.78 | Ambiguous | 0.691 | Likely Pathogenic | -2.72 | Deleterious | 0.868 | Possibly Damaging | 0.504 | Possibly Damaging | 5.57 | Benign | 0.05 | Affected | 0.1975 | 0.3619 | 3 | 2 | 0.0 | -14.03 | ||||||||||||||||||||||||||||||
| c.1240A>G | M414V 2D  AISynGAP1 M414V is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools show a split: benign calls come from REVEL, SIFT, FATHMM, and AlphaMissense‑Optimized; pathogenic calls come from PROVEAN, polyPhen‑2 (HumDiv and HumVar), and ESM1b; the remaining tools (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Default) are inconclusive. The SGM consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, yields a pathogenic majority. High‑accuracy assessments give AlphaMissense‑Optimized benign, SGM consensus pathogenic, and Foldetta uncertain. Because the high‑accuracy predictions are divided and the overall tool set is evenly split, there is no definitive evidence for pathogenicity or benignity. Thus, the variant is most likely inconclusive, and this lack of consensus does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.081712 | Structured | 0.329108 | Uncertain | 0.914 | 0.217 | 0.000 | Uncertain | 1 | -8.003 | Likely Pathogenic | 0.541 | Ambiguous | Likely Benign | 1.81 | Ambiguous | 0.4 | 1.73 | Ambiguous | 1.77 | Ambiguous | 0.95 | Ambiguous | 0.261 | Likely Benign | -2.95 | Deleterious | 0.999 | Probably Damaging | 0.987 | Probably Damaging | 3.43 | Benign | 0.24 | Tolerated | 0.2585 | 0.3482 | 2 | 1 | 2.3 | -32.06 | ||||||||||||||||||||||||||||
| c.1639T>A | C547S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C547S is not reported in ClinVar (ClinVar status: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a pathogenic effect include SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. No tools predict a benign outcome. Predictions that are uncertain or inconclusive are FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the preponderance of evidence points to a pathogenic effect. This conclusion is consistent with the lack of ClinVar annotation and does not contradict any existing database status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.045352 | Structured | 0.007912 | Uncertain | 0.971 | 0.275 | 0.000 | -11.888 | Likely Pathogenic | 0.946 | Likely Pathogenic | Ambiguous | 0.85 | Ambiguous | 0.0 | 1.73 | Ambiguous | 1.29 | Ambiguous | 1.76 | Destabilizing | 0.884 | Likely Pathogenic | -9.64 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.27 | Pathogenic | 0.04 | Affected | 0.4542 | 0.1792 | 0 | -1 | -3.3 | -16.06 | |||||||||||||||||||||||||||||
| c.1640G>C | C547S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C547S is not reported in ClinVar (ClinVar status: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a pathogenic effect include SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. No tools predict a benign outcome. Predictions that are uncertain or inconclusive are FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the preponderance of evidence points to a pathogenic effect. This conclusion is consistent with the lack of ClinVar annotation and does not contradict any existing database status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.045352 | Structured | 0.007912 | Uncertain | 0.971 | 0.275 | 0.000 | -11.888 | Likely Pathogenic | 0.946 | Likely Pathogenic | Ambiguous | 0.85 | Ambiguous | 0.0 | 1.73 | Ambiguous | 1.29 | Ambiguous | 1.76 | Destabilizing | 0.825 | Likely Pathogenic | -9.64 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.27 | Pathogenic | 0.04 | Affected | 0.4542 | 0.1792 | 0 | -1 | -3.3 | -16.06 | |||||||||||||||||||||||||||||
| c.2171C>G | A724G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A724G is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, SIFT, and AlphaMissense‑Optimized. Those that predict a pathogenic effect comprise SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, and AlphaMissense‑Default. Predictions from FoldX, Rosetta, Foldetta, and premPS are uncertain and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely pathogenic, and Foldetta as uncertain. Based on the overall distribution of predictions, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.476583 | Structured | 0.458050 | Uncertain | 0.923 | 0.483 | 0.250 | -8.908 | Likely Pathogenic | 0.580 | Likely Pathogenic | Likely Benign | 1.45 | Ambiguous | 0.1 | 1.73 | Ambiguous | 1.59 | Ambiguous | 0.56 | Ambiguous | 0.286 | Likely Benign | -3.10 | Deleterious | 0.999 | Probably Damaging | 0.995 | Probably Damaging | 2.07 | Pathogenic | 0.08 | Tolerated | 0.2001 | 0.3609 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.2186A>C | N729T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N729T is not reported in ClinVar (ClinVar status: not listed) but is present in gnomAD (gnomAD ID 6‑33441651‑A‑C). Consensus among the majority of in‑silico predictors is benign: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the change as benign. No tool predicts pathogenicity. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized predicts benign; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates “Likely Benign”; Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, yields an uncertain result. Overall, the evidence strongly supports a benign effect, and this conclusion does not contradict ClinVar status, which has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | 0.750527 | Disordered | 0.426547 | Uncertain | 0.651 | 0.583 | 0.625 | 6-33441651-A-C | 1 | 6.20e-7 | -1.952 | Likely Benign | 0.103 | Likely Benign | Likely Benign | 0.52 | Ambiguous | 0.3 | 1.73 | Ambiguous | 1.13 | Ambiguous | -0.34 | Likely Benign | 0.052 | Likely Benign | -0.52 | Neutral | 0.123 | Benign | 0.042 | Benign | 3.33 | Benign | 1.00 | Tolerated | 3.59 | 7 | 0.1201 | 0.4805 | 0 | 0 | 2.8 | -13.00 | |||||||||||||||||||||||||
| c.1576G>C | V526L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V526L is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include premPS and SIFT, whereas the majority of algorithms—SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized—classify the change as pathogenic. FoldX, Rosetta, and Foldetta provide uncertain or inconclusive results and are treated as unavailable evidence. High‑accuracy assessments reinforce the pathogenic prediction: AlphaMissense‑Optimized is pathogenic, and the SGM Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates pathogenicity; Foldetta remains uncertain. Overall, the preponderance of evidence points to a pathogenic effect for V526L, and this conclusion does not contradict the absence of a ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.139895 | Structured | 0.023118 | Uncertain | 0.943 | 0.403 | 0.000 | -9.289 | Likely Pathogenic | 0.982 | Likely Pathogenic | Likely Pathogenic | 1.31 | Ambiguous | 0.5 | 1.74 | Ambiguous | 1.53 | Ambiguous | 0.38 | Likely Benign | 0.643 | Likely Pathogenic | -2.97 | Deleterious | 0.929 | Possibly Damaging | 0.917 | Probably Damaging | -1.16 | Pathogenic | 0.17 | Tolerated | 0.0754 | 0.3894 | 2 | 1 | -0.4 | 14.03 | |||||||||||||||||||||||||||||
| c.1576G>T | V526L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V526L is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include premPS and SIFT, whereas the majority of algorithms—SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized—classify the change as pathogenic. FoldX, Rosetta, and Foldetta provide uncertain or inconclusive results and are treated as unavailable evidence. High‑accuracy assessments reinforce the pathogenic prediction: AlphaMissense‑Optimized is pathogenic, and the SGM Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates pathogenicity; Foldetta remains uncertain. Overall, the preponderance of evidence points to a pathogenic effect for V526L, and this conclusion does not contradict the absence of a ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.139895 | Structured | 0.023118 | Uncertain | 0.943 | 0.403 | 0.000 | -9.289 | Likely Pathogenic | 0.982 | Likely Pathogenic | Likely Pathogenic | 1.31 | Ambiguous | 0.5 | 1.74 | Ambiguous | 1.53 | Ambiguous | 0.38 | Likely Benign | 0.643 | Likely Pathogenic | -2.97 | Deleterious | 0.929 | Possibly Damaging | 0.917 | Probably Damaging | -1.16 | Pathogenic | 0.17 | Tolerated | 0.0754 | 0.3894 | 2 | 1 | -0.4 | 14.03 | |||||||||||||||||||||||||||||
| c.1670C>G | S557C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S557C is not reported in ClinVar and is absent from gnomAD. Prediction tools that indicate a benign effect include premPS and AlphaMissense‑Optimized, whereas the majority of other in silico predictors (REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default) classify the change as pathogenic. FoldX, Rosetta, and Foldetta provide uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized predicting benign, while the SGM‑Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—predicts pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta outputs, remains uncertain. Overall, the preponderance of evidence from multiple pathogenic predictors and the SGM‑Consensus suggests the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation because the variant is not yet reported there. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.028107 | Structured | 0.010261 | Uncertain | 0.924 | 0.215 | 0.000 | -9.845 | Likely Pathogenic | 0.577 | Likely Pathogenic | Likely Benign | 1.43 | Ambiguous | 0.1 | 1.74 | Ambiguous | 1.59 | Ambiguous | 0.49 | Likely Benign | 0.923 | Likely Pathogenic | -4.52 | Deleterious | 0.999 | Probably Damaging | 0.993 | Probably Damaging | -1.77 | Pathogenic | 0.00 | Affected | 0.1287 | 0.5678 | 0 | -1 | 3.3 | 16.06 | |||||||||||||||||||||||||||||
| c.1772C>G | A591G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A591G is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote). Only PROVEAN predicts a pathogenic outcome. Predictions that are inconclusive or unavailable are FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus as Likely Benign, and Foldetta as uncertain. Overall, the majority of evidence points to a benign impact for A591G, and this conclusion does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.018787 | Structured | 0.093848 | Uncertain | 0.882 | 0.185 | 0.000 | -6.596 | Likely Benign | 0.233 | Likely Benign | Likely Benign | 1.22 | Ambiguous | 0.2 | 1.74 | Ambiguous | 1.48 | Ambiguous | 0.83 | Ambiguous | 0.142 | Likely Benign | -2.77 | Deleterious | 0.007 | Benign | 0.009 | Benign | 3.60 | Benign | 0.16 | Tolerated | 0.2117 | 0.2228 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.662A>G | E221G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E221G missense variant is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include polyPhen‑2 HumVar and FATHMM, while the majority of other in silico predictors (REVEL, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus) indicate a pathogenic impact; FoldX, Rosetta, Foldetta, and premPS are inconclusive. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Based on the collective evidence, the variant is most likely pathogenic, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.127496 | Structured | 0.413334 | Uncertain | 0.891 | 0.283 | 0.000 | Uncertain | 1 | -12.221 | Likely Pathogenic | 0.992 | Likely Pathogenic | Likely Pathogenic | 1.40 | Ambiguous | 0.1 | 1.74 | Ambiguous | 1.57 | Ambiguous | 0.71 | Ambiguous | 0.863 | Likely Pathogenic | -5.56 | Deleterious | 0.596 | Possibly Damaging | 0.201 | Benign | 5.79 | Benign | 0.00 | Affected | 0.2611 | 0.6370 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||
| c.754C>G | P252A 2D  3DClick to see structure in 3D Viewer AISynGAP1 P252A is not reported in ClinVar and is absent from gnomAD. Prediction tools cluster into two groups: a single benign call from FATHMM, and a majority of pathogenic calls from REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Uncertain results are reported by FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as pathogenic, and Foldetta as inconclusive. Overall, the preponderance of evidence points to a pathogenic effect for P252A, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.461924 | Structured | 0.211606 | Uncertain | 0.753 | 0.304 | 0.250 | -10.587 | Likely Pathogenic | 0.977 | Likely Pathogenic | Likely Pathogenic | 0.97 | Ambiguous | 0.1 | 1.74 | Ambiguous | 1.36 | Ambiguous | 0.69 | Ambiguous | 0.831 | Likely Pathogenic | -7.35 | Deleterious | 0.941 | Possibly Damaging | 0.607 | Possibly Damaging | 5.87 | Benign | 0.03 | Affected | 0.3490 | 0.4161 | 1 | -1 | 3.4 | -26.04 | |||||||||||||||||||||||||||||
| c.821T>A | L274Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L274Q is reported in ClinVar with an uncertain significance (ClinVar ID 1810279.0) and is not found in gnomAD. Functional prediction tools uniformly indicate a deleterious effect: REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the change as pathogenic, while Rosetta remains inconclusive. No tool predicts a benign outcome. High‑accuracy assessments corroborate this trend: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, is pathogenic. Consequently, the variant is most likely pathogenic, and this assessment does not contradict the current ClinVar status of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.066181 | Structured | 0.377483 | Uncertain | 0.866 | 0.195 | 0.250 | Uncertain | 1 | -15.518 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 2.54 | Destabilizing | 0.3 | 1.74 | Ambiguous | 2.14 | Destabilizing | 1.97 | Destabilizing | 0.774 | Likely Pathogenic | -5.42 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 0.00 | Pathogenic | 0.00 | Affected | 3.38 | 19 | 0.1128 | 0.0688 | -2 | -2 | -7.3 | 14.97 | 245.9 | 1.8 | 0.0 | 0.0 | 0.1 | 0.2 | X | X | X | Potentially Pathogenic | The aliphatic side chain of Leu274, located in a β hairpin loop (res. Glu273-Lys278) connecting two anti-parallel β sheet strands, packs against multiple hydrophobic residues facing the β sheet (e.g., Ala271, Leu327, Tyr280, Val306). The hydrophilic carboxamide group of the Gln274 side chain is not suitable for this hydrophobic niche, causing nearby residues to adjust to make room for the hydrophilic glutamine. Additionally, a new hydrogen bond forms with the backbone carboxyl group of Arg272 in another β strand (res. Glu273-Arg259).As a result, the backbone amide group of Ala399 and the carbonyl group of Arg272, which connect two β strands at the β sheet end, form fewer hydrogen bonds in the variant than in the WT simulations. Although no major secondary structure disruption is observed in the variant simulations, the residue swap could profoundly affect the C2 domain folding, as the hydrophobic packing of Leu274 is crucial for maintaining the loop's contact with the rest of the C2 domain. Lastly, because the Leu274-containing loop faces the membrane surface, the residue swap could also negatively impact the SynGAP-membrane association. | ||||||||||||||
| c.1013A>G | D338G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D338G missense variant is not reported in ClinVar and has no gnomAD entry. Prediction tools that agree on a benign effect include REVEL, premPS, and polyPhen‑2 HumVar. Those that predict a pathogenic effect comprise SGM‑Consensus (Likely Pathogenic), PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. Because the majority of available predictors (seven versus three) indicate a deleterious impact, the variant is most likely pathogenic, and this assessment does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.335645 | Structured | 0.363354 | Uncertain | 0.460 | 0.438 | 0.375 | -8.875 | Likely Pathogenic | 0.871 | Likely Pathogenic | Ambiguous | 1.33 | Ambiguous | 0.5 | 1.75 | Ambiguous | 1.54 | Ambiguous | 0.15 | Likely Benign | 0.487 | Likely Benign | -5.51 | Deleterious | 0.771 | Possibly Damaging | 0.315 | Benign | 1.69 | Pathogenic | 0.01 | Affected | 0.4014 | 0.5934 | 1 | -1 | 3.1 | -58.04 | |||||||||||||||||||||||||||||
| c.1139G>T | G380V 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant G380V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show mixed results: benign predictions come from premPS, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized, while pathogenic predictions are reported by REVEL, FoldX, polyPhen‑2 HumDiv, and SIFT; Rosetta is uncertain. The SGM‑Consensus, derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, indicates a likely benign effect. High‑accuracy assessments further support this: AlphaMissense‑Optimized predicts benign, the SGM‑Consensus (majority of the four high‑accuracy tools) is benign, whereas Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, predicts a pathogenic impact. Overall, the majority of evidence points to a benign effect, and this does not contradict the lack of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.724957 | Disordered | 0.432982 | Uncertain | 0.316 | 0.939 | 0.750 | -6.234 | Likely Benign | 0.172 | Likely Benign | Likely Benign | 7.60 | Destabilizing | 2.5 | 1.75 | Ambiguous | 4.68 | Destabilizing | -0.17 | Likely Benign | 0.574 | Likely Pathogenic | -0.70 | Neutral | 0.816 | Possibly Damaging | 0.210 | Benign | 2.54 | Benign | 0.02 | Affected | 0.1618 | 0.3708 | -1 | -3 | 4.6 | 42.08 | |||||||||||||||||||||||||||||
| c.1232T>C | I411T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 I411T missense variant is not reported in ClinVar (status: None) and has no entries in gnomAD. Prediction tools that agree on a benign effect are limited to FATHMM, while the majority of tools predict a pathogenic impact: SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Rosetta’s assessment is uncertain. High‑accuracy methods give consistent pathogenic predictions: AlphaMissense‑Optimized is Pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) is Pathogenic. No prediction or stability result is missing or inconclusive. Based on the collective evidence, the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.116183 | Structured | 0.339366 | Uncertain | 0.927 | 0.198 | 0.000 | -11.258 | Likely Pathogenic | 0.992 | Likely Pathogenic | Likely Pathogenic | 2.86 | Destabilizing | 0.0 | 1.75 | Ambiguous | 2.31 | Destabilizing | 1.86 | Destabilizing | 0.579 | Likely Pathogenic | -4.54 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.31 | Benign | 0.00 | Affected | 0.0903 | 0.1389 | 0 | -1 | -5.2 | -12.05 | |||||||||||||||||||||||||||||
| c.1580A>C | D527A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D527A missense variant is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only premPS, while the remaining evaluated methods (SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized) uniformly predict pathogenicity. FoldX, Rosetta, and Foldetta are inconclusive and are treated as unavailable. High‑accuracy assessments further support a deleterious outcome: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta remains uncertain. Overall, the variant is most likely pathogenic based on the consensus of predictive tools, and this assessment does not contradict any ClinVar status, as none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.139895 | Structured | 0.021908 | Uncertain | 0.913 | 0.408 | 0.000 | -15.473 | Likely Pathogenic | 0.970 | Likely Pathogenic | Likely Pathogenic | 0.81 | Ambiguous | 0.9 | 1.75 | Ambiguous | 1.28 | Ambiguous | -0.24 | Likely Benign | 0.929 | Likely Pathogenic | -7.79 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | -2.39 | Pathogenic | 0.00 | Affected | 0.2850 | 0.3799 | 0 | -2 | 5.3 | -44.01 | |||||||||||||||||||||||||||||
| c.1624A>G | N542D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N542D is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only SIFT, whereas the majority of tools (REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus) predict a pathogenic impact; Rosetta and Foldetta are uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as likely pathogenic, and Foldetta as uncertain. Taken together, the preponderance of evidence indicates that the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status, as no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.053060 | Structured | 0.026143 | Uncertain | 0.953 | 0.331 | 0.000 | -13.269 | Likely Pathogenic | 0.987 | Likely Pathogenic | Likely Pathogenic | 2.13 | Destabilizing | 0.3 | 1.75 | Ambiguous | 1.94 | Ambiguous | 1.05 | Destabilizing | 0.796 | Likely Pathogenic | -4.51 | Deleterious | 1.000 | Probably Damaging | 0.997 | Probably Damaging | -1.40 | Pathogenic | 0.08 | Tolerated | 0.1664 | 0.3176 | 2 | 1 | 0.0 | 0.98 | |||||||||||||||||||||||||||||
| c.1418T>C | V473A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 V473A missense variant is not reported in ClinVar and has no entry in gnomAD. Prediction tools that agree on a benign effect are REVEL and FATHMM. Tools that agree on a pathogenic effect include SGM‑Consensus (Likely Pathogenic), premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, and AlphaMissense‑Default. FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized are all uncertain or unavailable, providing no decisive evidence. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. Overall, the majority of available predictions support a pathogenic impact. This conclusion is consistent with the lack of ClinVar annotation (no contradiction). Thus, the variant is most likely pathogenic based on current computational evidence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.191378 | Structured | 0.362529 | Uncertain | 0.884 | 0.239 | 0.000 | -10.867 | Likely Pathogenic | 0.925 | Likely Pathogenic | Ambiguous | 1.88 | Ambiguous | 0.0 | 1.76 | Ambiguous | 1.82 | Ambiguous | 2.17 | Destabilizing | 0.485 | Likely Benign | -3.95 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.18 | Benign | 0.00 | Affected | 0.2529 | 0.2521 | 0 | 0 | -2.4 | -28.05 | |||||||||||||||||||||||||||||
| c.1487A>G | E496G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E496G missense variant is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that assess the variant’s effect fall into two groups: no tool predicts a benign outcome, while eight tools (REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default) all predict a pathogenic effect. The SGM‑Consensus, which is a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also indicates a likely pathogenic outcome. High‑accuracy assessments are mixed: AlphaMissense‑Optimized is uncertain, the SGM‑Consensus remains likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is uncertain. Overall, the preponderance of evidence points to a pathogenic effect, contradicting the current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.102787 | Structured | 0.383296 | Uncertain | 0.945 | 0.179 | 0.000 | Uncertain | 1 | -13.529 | Likely Pathogenic | 0.850 | Likely Pathogenic | Ambiguous | 1.83 | Ambiguous | 0.1 | 1.76 | Ambiguous | 1.80 | Ambiguous | 0.92 | Ambiguous | 0.825 | Likely Pathogenic | -6.16 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | -1.45 | Pathogenic | 0.02 | Affected | 3.37 | 35 | 0.2435 | 0.3473 | 0 | -2 | 3.1 | -72.06 | 173.9 | 103.1 | 0.0 | 0.0 | -0.7 | 0.0 | X | X | Potentially Pathogenic | Glu496 is located in the α-helix (res. Leu489-Glu519), and its carboxylate group forms salt bridges with the neighbouring residues Lys492 and Arg499 in the WT simulations. Glu496 also forms a hydrogen bond with Ser449 on an opposing helix (res. Val441-Ser457). In the variant simulations, Gly496 cannot form these salt bridges, which could weaken the secondary structure. Additionally, the loss of the hydrogen bond with Ser449 on the opposite helix can weaken the tertiary structure assembly. Moreover, glycine is an α-helix breaker, and it is seen to weaken the integrity of the helix as the hydrogen bonding between the backbone atoms of Gly496 and Ala493 breaks down. Also, due to its location at the GAP-Ras interface, the interaction of Glu496 with Arg499 and Lys492 might play a role in complex association and stability, which cannot be fully addressed using the SynGAP solvent-only simulations. | |||||||||||||||
| c.2050G>A | D684N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant D684N is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that indicate a benign effect include REVEL, premPS, and FATHMM, whereas the majority of tools predict a pathogenic outcome: PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized classifies the variant as pathogenic, the SGM‑Consensus also reports it as likely pathogenic, and the Foldetta stability analysis is inconclusive. Protein‑stability predictors FoldX and Rosetta likewise return uncertain results. Overall, the preponderance of evidence points to a pathogenic effect, which contradicts the current ClinVar designation of uncertainty. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.254060 | Structured | 0.153798 | Uncertain | 0.870 | 0.282 | 0.000 | Uncertain | 1 | -13.155 | Likely Pathogenic | 0.985 | Likely Pathogenic | Likely Pathogenic | 1.47 | Ambiguous | 0.8 | 1.76 | Ambiguous | 1.62 | Ambiguous | 0.37 | Likely Benign | 0.382 | Likely Benign | -4.99 | Deleterious | 0.999 | Probably Damaging | 0.746 | Possibly Damaging | 3.39 | Benign | 0.01 | Affected | 0.1157 | 0.6373 | 2 | 1 | 0.0 | -0.98 | |||||||||||||||||||||||||||
| c.643G>T | G215C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G215C is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are premPS and FATHMM; those that agree on a pathogenic effect include SGM‑Consensus, REVEL, FoldX, Foldetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Rosetta gives an uncertain result and is listed separately. High‑accuracy methods all predict pathogenicity: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, is pathogenic. Based on the overwhelming agreement among both general and high‑accuracy predictors, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.155435 | Structured | 0.382818 | Uncertain | 0.791 | 0.291 | 0.000 | -12.295 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 2.39 | Destabilizing | 0.2 | 1.76 | Ambiguous | 2.08 | Destabilizing | 0.33 | Likely Benign | 0.869 | Likely Pathogenic | -7.69 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 5.56 | Benign | 0.00 | Affected | 0.1274 | 0.4640 | -3 | -3 | 2.9 | 46.09 | |||||||||||||||||||||||||||||
| c.772C>T | R258C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 R258C missense variant is listed in ClinVar with an “Uncertain” status and is present in gnomAD (variant ID 6‑33437677‑C‑T). Prediction tools that agree on a benign effect include only FATHMM. All other evaluated predictors—REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN)—indicate a pathogenic or likely pathogenic impact. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM‑Consensus as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Based on the preponderance of pathogenic predictions, the variant is most likely pathogenic, which does not contradict its current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.295083 | Structured | 0.293667 | Uncertain | 0.894 | 0.260 | 0.250 | Uncertain | 1 | 6-33437677-C-T | 1 | 6.20e-7 | -10.285 | Likely Pathogenic | 0.790 | Likely Pathogenic | Ambiguous | 1.17 | Ambiguous | 0.4 | 1.76 | Ambiguous | 1.47 | Ambiguous | 0.87 | Ambiguous | 0.771 | Likely Pathogenic | -6.79 | Deleterious | 1.000 | Probably Damaging | 0.993 | Probably Damaging | 5.77 | Benign | 0.00 | Affected | 3.39 | 15 | 0.3307 | 0.3411 | -3 | -4 | 7.0 | -53.05 | ||||||||||||||||||||||
| c.1196C>G | A399G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A399G is not reported in ClinVar and is absent from gnomAD. Across a panel of in silico predictors, all available pathogenicity scores classify the substitution as benign: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool predicts pathogenicity. Uncertain results from FoldX, Rosetta, Foldetta, and premPS are treated as unavailable. High‑accuracy assessments reinforce the benign prediction: AlphaMissense‑Optimized is benign; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also indicates benign. Foldetta, which integrates FoldX‑MD and Rosetta outputs, is inconclusive and therefore not considered evidence. Consequently, the variant is most likely benign, and this assessment does not contradict any ClinVar annotation (none present). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.394753 | Structured | 0.407674 | Uncertain | 0.939 | 0.490 | 0.125 | -6.513 | Likely Benign | 0.215 | Likely Benign | Likely Benign | 1.03 | Ambiguous | 0.1 | 1.77 | Ambiguous | 1.40 | Ambiguous | 0.74 | Ambiguous | 0.153 | Likely Benign | -1.59 | Neutral | 0.062 | Benign | 0.024 | Benign | 5.39 | Benign | 0.13 | Tolerated | 0.2174 | 0.4532 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.1486G>A | E496K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E496K is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only FoldX, whereas the majority of tools (REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus) predict a pathogenic impact. Uncertain predictions come from Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as likely pathogenic, and Foldetta as inconclusive. Overall, the evidence strongly favors a pathogenic classification for E496K, and this conclusion does not contradict the ClinVar status, which currently contains no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.102787 | Structured | 0.383296 | Uncertain | 0.945 | 0.179 | 0.000 | -15.795 | Likely Pathogenic | 0.961 | Likely Pathogenic | Likely Pathogenic | 0.38 | Likely Benign | 0.1 | 1.77 | Ambiguous | 1.08 | Ambiguous | 0.76 | Ambiguous | 0.743 | Likely Pathogenic | -3.58 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | -1.40 | Pathogenic | 0.04 | Affected | 0.1810 | 0.3528 | 0 | 1 | -0.4 | -0.94 | |||||||||||||||||||||||||||||
| c.1703T>C | V568A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 V568A missense variant is not reported in ClinVar (ClinVar status: none) but is present in the gnomAD database (gnomAD ID: 6‑33440755‑T‑C). Prediction tools that agree on a benign effect include only SIFT, whereas the majority of tools predict a pathogenic impact: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus (Likely Pathogenic). Uncertain or inconclusive results come from FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM Consensus (derived from the unanimous pathogenic vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta as uncertain. Based on the preponderance of pathogenic predictions and the consensus from high‑accuracy tools, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status (which has no entry). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.024826 | Structured | 0.053503 | Uncertain | 0.937 | 0.257 | 0.000 | 6-33440755-T-C | 2 | 1.25e-6 | -10.929 | Likely Pathogenic | 0.946 | Likely Pathogenic | Ambiguous | 1.90 | Ambiguous | 0.1 | 1.77 | Ambiguous | 1.84 | Ambiguous | 2.16 | Destabilizing | 0.834 | Likely Pathogenic | -3.82 | Deleterious | 0.999 | Probably Damaging | 0.990 | Probably Damaging | -1.38 | Pathogenic | 0.06 | Tolerated | 3.37 | 35 | 0.2413 | 0.1961 | 0 | 0 | -2.4 | -28.05 | ||||||||||||||||||||||||
| c.1192C>T | P398S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P398S missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include polyPhen‑2 (HumVar), ESM1b, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are REVEL, PROVEAN, polyPhen‑2 (HumDiv), and SIFT. The remaining tools (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Default) give uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as benign, and Foldetta as uncertain. Overall, the majority of definitive predictions lean toward a benign impact, and this conclusion does not contradict the lack of ClinVar annotation. Thus, the variant is most likely benign based on current computational evidence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.436924 | Structured | 0.401041 | Uncertain | 0.891 | 0.525 | 0.250 | -6.757 | Likely Benign | 0.490 | Ambiguous | Likely Benign | 1.86 | Ambiguous | 0.3 | 1.78 | Ambiguous | 1.82 | Ambiguous | 0.85 | Ambiguous | 0.544 | Likely Pathogenic | -5.58 | Deleterious | 0.478 | Possibly Damaging | 0.130 | Benign | 5.68 | Benign | 0.03 | Affected | 0.3619 | 0.5737 | 1 | -1 | 0.8 | -10.04 | ||||||||||||||||||||||||||||||
| c.1246C>G | L416V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L416V is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, SIFT, AlphaMissense‑Default, AlphaMissense‑Optimized, and FATHMM. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv and polyPhen‑2 HumVar. Predictions that are inconclusive or unavailable are FoldX, Rosetta, Foldetta, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Benign, and Foldetta as uncertain. Overall, the majority of evidence points to a benign impact for the variant. This conclusion is not contradicted by any ClinVar annotation, as the variant is not present in that database. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.104810 | Structured | 0.336105 | Uncertain | 0.935 | 0.227 | 0.000 | -7.861 | In-Between | 0.310 | Likely Benign | Likely Benign | 1.46 | Ambiguous | 0.0 | 1.78 | Ambiguous | 1.62 | Ambiguous | 0.45 | Likely Benign | 0.124 | Likely Benign | -1.47 | Neutral | 0.995 | Probably Damaging | 0.970 | Probably Damaging | 3.42 | Benign | 0.53 | Tolerated | 0.1563 | 0.3550 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.1313C>G | A438G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A438G is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote). Only PROVEAN predicts a pathogenic outcome. Predictions that are inconclusive or unavailable are FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus as Likely Benign, and Foldetta as uncertain. Overall, the majority of evidence points to a benign impact for A438G, and this conclusion is consistent with the lack of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.147574 | Structured | 0.290154 | Uncertain | 0.929 | 0.293 | 0.000 | -5.790 | Likely Benign | 0.182 | Likely Benign | Likely Benign | 1.08 | Ambiguous | 0.1 | 1.78 | Ambiguous | 1.43 | Ambiguous | 0.80 | Ambiguous | 0.034 | Likely Benign | -2.51 | Deleterious | 0.247 | Benign | 0.037 | Benign | 4.12 | Benign | 0.11 | Tolerated | 0.1788 | 0.2719 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.1488G>C | E496D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E496D missense variant is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only FoldX, whereas the majority of tools predict a pathogenic impact: SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. Uncertain predictions come from Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (derived from a unanimous majority of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the preponderance of evidence points to a pathogenic effect. This conclusion is consistent with the absence of ClinVar annotation, so there is no contradiction with existing clinical data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.102787 | Structured | 0.383296 | Uncertain | 0.945 | 0.179 | 0.000 | -10.552 | Likely Pathogenic | 0.922 | Likely Pathogenic | Ambiguous | 0.43 | Likely Benign | 0.2 | 1.78 | Ambiguous | 1.11 | Ambiguous | 1.18 | Destabilizing | 0.583 | Likely Pathogenic | -2.78 | Deleterious | 0.996 | Probably Damaging | 0.989 | Probably Damaging | -1.45 | Pathogenic | 0.04 | Affected | 3.37 | 35 | 0.1576 | 0.1912 | 2 | 3 | 0.0 | -14.03 | |||||||||||||||||||||||||||
| c.1488G>T | E496D 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant E496D is reported in gnomAD (ID 6‑33438520‑G‑T) but has no ClinVar entry. Functional prediction tools cluster into two groups: benign predictions are provided only by FoldX, whereas the remaining tools—REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus—classify the change as pathogenic. Uncertain results come from Rosetta, Foldetta, and AlphaMissense‑Optimized and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Overall, the preponderance of evidence points to a pathogenic effect, and this conclusion does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.102787 | Structured | 0.383296 | Uncertain | 0.945 | 0.179 | 0.000 | 6-33438520-G-T | 2 | 1.24e-6 | -10.552 | Likely Pathogenic | 0.922 | Likely Pathogenic | Ambiguous | 0.43 | Likely Benign | 0.2 | 1.78 | Ambiguous | 1.11 | Ambiguous | 1.18 | Destabilizing | 0.583 | Likely Pathogenic | -2.78 | Deleterious | 0.996 | Probably Damaging | 0.989 | Probably Damaging | -1.45 | Pathogenic | 0.04 | Affected | 3.37 | 35 | 0.1576 | 0.1912 | 2 | 3 | 0.0 | -14.03 | ||||||||||||||||||||||||
| c.1499T>A | L500Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L500Q has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include only AlphaMissense‑Optimized. All other evaluated predictors—SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default—consistently classify the variant as pathogenic. FoldX, Rosetta, and Foldetta provide uncertain or inconclusive stability results and are treated as unavailable. High‑accuracy assessments further clarify the picture: AlphaMissense‑Optimized (benign) stands alone, whereas the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, predicts pathogenic. Foldetta remains uncertain. Overall, the preponderance of evidence from multiple independent pathogenic predictors indicates that the variant is most likely pathogenic, and this conclusion does not contradict ClinVar status, which currently has no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.066181 | Structured | 0.382942 | Uncertain | 0.893 | 0.150 | 0.000 | -13.431 | Likely Pathogenic | 0.649 | Likely Pathogenic | Likely Benign | 1.92 | Ambiguous | 0.1 | 1.78 | Ambiguous | 1.85 | Ambiguous | 1.49 | Destabilizing | 0.878 | Likely Pathogenic | -5.97 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.35 | Pathogenic | 0.00 | Affected | 0.0950 | 0.0488 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.1878T>G | I626M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I626M is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. Tools that agree on a pathogenic effect include SGM‑Consensus, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. FoldX, Rosetta, Foldetta, and premPS give uncertain results and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized predicts benign, while the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts likely pathogenic; Foldetta’s stability prediction is uncertain. Overall, the majority of evidence points to a pathogenic impact. This conclusion does not contradict ClinVar status, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.109221 | Structured | 0.040732 | Uncertain | 0.970 | 0.223 | 0.000 | -11.407 | Likely Pathogenic | 0.618 | Likely Pathogenic | Likely Benign | 0.72 | Ambiguous | 0.1 | 1.78 | Ambiguous | 1.25 | Ambiguous | 0.92 | Ambiguous | 0.405 | Likely Benign | -2.76 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.06 | Benign | 0.02 | Affected | 0.0661 | 0.1959 | 2 | 1 | -2.6 | 18.03 | |||||||||||||||||||||||||||||
| c.1700A>C | E567A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E567A is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, FoldX, SIFT, and FATHMM, while pathogenic predictions arise from SGM‑Consensus, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and AlphaMissense‑Default. Four tools (Foldetta, premPS, Rosetta, AlphaMissense‑Optimized) yield uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as Likely Pathogenic (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN), and Foldetta as uncertain. Overall, the majority of evidence points toward a pathogenic effect. This conclusion is not contradicted by ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.021816 | Structured | 0.051008 | Uncertain | 0.916 | 0.234 | 0.000 | -12.854 | Likely Pathogenic | 0.902 | Likely Pathogenic | Ambiguous | 0.43 | Likely Benign | 0.1 | 1.79 | Ambiguous | 1.11 | Ambiguous | 0.63 | Ambiguous | 0.418 | Likely Benign | -5.74 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.43 | Benign | 0.17 | Tolerated | 0.3170 | 0.5189 | 0 | -1 | 5.3 | -58.04 | |||||||||||||||||||||||||||||
| c.1761G>C | R587S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R587S is not reported in ClinVar and is absent from gnomAD. Prediction tools cluster into two groups: the single benign prediction comes from SIFT, while the remaining tools—REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and the SGM Consensus—indicate pathogenicity. High‑accuracy assessments further support a deleterious effect: the SGM Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is “Likely Pathogenic”; AlphaMissense‑Optimized is “Uncertain”; and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is also “Uncertain.” Taken together, the preponderance of evidence points to a pathogenic impact for R587S. This conclusion is not contradicted by ClinVar status, which currently has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.054297 | Structured | 0.077330 | Uncertain | 0.862 | 0.216 | 0.000 | -12.264 | Likely Pathogenic | 0.830 | Likely Pathogenic | Ambiguous | 0.84 | Ambiguous | 0.1 | 1.79 | Ambiguous | 1.32 | Ambiguous | 1.17 | Destabilizing | 0.508 | Likely Pathogenic | -4.84 | Deleterious | 0.990 | Probably Damaging | 0.779 | Possibly Damaging | -1.20 | Pathogenic | 0.09 | Tolerated | 3.37 | 35 | 0.2852 | 0.4165 | -1 | 0 | 3.7 | -69.11 | |||||||||||||||||||||||||||
| c.1761G>T | R587S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 R587S missense variant is catalogued in gnomAD (ID 6‑33440813‑G‑T) but has no ClinVar entry. Functional prediction tools largely agree on a deleterious effect: pathogenic calls come from REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, and AlphaMissense‑Default, while only SIFT predicts a benign outcome. Uncertain results are reported by FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments reinforce the pathogenic signal: AlphaMissense‑Optimized remains uncertain, but the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—classifies the variant as pathogenic, and Foldetta likewise yields an uncertain stability change. Overall, the preponderance of evidence indicates that R587S is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.054297 | Structured | 0.077330 | Uncertain | 0.862 | 0.216 | 0.000 | 6-33440813-G-T | 4 | 2.48e-6 | -12.264 | Likely Pathogenic | 0.830 | Likely Pathogenic | Ambiguous | 0.84 | Ambiguous | 0.1 | 1.79 | Ambiguous | 1.32 | Ambiguous | 1.17 | Destabilizing | 0.508 | Likely Pathogenic | -4.84 | Deleterious | 0.990 | Probably Damaging | 0.779 | Possibly Damaging | -1.20 | Pathogenic | 0.09 | Tolerated | 3.37 | 35 | 0.2852 | 0.4165 | -1 | 0 | 3.7 | -69.11 | ||||||||||||||||||||||||
| c.1948A>C | N650H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N650H lies in the GAP domain. ClinVar has no entry for this variant, and it is not reported in gnomAD. Prediction tools that agree on a benign effect are REVEL and FATHMM. Tools that predict a pathogenic effect include PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, FoldX, and the SGM‑Consensus (majority vote). Uncertain or inconclusive results come from Rosetta, Foldetta, and premPS. High‑accuracy assessments: AlphaMissense‑Optimized predicts pathogenicity; the SGM‑Consensus also indicates likely pathogenic; Foldetta, which integrates FoldX‑MD and Rosetta outputs, is inconclusive. Based on the preponderance of pathogenic predictions and the high‑accuracy tools’ results, the variant is most likely pathogenic. This conclusion does not contradict ClinVar status, as no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.086953 | Structured | 0.361944 | Uncertain | 0.961 | 0.357 | 0.000 | -14.187 | Likely Pathogenic | 0.986 | Likely Pathogenic | Likely Pathogenic | 2.14 | Destabilizing | 0.3 | 1.79 | Ambiguous | 1.97 | Ambiguous | 0.55 | Ambiguous | 0.465 | Likely Benign | -4.98 | Deleterious | 0.999 | Probably Damaging | 0.929 | Probably Damaging | 3.01 | Benign | 0.05 | Affected | 0.1990 | 0.4784 | 2 | 1 | 0.3 | 23.04 | |||||||||||||||||||||||||||||
| c.737T>A | L246Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 L246Q missense variant is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Prediction tools that agree on a benign effect are limited to FATHMM, while the remaining tools—SGM‑Consensus, REVEL, FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—consistently predict a pathogenic impact; Rosetta is uncertain and is not grouped. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. Taken together, the overwhelming majority of evidence indicates that the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status, as none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.472492 | Structured | 0.302312 | Uncertain | 0.859 | 0.364 | 0.000 | -15.420 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 2.82 | Destabilizing | 0.3 | 1.79 | Ambiguous | 2.31 | Destabilizing | 1.69 | Destabilizing | 0.921 | Likely Pathogenic | -5.43 | Deleterious | 0.997 | Probably Damaging | 0.916 | Probably Damaging | 4.67 | Benign | 0.00 | Affected | 0.1092 | 0.0758 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.1066C>G | R356G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 R356G missense variant is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are REVEL and SIFT, while the majority of tools predict a pathogenic outcome: SGM‑Consensus, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, and AlphaMissense‑Default. FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized are uncertain and therefore treated as unavailable evidence. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as Likely Pathogenic (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN), and Foldetta as uncertain. Overall, the preponderance of pathogenic predictions indicates that the variant is most likely pathogenic, and this conclusion does not contradict the current ClinVar status, which contains no classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.219301 | Structured | 0.395028 | Uncertain | 0.802 | 0.373 | 0.250 | -12.305 | Likely Pathogenic | 0.883 | Likely Pathogenic | Ambiguous | 1.02 | Ambiguous | 0.0 | 1.80 | Ambiguous | 1.41 | Ambiguous | 1.06 | Destabilizing | 0.271 | Likely Benign | -6.20 | Deleterious | 0.993 | Probably Damaging | 0.982 | Probably Damaging | 1.95 | Pathogenic | 0.08 | Tolerated | 0.3505 | 0.3793 | -3 | -2 | 4.1 | -99.14 | |||||||||||||||||||||||||||||
| c.1343C>T | A448V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A448V is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include REVEL and FATHMM, whereas a majority of tools predict a pathogenic impact: FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Pathogenic). Two tools (Rosetta and premPS) yield uncertain results. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. Overall, the preponderance of evidence indicates that A448V is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.225814 | Structured | 0.292774 | Uncertain | 0.973 | 0.257 | 0.000 | -10.372 | Likely Pathogenic | 0.992 | Likely Pathogenic | Likely Pathogenic | 3.31 | Destabilizing | 0.2 | 1.80 | Ambiguous | 2.56 | Destabilizing | 0.66 | Ambiguous | 0.487 | Likely Benign | -3.95 | Deleterious | 0.998 | Probably Damaging | 0.955 | Probably Damaging | 3.26 | Benign | 0.02 | Affected | 0.0950 | 0.4955 | 0 | 0 | 2.4 | 28.05 | |||||||||||||||||||||||||||||
| c.1762C>A | L588I 2D  AISynGAP1 missense variant L588I has no ClinVar entry and is not reported in gnomAD. Prediction tools that classify the variant as benign include PROVEAN. Those that predict pathogenicity are SGM‑Consensus, REVEL, FoldX, Foldetta, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. Uncertain results come from Rosetta, premPS, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as pathogenic. Overall, the majority of evidence supports a pathogenic effect, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.038042 | Structured | 0.082229 | Uncertain | 0.887 | 0.214 | 0.000 | -12.454 | Likely Pathogenic | 0.874 | Likely Pathogenic | Ambiguous | 2.54 | Destabilizing | 1.2 | 1.80 | Ambiguous | 2.17 | Destabilizing | 0.88 | Ambiguous | 0.607 | Likely Pathogenic | -1.99 | Neutral | 0.999 | Probably Damaging | 0.997 | Probably Damaging | -1.27 | Pathogenic | 0.04 | Affected | 0.0949 | 0.2433 | 2 | 2 | 0.7 | 0.00 | |||||||||||||||||||||||||||||
| c.1816A>C | S606R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S606R is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, FoldX, SIFT, and FATHMM, whereas a separate group predicts pathogenicity: PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Default. Tools with uncertain outcomes are Foldetta, premPS, and Rosetta. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is Likely Pathogenic, and Foldetta remains inconclusive. Overall, the majority of high‑confidence predictions and the consensus vote indicate a pathogenic effect. Because ClinVar contains no entry for this variant, there is no contradiction between the computational evidence and clinical annotation. Thus, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.041405 | Structured | 0.191720 | Uncertain | 0.875 | 0.247 | 0.000 | -12.900 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 0.15 | Likely Benign | 0.4 | 1.80 | Ambiguous | 0.98 | Ambiguous | 0.82 | Ambiguous | 0.252 | Likely Benign | -4.98 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | 3.38 | Benign | 0.08 | Tolerated | 0.0819 | 0.2750 | 0 | -1 | -3.7 | 69.11 | |||||||||||||||||||||||||||||
| c.1818T>A | S606R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S606R is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, FoldX, SIFT, and FATHMM, whereas a separate group predicts pathogenicity: PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Default. Tools with uncertain outcomes are Foldetta, premPS, and Rosetta. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is Likely Pathogenic, and Foldetta remains inconclusive. Overall, the majority of high‑confidence predictions and the consensus vote indicate a pathogenic effect. Because ClinVar contains no entry for this variant, there is no contradiction between the computational evidence and clinical annotation. Thus, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.041405 | Structured | 0.191720 | Uncertain | 0.875 | 0.247 | 0.000 | -12.900 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 0.15 | Likely Benign | 0.4 | 1.80 | Ambiguous | 0.98 | Ambiguous | 0.82 | Ambiguous | 0.246 | Likely Benign | -4.98 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | 3.38 | Benign | 0.08 | Tolerated | 0.0819 | 0.2750 | 0 | -1 | -3.7 | 69.11 | |||||||||||||||||||||||||||||
| c.1818T>G | S606R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S606R is not reported in ClinVar or gnomAD. Prediction tools that indicate a benign effect include REVEL, FoldX, SIFT, and FATHMM, whereas a majority predict pathogenicity: PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is labeled Likely Pathogenic, while Foldetta, which integrates FoldX‑MD and Rosetta outputs, is uncertain. High‑accuracy methods confirm the pathogenic trend: AlphaMissense‑Optimized is pathogenic, SGM Consensus is Likely Pathogenic, and Foldetta remains uncertain. Overall, the evidence points to the variant being most likely pathogenic, and this assessment does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.041405 | Structured | 0.191720 | Uncertain | 0.875 | 0.247 | 0.000 | -12.900 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 0.15 | Likely Benign | 0.4 | 1.80 | Ambiguous | 0.98 | Ambiguous | 0.82 | Ambiguous | 0.246 | Likely Benign | -4.98 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | 3.38 | Benign | 0.08 | Tolerated | 0.0819 | 0.2750 | 0 | -1 | -3.7 | 69.11 | |||||||||||||||||||||||||||||
| c.1999A>T | I667F 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 I667F missense variant is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Prediction tools largely agree on a deleterious effect: benign calls are limited to FATHMM, while the remaining 11 predictors—SGM‑Consensus, REVEL, FoldX, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and Foldetta—classify the change as pathogenic. High‑accuracy methods reinforce this view: AlphaMissense‑Optimized predicts pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also predicts pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) reports a pathogenic effect. No prediction is inconclusive. Consequently, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.142424 | Structured | 0.083597 | Uncertain | 0.927 | 0.379 | 0.000 | -14.389 | Likely Pathogenic | 0.963 | Likely Pathogenic | Likely Pathogenic | 7.55 | Destabilizing | 0.3 | 1.80 | Ambiguous | 4.68 | Destabilizing | 0.86 | Ambiguous | 0.569 | Likely Pathogenic | -3.98 | Deleterious | 0.998 | Probably Damaging | 0.790 | Possibly Damaging | 2.96 | Benign | 0.00 | Affected | 0.0668 | 0.2648 | 1 | 0 | -1.7 | 34.02 | |||||||||||||||||||||||||||||
| c.2055G>C | L685F 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L685F is not reported in ClinVar and has no gnomAD entry. Functional prediction tools cluster into two groups: benign predictions come from REVEL, premPS, and FATHMM, whereas pathogenic predictions arise from SGM‑Consensus, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized classifies the change as pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates likely pathogenic, while Foldetta’s stability analysis is inconclusive. FoldX and Rosetta predictions are uncertain and are treated as unavailable. Overall, the preponderance of evidence points to a pathogenic impact for the variant, and this conclusion does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.175930 | Structured | 0.162061 | Uncertain | 0.913 | 0.280 | 0.000 | -12.304 | Likely Pathogenic | 0.992 | Likely Pathogenic | Likely Pathogenic | 1.62 | Ambiguous | 0.2 | 1.80 | Ambiguous | 1.71 | Ambiguous | 0.50 | Likely Benign | 0.300 | Likely Benign | -3.99 | Deleterious | 0.999 | Probably Damaging | 0.895 | Possibly Damaging | 3.33 | Benign | 0.01 | Affected | 0.0724 | 0.2367 | 2 | 0 | -1.0 | 34.02 | |||||||||||||||||||||||||||||
| c.2055G>T | L685F 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L685F is not reported in ClinVar and has no gnomAD entry. Functional prediction tools cluster into two groups: benign predictions come from REVEL, premPS, and FATHMM, whereas pathogenic predictions arise from SGM‑Consensus, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized classifies the change as pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates likely pathogenic, while Foldetta’s stability analysis is inconclusive. FoldX and Rosetta predictions are uncertain and are treated as unavailable. Overall, the preponderance of evidence points to a pathogenic impact for the variant, and this conclusion does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.175930 | Structured | 0.162061 | Uncertain | 0.913 | 0.280 | 0.000 | -12.304 | Likely Pathogenic | 0.992 | Likely Pathogenic | Likely Pathogenic | 1.62 | Ambiguous | 0.2 | 1.80 | Ambiguous | 1.71 | Ambiguous | 0.50 | Likely Benign | 0.300 | Likely Benign | -3.99 | Deleterious | 0.999 | Probably Damaging | 0.895 | Possibly Damaging | 3.33 | Benign | 0.01 | Affected | 0.0724 | 0.2367 | 2 | 0 | -1.0 | 34.02 | |||||||||||||||||||||||||||||
| c.857T>G | L286R 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L286R is not reported in ClinVar and is absent from gnomAD. Functional prediction tools uniformly indicate a deleterious effect: REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the change as pathogenic, while Rosetta remains uncertain. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also reports a likely pathogenic outcome. High‑accuracy assessments corroborate this: AlphaMissense‑Optimized predicts pathogenicity; the SGM‑Consensus (majority vote) is likely pathogenic; and Foldetta, which integrates FoldX‑MD (pathogenic) and Rosetta (uncertain), reports a pathogenic effect. Consequently, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation, as none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.122885 | Structured | 0.385647 | Uncertain | 0.932 | 0.260 | 0.000 | -15.563 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 2.90 | Destabilizing | 0.3 | 1.80 | Ambiguous | 2.35 | Destabilizing | 1.97 | Destabilizing | 0.868 | Likely Pathogenic | -5.52 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.49 | Pathogenic | 0.00 | Affected | 0.1370 | 0.0600 | -3 | -2 | -8.3 | 43.03 | |||||||||||||||||||||||||||||
| c.1351C>G | L451V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L451V is not reported in ClinVar (ClinVar status: none) and is absent from gnomAD (gnomAD ID: none). Prediction tools that agree on a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. In contrast, the majority of tools predict a pathogenic impact: SGM‑Consensus (Likely Pathogenic), FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. Rosetta is uncertain and does not contribute to the consensus. High‑accuracy methods give the following results: AlphaMissense‑Optimized predicts benign; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts likely pathogenic; and Foldetta predicts pathogenic. Overall, the preponderance of evidence points to a pathogenic effect for L451V. This conclusion is not contradicted by ClinVar, which has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.281712 | Structured | 0.314017 | Uncertain | 0.978 | 0.232 | 0.000 | -10.732 | Likely Pathogenic | 0.635 | Likely Pathogenic | Likely Benign | 2.83 | Destabilizing | 0.1 | 1.81 | Ambiguous | 2.32 | Destabilizing | 1.39 | Destabilizing | 0.325 | Likely Benign | -2.90 | Deleterious | 0.995 | Probably Damaging | 0.970 | Probably Damaging | 2.54 | Benign | 0.01 | Affected | 0.1065 | 0.2856 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.1502T>C | I501T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I501T is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict a pathogenic outcome are FoldX, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and premPS, while Rosetta remains uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as likely benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as pathogenic. Overall, the majority of predictions lean toward a benign effect, and this does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.079919 | Structured | 0.366596 | Uncertain | 0.886 | 0.153 | 0.000 | Uncertain | 1 | -5.996 | Likely Benign | 0.252 | Likely Benign | Likely Benign | 2.40 | Destabilizing | 0.1 | 1.81 | Ambiguous | 2.11 | Destabilizing | 1.57 | Destabilizing | 0.362 | Likely Benign | -3.48 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.44 | Benign | 0.16 | Tolerated | 3.37 | 35 | 0.0972 | 0.0640 | 0 | -1 | -5.2 | -12.05 | 214.5 | 26.9 | 0.0 | 0.0 | 0.5 | 0.0 | X | Potentially Pathogenic | Ile501 is located near a hinge in the middle of an α-helix (res. Leu489-Glu519). The sec-butyl side chain of Ile501 is hydrophobically packed with other residues in the inter-helix space (e.g., Leu500, Tyr497, Phe679) in the WT simulations. In the variant simulations, the hydroxyl group of Thr501 forms a hydrogen bond with the backbone atoms of Tyr497 on the same α-helix, which may weaken the α-helix integrity. Additionally, the polar hydroxyl group of Thr501 is not suitable for the hydrophobic inter-helix space, and thus, the residue swap could affect protein folding. However, Ile501 is followed by Gly502, which facilitates a hinge in the middle of the α-helix, making further weakening caused by Thr501 unlikely to be harmful to the α-helix integrity. | ||||||||||||||||
| c.1727G>T | C576F 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C576F is not reported in ClinVar (ClinVar ID = None) and has no entries in gnomAD (gnomAD ID = None). Prediction tools that agree on a benign effect are limited to FATHMM, which scores the variant as benign. All other evaluated algorithms—SGM‑Consensus, REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and Foldetta—consistently predict a pathogenic or likely pathogenic impact. Uncertain predictions from Rosetta and premPS are treated as unavailable. High‑accuracy methods reinforce this view: AlphaMissense‑Optimized reports pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates likely pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenic. Based on the overwhelming agreement among high‑confidence tools, the variant is most likely pathogenic, and this assessment does not contradict the absence of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.113710 | Structured | 0.017684 | Uncertain | 0.913 | 0.245 | 0.000 | -13.467 | Likely Pathogenic | 0.994 | Likely Pathogenic | Likely Pathogenic | 5.04 | Destabilizing | 0.5 | 1.81 | Ambiguous | 3.43 | Destabilizing | 0.58 | Ambiguous | 0.516 | Likely Pathogenic | -9.93 | Deleterious | 0.999 | Probably Damaging | 0.996 | Probably Damaging | 3.40 | Benign | 0.02 | Affected | 0.1626 | 0.3527 | -4 | -2 | 0.3 | 44.04 | |||||||||||||||||||||||||||||
| c.2074C>A | L692M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L692M is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, PROVEAN, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are premPS, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive (2 vs 2) and Foldetta (combining FoldX‑MD and Rosetta outputs) is also inconclusive. Thus, the overall evidence slightly favors pathogenicity, with a majority of standard tools predicting a deleterious impact. The variant is most likely pathogenic based on current predictions, and this assessment does not contradict the ClinVar status, which has no entry for this change. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.064632 | Structured | 0.295225 | Uncertain | 0.966 | 0.243 | 0.000 | -9.659 | Likely Pathogenic | 0.746 | Likely Pathogenic | Likely Benign | 0.49 | Likely Benign | 0.0 | 1.81 | Ambiguous | 1.15 | Ambiguous | 1.01 | Destabilizing | 0.302 | Likely Benign | -1.99 | Neutral | 1.000 | Probably Damaging | 0.996 | Probably Damaging | 3.07 | Benign | 0.01 | Affected | 0.0754 | 0.2585 | 4 | 2 | -1.9 | 18.03 | ||||||||||||||||||||||||||||||
| c.754C>T | P252S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P252S is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only FATHMM, while the majority of tools (SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic impact. Predictions that are uncertain (FoldX, Rosetta, Foldetta, premPS) are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta as uncertain. Overall, the evidence strongly favors a pathogenic effect for P252S, and this conclusion does not contradict the ClinVar status, which is currently unreported. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.461924 | Structured | 0.211606 | Uncertain | 0.753 | 0.304 | 0.250 | -10.926 | Likely Pathogenic | 0.996 | Likely Pathogenic | Likely Pathogenic | 0.76 | Ambiguous | 0.1 | 1.81 | Ambiguous | 1.29 | Ambiguous | 0.79 | Ambiguous | 0.840 | Likely Pathogenic | -7.35 | Deleterious | 0.941 | Possibly Damaging | 0.531 | Possibly Damaging | 5.79 | Benign | 0.05 | Affected | 0.3508 | 0.4361 | 1 | -1 | 0.8 | -10.04 | |||||||||||||||||||||||||||||
| c.1649C>G | A550G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 A550G missense variant is not reported in ClinVar (ClinVar ID None) and has no entries in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only AlphaMissense‑Optimized. Tools that agree on a pathogenic effect are SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. FoldX, Rosetta, Foldetta, and premPS are inconclusive and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized predicting benign, while the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta outputs, is uncertain. Overall, the majority of reliable predictors indicate a pathogenic effect. This conclusion does not contradict ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.018106 | Structured | 0.007241 | Uncertain | 0.954 | 0.265 | 0.000 | -15.086 | Likely Pathogenic | 0.776 | Likely Pathogenic | Likely Benign | 1.73 | Ambiguous | 0.0 | 1.82 | Ambiguous | 1.78 | Ambiguous | 0.85 | Ambiguous | 0.769 | Likely Pathogenic | -3.79 | Deleterious | 0.999 | Probably Damaging | 0.932 | Probably Damaging | -1.31 | Pathogenic | 0.01 | Affected | 0.1595 | 0.2758 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.967C>A | L323M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 L323M missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, PROVEAN, and AlphaMissense‑Optimized. Those that predict a pathogenic outcome are polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM. The remaining tools—Foldetta, premPS, AlphaMissense‑Default, ESM1b, and Rosetta—return uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive, and Foldetta is also inconclusive. Taken together, the evidence leans toward a benign impact for the variant, and this assessment does not contradict any ClinVar annotation, as none exists for this change. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.268042 | Structured | 0.428564 | Uncertain | 0.956 | 0.369 | 0.000 | -7.065 | In-Between | 0.427 | Ambiguous | Likely Benign | 0.11 | Likely Benign | 0.1 | 1.82 | Ambiguous | 0.97 | Ambiguous | 0.93 | Ambiguous | 0.270 | Likely Benign | -1.02 | Neutral | 0.997 | Probably Damaging | 0.939 | Probably Damaging | 0.64 | Pathogenic | 0.03 | Affected | 0.0637 | 0.3010 | 4 | 2 | -1.9 | 18.03 | ||||||||||||||||||||||||||||||
| c.2009T>C | L670P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L670P is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, premPS, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) all indicate benign. In contrast, polyPhen‑2 (HumDiv and HumVar) predict a pathogenic impact. Stability‑based methods are inconclusive: FoldX, Rosetta, and the combined Foldetta output are uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely benign, and Foldetta as uncertain. Overall, the majority of evidence supports a benign classification, and this is consistent with the absence of a ClinVar assertion; there is no contradiction with ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.161087 | Structured | 0.090855 | Uncertain | 0.812 | 0.385 | 0.000 | -4.916 | Likely Benign | 0.237 | Likely Benign | Likely Benign | 0.56 | Ambiguous | 0.3 | 1.83 | Ambiguous | 1.20 | Ambiguous | -0.25 | Likely Benign | 0.211 | Likely Benign | 2.42 | Neutral | 1.000 | Probably Damaging | 0.996 | Probably Damaging | 3.41 | Benign | 0.26 | Tolerated | 0.3590 | 0.1474 | -3 | -3 | -5.4 | -16.04 | |||||||||||||||||||||||||||||
| c.712G>A | E238K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E238K missense change is not reported in ClinVar and is absent from gnomAD. In silico predictors cluster into two groups: a single benign call from FATHMM, and a consensus of pathogenic predictions from REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Four tools (FoldX, Rosetta, Foldetta, premPS) returned uncertain results and are not considered evidence. High‑accuracy methods reinforce the pathogenic signal: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, while Foldetta remains uncertain. Overall, the preponderance of evidence points to a pathogenic effect for E238K. This conclusion is not contradicted by ClinVar, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.194234 | Structured | 0.332638 | Uncertain | 0.796 | 0.326 | 0.000 | -13.475 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 0.56 | Ambiguous | 0.4 | 1.83 | Ambiguous | 1.20 | Ambiguous | 0.83 | Ambiguous | 0.858 | Likely Pathogenic | -3.63 | Deleterious | 0.995 | Probably Damaging | 0.695 | Possibly Damaging | 5.46 | Benign | 0.01 | Affected | 4.29 | 391 | 0.2812 | 0.5524 | 0 | 1 | -0.4 | -0.94 | 209.0 | 55.9 | 0.0 | 0.0 | -0.1 | 0.0 | X | Potentially Pathogenic | The negatively charged residue Glu238, located in an α helix (res. Ala236-Val250), is replaced by the positively charged residue Lys238. This charge reversal removes the periodically formed salt bridge between the carboxylate group of Glu238 and the guanidinium group of Arg234 observed in the WT simulations. In the variant simulations, both Lys238 and Arg234 form alternative salt bridges with the carboxylate group of Glu680 in the GAP domain loop. Although not visible in the simulations, the absence of the Glu238-Arg234 salt bridge could weaken the integrity of the α helix (residues Ala236-Val250) and potentially affect the tertiary assembly between the PH and GAP domains. | ||||||||||||||||||
| c.875C>G | A292G 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant A292G is not reported in ClinVar and is absent from gnomAD. Prediction tools that indicate a benign effect include REVEL, SIFT, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect comprise premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, and AlphaMissense‑Default. The high‑accuracy consensus from AlphaMissense‑Optimized is benign, whereas the SGM Consensus—derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—classifies the variant as pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta outputs, is uncertain, and individual FoldX and Rosetta predictions are also inconclusive. Overall, the majority of evidence points to a pathogenic impact, and this assessment does not contradict any ClinVar annotation because no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.247041 | Structured | 0.362042 | Uncertain | 0.929 | 0.256 | 0.000 | -9.692 | Likely Pathogenic | 0.713 | Likely Pathogenic | Likely Benign | 0.88 | Ambiguous | 0.1 | 1.83 | Ambiguous | 1.36 | Ambiguous | 1.05 | Destabilizing | 0.323 | Likely Benign | -3.68 | Deleterious | 0.999 | Probably Damaging | 0.996 | Probably Damaging | 1.74 | Pathogenic | 0.10 | Tolerated | 0.2187 | 0.5054 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.931C>G | H311D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant H311D is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that assess evolutionary conservation and protein function uniformly indicate a deleterious effect: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the substitution as pathogenic. The majority‑vote consensus (SGM‑Consensus) also reports it as likely pathogenic. Tools that evaluate structural stability give inconclusive results: FoldX, Rosetta, Foldetta, and premPS are listed as uncertain. High‑accuracy assessments further support a damaging outcome: AlphaMissense‑Optimized predicts pathogenic, SGM‑Consensus (derived from the same set of predictors) reports likely pathogenic, while Foldetta’s combined FoldX‑MD/Rosetta stability analysis remains uncertain. Overall, the preponderance of evidence from multiple independent predictors points to a pathogenic effect, which is consistent with the absence of a benign ClinVar annotation and the lack of population frequency data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.229226 | Structured | 0.354792 | Uncertain | 0.902 | 0.314 | 0.125 | -13.513 | Likely Pathogenic | 0.976 | Likely Pathogenic | Likely Pathogenic | 1.27 | Ambiguous | 0.0 | 1.83 | Ambiguous | 1.55 | Ambiguous | 0.89 | Ambiguous | 0.633 | Likely Pathogenic | -6.94 | Deleterious | 0.997 | Probably Damaging | 0.994 | Probably Damaging | 1.94 | Pathogenic | 0.02 | Affected | 0.2301 | 0.2008 | 1 | -1 | -0.3 | -22.05 | |||||||||||||||||||||||||||||
| c.1523A>C | D508A 2D  3DClick to see structure in 3D Viewer AISynGAP1 D508A is not reported in ClinVar and is absent from gnomAD. Prediction tools that classify the variant as benign include REVEL, FoldX, premPS, SIFT, FATHMM, and AlphaMissense‑Optimized. Those that predict pathogenicity are SGM Consensus, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Default. The high‑accuracy methods give a split result: AlphaMissense‑Optimized predicts benign, SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts pathogenic, and Foldetta (combining FoldX‑MD and Rosetta) is uncertain. No prediction or stability result is missing; all available outputs are considered. Overall, the predictions are evenly divided between benign and pathogenic, with no clear consensus. Therefore, the variant is inconclusive; it is not contradictory to the ClinVar status, which has no entry for this change. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.019401 | Structured | 0.255890 | Uncertain | 0.890 | 0.228 | 0.000 | -11.434 | Likely Pathogenic | 0.704 | Likely Pathogenic | Likely Benign | -0.18 | Likely Benign | 0.2 | 1.84 | Ambiguous | 0.83 | Ambiguous | 0.09 | Likely Benign | 0.339 | Likely Benign | -7.37 | Deleterious | 0.988 | Probably Damaging | 0.762 | Possibly Damaging | 3.37 | Benign | 0.06 | Tolerated | 0.3758 | 0.4241 | 0 | -2 | 5.3 | -44.01 | |||||||||||||||||||||||||||||
| c.1535A>G | E512G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E512G is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, and FATHMM, whereas a majority of tools (SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, and AlphaMissense‑Default) predict a pathogenic impact. The remaining tools—FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized—return uncertain or inconclusive results and are treated as unavailable evidence. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as Likely Pathogenic (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN), and Foldetta as uncertain. Overall, the preponderance of evidence points to a pathogenic effect for E512G, and this conclusion does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.092881 | Structured | 0.247079 | Uncertain | 0.923 | 0.273 | 0.000 | -11.996 | Likely Pathogenic | 0.817 | Likely Pathogenic | Ambiguous | 0.86 | Ambiguous | 0.1 | 1.84 | Ambiguous | 1.35 | Ambiguous | 0.17 | Likely Benign | 0.381 | Likely Benign | -6.46 | Deleterious | 0.997 | Probably Damaging | 0.915 | Probably Damaging | 3.30 | Benign | 0.02 | Affected | 0.3262 | 0.4185 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.1867C>G | L623V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L623V is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only REVEL, whereas the majority of tools predict a pathogenic impact: SGM‑Consensus, FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. Rosetta and AlphaMissense‑Optimized give uncertain results. High‑accuracy assessments further support pathogenicity: the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic, Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic, and AlphaMissense‑Optimized remains uncertain. Overall, the preponderance of evidence indicates that the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.175930 | Structured | 0.060667 | Uncertain | 0.962 | 0.211 | 0.000 | -12.802 | Likely Pathogenic | 0.896 | Likely Pathogenic | Ambiguous | 3.96 | Destabilizing | 0.3 | 1.84 | Ambiguous | 2.90 | Destabilizing | 1.45 | Destabilizing | 0.416 | Likely Benign | -2.99 | Deleterious | 0.998 | Probably Damaging | 0.992 | Probably Damaging | 1.60 | Pathogenic | 0.01 | Affected | 0.1653 | 0.3588 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.1214G>A | R405H 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant R405H is listed in ClinVar with an uncertain significance (ClinVar ID 863440.0) and is present in gnomAD (variant ID 6‑33438119‑G‑A). Functional prediction tools that agree on a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. In contrast, the majority of tools predict a pathogenic impact: FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and the SGM‑Consensus score (Likely Pathogenic). High‑accuracy assessments further support a deleterious outcome: AlphaMissense‑Optimized reports a benign change, whereas the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) and Foldetta (combining FoldX‑MD and Rosetta) both predict pathogenicity. Overall, the preponderance of evidence indicates that R405H is most likely pathogenic, which does not contradict the current ClinVar status of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.250310 | Structured | 0.404888 | Uncertain | 0.949 | 0.315 | 0.000 | Conflicting | 2 | 6-33438119-G-A | 4 | 2.48e-6 | -9.081 | Likely Pathogenic | 0.706 | Likely Pathogenic | Likely Benign | 2.79 | Destabilizing | 0.6 | 1.85 | Ambiguous | 2.32 | Destabilizing | 1.26 | Destabilizing | 0.371 | Likely Benign | -4.54 | Deleterious | 1.000 | Probably Damaging | 0.991 | Probably Damaging | 3.65 | Benign | 0.01 | Affected | 3.38 | 28 | 0.3395 | 0.2363 | 2 | 0 | 1.3 | -19.05 | 214.0 | 102.2 | -0.1 | 0.0 | -0.7 | 0.1 | X | Potentially Pathogenic | The guanidinium group of Arg405, located in an anti-parallel β sheet strand of the C2 domain (res. Pro398-Ile411), forms a salt bridge with the carboxylate group of the Glu446 side chain from an opposing α helix (res. Val441-Ser457) in the GAP domain. The positively charged Arg405 side chain also stacks with the aromatic ring of the Phe358 side chain from a loop preceding the β strand (res. Thr359-Thr366), which could assist in maintaining the anti-parallel strand arrangement.In the variant simulations, the imidazole ring of His405 does not stack with the aromatic ring of Phe358 nor form any lasting H-bonds with the loop residues. The imidazole ring of His405 (neutral and epsilon protonated in the simulations) is unable to form a salt bridge with Glu446, which could affect the tertiary structure assembly, although this is not apparent based on the variant simulations. | |||||||||||||
| c.1238C>A | P413H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P413H is not reported in ClinVar (ClinVar status: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include only FATHMM, while the remaining tools—SGM‑Consensus, REVEL, FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—consistently predict a pathogenic or likely pathogenic impact; Rosetta is uncertain. High‑accuracy methods give the following results: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates likely pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. No predictions are missing or inconclusive. Based on the overwhelming agreement among pathogenic predictors and the high‑accuracy tools, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which is currently unreported. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.113710 | Structured | 0.332472 | Uncertain | 0.927 | 0.201 | 0.000 | -13.376 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 4.89 | Destabilizing | 1.0 | 1.85 | Ambiguous | 3.37 | Destabilizing | 1.07 | Destabilizing | 0.546 | Likely Pathogenic | -8.29 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 3.16 | Benign | 0.00 | Affected | 0.1745 | 0.3480 | 0 | -2 | -1.6 | 40.02 | |||||||||||||||||||||||||||||
| c.2086C>G | L696V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 L696V variant is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that indicate a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized, whereas the majority of other in silico predictors (FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) report a pathogenic outcome; Rosetta remains inconclusive. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized predicts benign, but the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—leans pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenic. Overall, the preponderance of evidence points to a pathogenic effect for the variant, which does not contradict the current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.200174 | Structured | 0.390093 | Uncertain | 0.962 | 0.267 | 0.000 | Uncertain | 1 | -11.909 | Likely Pathogenic | 0.745 | Likely Pathogenic | Likely Benign | 2.35 | Destabilizing | 0.1 | 1.85 | Ambiguous | 2.10 | Destabilizing | 1.46 | Destabilizing | 0.351 | Likely Benign | -2.79 | Deleterious | 0.992 | Probably Damaging | 0.970 | Probably Damaging | 3.16 | Benign | 0.00 | Affected | 3.46 | 13 | 0.1307 | 0.2830 | 1 | 2 | 0.4 | -14.03 | |||||||||||||||||||||||||
| c.598T>G | L200V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L200V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Benign). Tools that predict a pathogenic outcome are FoldX, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and the Foldetta stability assessment. Uncertain results come from Rosetta and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Benign, and Foldetta as pathogenic. Overall, the majority of evidence points to a benign effect; there is no ClinVar annotation to contradict this conclusion. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | PH | 0.366687 | Structured | 0.428168 | Uncertain | 0.687 | 0.453 | 0.125 | -6.917 | Likely Benign | 0.213 | Likely Benign | Likely Benign | 2.15 | Destabilizing | 0.2 | 1.85 | Ambiguous | 2.00 | Destabilizing | 0.92 | Ambiguous | 0.098 | Likely Benign | -1.07 | Neutral | 0.990 | Probably Damaging | 0.760 | Possibly Damaging | 4.09 | Benign | 0.24 | Tolerated | 0.1593 | 0.3607 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.626T>C | V209A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V209A is reported in gnomAD (ID 6‑33435268‑T‑C) but has no ClinVar entry. Prediction tools that agree on a benign effect include REVEL and FATHMM. In contrast, a majority of tools predict a pathogenic impact: premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Pathogenic). Stability‑based methods FoldX, Rosetta, and the combined Foldetta score are uncertain and therefore not considered evidence for or against pathogenicity. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. Overall, the preponderance of evidence from multiple independent predictors indicates that V209A is most likely pathogenic, and this conclusion does not contradict any ClinVar status because no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.247041 | Structured | 0.397624 | Uncertain | 0.874 | 0.331 | 0.125 | 6-33435268-T-C | 1 | 6.20e-7 | -9.796 | Likely Pathogenic | 0.976 | Likely Pathogenic | Likely Pathogenic | 1.56 | Ambiguous | 0.3 | 1.85 | Ambiguous | 1.71 | Ambiguous | 1.60 | Destabilizing | 0.236 | Likely Benign | -2.79 | Deleterious | 0.958 | Probably Damaging | 0.581 | Possibly Damaging | 3.70 | Benign | 0.02 | Affected | 3.41 | 13 | 0.2237 | 0.1919 | 0 | 0 | -2.4 | -28.05 | ||||||||||||||||||||||||
| c.602A>C | D201A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D201A variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, SIFT, and FATHMM, whereas a pathogenic consensus is reached by PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and AlphaMissense‑Default. Uncertain or unavailable results come from Rosetta, Foldetta, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized predicts a benign outcome, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) leans toward pathogenic, and Foldetta remains inconclusive. Overall, the majority of standard tools favor a benign classification, but the high‑accuracy consensus indicates a pathogenic signal, leaving the variant’s impact uncertain. The predictions do not contradict ClinVar status, as no ClinVar assertion exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.366687 | Structured | 0.428570 | Uncertain | 0.698 | 0.447 | 0.125 | -7.793 | In-Between | 0.769 | Likely Pathogenic | Likely Benign | 0.45 | Likely Benign | 0.1 | 1.86 | Ambiguous | 1.16 | Ambiguous | 0.23 | Likely Benign | 0.261 | Likely Benign | -3.81 | Deleterious | 0.989 | Probably Damaging | 0.828 | Possibly Damaging | 4.11 | Benign | 0.09 | Tolerated | 0.3177 | 0.5050 | 0 | -2 | 5.3 | -44.01 | ||||||||||||||||||||||||||||||
| c.779T>G | V260G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 V260G missense variant has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include FATHMM and AlphaMissense‑Optimized, while the majority of tools (REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) predict a pathogenic impact. FoldX, Rosetta, and Foldetta are uncertain and therefore not considered evidence. High‑accuracy assessments show AlphaMissense‑Optimized as benign, whereas the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—leans pathogenic (3 pathogenic vs. 1 benign). Foldetta remains uncertain. Overall, the preponderance of evidence points to a pathogenic effect for V260G, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.254060 | Structured | 0.382651 | Uncertain | 0.888 | 0.259 | 0.250 | -9.300 | Likely Pathogenic | 0.644 | Likely Pathogenic | Likely Benign | 1.00 | Ambiguous | 0.3 | 1.86 | Ambiguous | 1.43 | Ambiguous | 1.40 | Destabilizing | 0.817 | Likely Pathogenic | -4.20 | Deleterious | 0.991 | Probably Damaging | 0.999 | Probably Damaging | 5.76 | Benign | 0.00 | Affected | 0.1844 | 0.1949 | -1 | -3 | -4.6 | -42.08 | ||||||||||||||||||||||||||||||
| c.806T>A | I269K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I269K is not reported in ClinVar and has no entries in gnomAD. Functional prediction tools largely agree on a deleterious effect: SIFT is the sole benign predictor, whereas the remaining methods—SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized—classify the change as pathogenic. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized predicts pathogenicity, the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates pathogenicity, and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is inconclusive. FoldX and Rosetta individually report uncertain effects, and Foldetta remains unavailable. Overall, the consensus of the majority of evidence points to a pathogenic impact, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.216401 | Structured | 0.343787 | Uncertain | 0.937 | 0.244 | 0.125 | -14.609 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 1.45 | Ambiguous | 0.1 | 1.86 | Ambiguous | 1.66 | Ambiguous | 1.55 | Destabilizing | 0.763 | Likely Pathogenic | -5.10 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | 1.76 | Pathogenic | 0.06 | Tolerated | 0.0878 | 0.0713 | -2 | -3 | -8.4 | 15.01 | |||||||||||||||||||||||||||||
| c.1046C>A | P349Q 2D  3DClick to see structure in 3D Viewer AISynGAP1 P349Q is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, and AlphaMissense‑Optimized, while pathogenic predictions arise from PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and FATHMM. The remaining tools (FoldX, Rosetta, premPS, AlphaMissense‑Default, Foldetta) give uncertain or inconclusive results. High‑accuracy assessments further clarify the variant’s likely effect: AlphaMissense‑Optimized predicts a benign outcome; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, indicates a pathogenic outcome; and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, remains uncertain. Overall, the majority of evidence points toward a pathogenic effect, and this conclusion does not conflict with the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.167087 | Structured | 0.348607 | Uncertain | 0.947 | 0.396 | 0.000 | -10.545 | Likely Pathogenic | 0.508 | Ambiguous | Likely Benign | 0.98 | Ambiguous | 0.1 | 1.87 | Ambiguous | 1.43 | Ambiguous | 0.91 | Ambiguous | 0.345 | Likely Benign | -6.40 | Deleterious | 1.000 | Probably Damaging | 0.997 | Probably Damaging | 1.54 | Pathogenic | 0.06 | Tolerated | 0.1480 | 0.4766 | 0 | -1 | -1.9 | 31.01 | |||||||||||||||||||||||||||||
| c.1237C>A | P413T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P413T missense variant is not reported in ClinVar (ClinVar ID None) and has no entries in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are limited to FATHMM, while the remaining tools—SGM‑Consensus, REVEL, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and Foldetta—consistently predict a pathogenic or likely pathogenic impact. High‑accuracy assessments further support this view: AlphaMissense‑Optimized classifies the variant as pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) yields a Likely Pathogenic verdict; and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, also predicts pathogenicity. No predictions are inconclusive or missing. Based on the collective evidence, the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.113710 | Structured | 0.332472 | Uncertain | 0.927 | 0.201 | 0.000 | -12.851 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 3.22 | Destabilizing | 0.2 | 1.87 | Ambiguous | 2.55 | Destabilizing | 0.93 | Ambiguous | 0.554 | Likely Pathogenic | -7.37 | Deleterious | 1.000 | Probably Damaging | 0.993 | Probably Damaging | 3.18 | Benign | 0.01 | Affected | 0.1663 | 0.4670 | 0 | -1 | 0.9 | 3.99 | |||||||||||||||||||||||||||||
| c.1543C>G | R515G 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant R515G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that reach consensus classify the change as pathogenic: REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) all indicate a deleterious effect. Tools with inconclusive results—Rosetta, Foldetta, and AlphaMissense‑Optimized—do not provide evidence for benignity. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as likely pathogenic, and Foldetta as uncertain. Taken together, the overwhelming majority of predictions support a pathogenic impact, and this conclusion does not conflict with the absence of a ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.055536 | Structured | 0.191256 | Uncertain | 0.924 | 0.275 | 0.000 | -11.562 | Likely Pathogenic | 0.807 | Likely Pathogenic | Ambiguous | 2.07 | Destabilizing | 0.3 | 1.87 | Ambiguous | 1.97 | Ambiguous | 1.29 | Destabilizing | 0.674 | Likely Pathogenic | -4.66 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.34 | Pathogenic | 0.04 | Affected | 0.2920 | 0.1998 | -3 | -2 | 4.1 | -99.14 | |||||||||||||||||||||||||||||
| c.1970G>C | W657S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant W657S is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, SIFT, and FATHMM, whereas a majority of tools (SGM‑Consensus, FoldX, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and Foldetta) predict a pathogenic impact; Rosetta remains uncertain. High‑accuracy methods all support pathogenicity: AlphaMissense‑Optimized scores the variant as pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenic. Based on the collective evidence, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status, as no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.030611 | Structured | 0.208729 | Uncertain | 0.941 | 0.245 | 0.000 | -11.817 | Likely Pathogenic | 0.986 | Likely Pathogenic | Likely Pathogenic | 2.27 | Destabilizing | 0.2 | 1.87 | Ambiguous | 2.07 | Destabilizing | 1.26 | Destabilizing | 0.334 | Likely Benign | -11.82 | Deleterious | 0.999 | Probably Damaging | 0.947 | Probably Damaging | 3.52 | Benign | 0.09 | Tolerated | 0.4299 | 0.0489 | -2 | -3 | 0.1 | -99.14 | |||||||||||||||||||||||||||||
| c.2173C>G | L725V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L725V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign calls come from REVEL and AlphaMissense‑Optimized, while pathogenic calls are made by PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and FATHMM. The remaining methods (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Default) are inconclusive. High‑accuracy assessments further clarify the picture: AlphaMissense‑Optimized predicts a benign effect, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates a likely pathogenic outcome, and Foldetta provides no definitive stability change. Overall, the majority of evidence points toward a pathogenic impact, and this conclusion does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | 0.557691 | Disordered | 0.455613 | Uncertain | 0.911 | 0.491 | 0.625 | -8.291 | Likely Pathogenic | 0.461 | Ambiguous | Likely Benign | 1.76 | Ambiguous | 0.1 | 1.87 | Ambiguous | 1.82 | Ambiguous | 0.77 | Ambiguous | 0.183 | Likely Benign | -2.69 | Deleterious | 0.993 | Probably Damaging | 0.992 | Probably Damaging | 1.36 | Pathogenic | 0.01 | Affected | 0.1739 | 0.3977 | 2 | 1 | 0.4 | -14.03 | ||||||||||||||||||||||||||||||
| c.650A>G | E217G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E217G missense variant is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include polyPhen‑2 HumVar, SIFT, and FATHMM, while those that agree on a pathogenic effect include REVEL, PROVEAN, polyPhen‑2 HumDiv, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Pathogenic). Predictions that are uncertain or inconclusive are FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized predicts a pathogenic impact, the SGM‑Consensus also indicates a likely pathogenic outcome, and Foldetta’s stability analysis is inconclusive. Overall, the majority of evidence points to a pathogenic effect for E217G, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.278302 | Structured | 0.404912 | Uncertain | 0.823 | 0.284 | 0.000 | -8.076 | Likely Pathogenic | 0.968 | Likely Pathogenic | Likely Pathogenic | 1.13 | Ambiguous | 0.4 | 1.87 | Ambiguous | 1.50 | Ambiguous | 0.59 | Ambiguous | 0.684 | Likely Pathogenic | -3.93 | Deleterious | 0.816 | Possibly Damaging | 0.307 | Benign | 5.82 | Benign | 0.06 | Tolerated | 0.3278 | 0.6941 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.762G>C | K254N 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant K254N is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools that classify the variant as benign include polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. The majority of other in silico predictors—REVEL, premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN)—indicate a pathogenic effect. Stability‑based methods FoldX, Rosetta, and Foldetta returned uncertain results and are therefore not considered evidence for or against pathogenicity. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as likely pathogenic, and Foldetta as unavailable. Overall, the preponderance of evidence supports a pathogenic classification, which contradicts the current ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.513880 | Disordered | 0.207751 | Uncertain | 0.799 | 0.285 | 0.375 | Uncertain | 1 | -13.306 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.73 | Ambiguous | 0.2 | 1.87 | Ambiguous | 1.30 | Ambiguous | 1.19 | Destabilizing | 0.757 | Likely Pathogenic | -4.23 | Deleterious | 0.384 | Benign | 0.070 | Benign | 5.93 | Benign | 0.01 | Affected | 3.39 | 15 | 0.3200 | 0.1488 | 1 | 0 | 0.4 | -14.07 | 215.3 | -21.0 | -1.0 | 1.7 | 0.2 | 0.3 | X | Potentially Pathogenic | The amino group of Lys254, located in an α-β loop connecting the PH and C2 domains (res. Lys251-Arg258), forms salt bridges with the carboxylate groups of Glu244 and Asp684. Since the neutral carboxamide group of the Asn254 side chain cannot form salt bridges with acidic residues, the residue swap potentially weakens the tertiary structure assembly and/or influences the loop positioning. Regardless, in both the variant and WT simulations, all hydrogen bonds formed by the residue’s side chain were broken, and the residue rotated outwards. The partially α helical conformation of the loop, which extends to a nearby α helix (res. Met414-Asn426), is dynamic, making it unclear if the mutation affects it. | ||||||||||||||||
| c.762G>T | K254N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K254N is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that classify the variant as benign include polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. Tools that predict a pathogenic effect are REVEL, premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM‑Consensus score, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is “Likely Pathogenic” (3 pathogenic vs. 1 benign). High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, SGM‑Consensus as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. No evidence from FoldX or Rosetta alone is conclusive. Based on the overall pattern of predictions, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because the variant is not currently catalogued in ClinVar. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.513880 | Disordered | 0.207751 | Uncertain | 0.799 | 0.285 | 0.375 | -13.306 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.73 | Ambiguous | 0.2 | 1.87 | Ambiguous | 1.30 | Ambiguous | 1.19 | Destabilizing | 0.757 | Likely Pathogenic | -4.23 | Deleterious | 0.384 | Benign | 0.070 | Benign | 5.93 | Benign | 0.01 | Affected | 3.39 | 15 | 0.3200 | 0.1488 | 1 | 0 | 0.4 | -14.07 | 215.3 | -21.0 | -1.0 | 1.7 | 0.2 | 0.3 | X | Potentially Pathogenic | The amino group of Lys254, located in an α-β loop connecting the PH and C2 domains (res. Lys251-Arg258), forms salt bridges with the carboxylate groups of Glu244 and Asp684. Since the neutral carboxamide group of the Asn254 side chain cannot form salt bridges with acidic residues, the residue swap potentially weakens the tertiary structure assembly and/or influences the loop positioning. Regardless, in both the variant and WT simulations, all hydrogen bonds formed by the residue’s side chain were broken, and the residue rotated outwards. The partially α helical conformation of the loop, which extends to a nearby α helix (res. Met414-Asn426), is dynamic, making it unclear if the mutation affects it. | ||||||||||||||||||
| c.959T>G | V320G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V320G is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools cluster into two agreement groups: the single benign prediction comes from REVEL, while the pathogenic group includes FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. Two tools give uncertain results: Rosetta and AlphaMissense‑Optimized. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is uncertain; the SGM Consensus—derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—returns pathogenic; and Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, also predicts pathogenic. Overall, the preponderance of evidence indicates that V320G is most likely pathogenic, and this conclusion does not contradict any existing ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.185198 | Structured | 0.419626 | Uncertain | 0.905 | 0.266 | 0.125 | -9.043 | Likely Pathogenic | 0.816 | Likely Pathogenic | Ambiguous | 2.15 | Destabilizing | 1.1 | 1.87 | Ambiguous | 2.01 | Destabilizing | 1.48 | Destabilizing | 0.438 | Likely Benign | -5.74 | Deleterious | 0.958 | Probably Damaging | 0.999 | Probably Damaging | 1.89 | Pathogenic | 0.02 | Affected | 0.1688 | 0.1949 | -1 | -3 | -4.6 | -42.08 | ||||||||||||||||||||||||||||||
| c.1211C>G | A404G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A404G is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are premPS and SIFT. FoldX, Rosetta, and Foldetta give uncertain or inconclusive results and are treated as unavailable. The high‑accuracy consensus (SGM‑Consensus) classifies the variant as Likely Benign, and AlphaMissense‑Optimized also predicts Benign. Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, remains uncertain. Overall, the majority of evidence points to a benign impact for A404G, and this assessment does not contradict any ClinVar status, as none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.232838 | Structured | 0.415505 | Uncertain | 0.965 | 0.355 | 0.000 | -5.277 | Likely Benign | 0.137 | Likely Benign | Likely Benign | 0.92 | Ambiguous | 0.0 | 1.88 | Ambiguous | 1.40 | Ambiguous | 1.18 | Destabilizing | 0.099 | Likely Benign | -2.34 | Neutral | 0.045 | Benign | 0.013 | Benign | 4.03 | Benign | 0.00 | Affected | 0.2370 | 0.5054 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.1408A>G | M470V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M470V is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Consensus from most in silico predictors indicates a pathogenic effect: SGM‑Consensus, REVEL, FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and FATHMM all score it as deleterious. Only two tools—SIFT and AlphaMissense‑Optimized—classify it as benign, while Rosetta and AlphaMissense‑Default remain inconclusive. High‑accuracy assessments reinforce the pathogenic prediction: AlphaMissense‑Optimized reports a benign outcome, but the SGM‑Consensus (derived from a majority of pathogenic calls among AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) and Foldetta (combining pathogenic FoldX with uncertain Rosetta) both predict pathogenicity. Overall, the preponderance of evidence supports a likely pathogenic classification, which does not conflict with the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.298791 | Structured | 0.351497 | Uncertain | 0.908 | 0.272 | 0.000 | Uncertain | 1 | -8.856 | Likely Pathogenic | 0.478 | Ambiguous | Likely Benign | 2.73 | Destabilizing | 0.1 | 1.88 | Ambiguous | 2.31 | Destabilizing | 1.31 | Destabilizing | 0.770 | Likely Pathogenic | -3.58 | Deleterious | 0.999 | Probably Damaging | 0.993 | Probably Damaging | -1.20 | Pathogenic | 0.15 | Tolerated | 3.37 | 34 | 0.2710 | 0.3256 | 1 | 2 | 2.3 | -32.06 | |||||||||||||||||||||||||
| c.1541T>C | I514T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I514T has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect are limited to FATHMM, whereas the majority of algorithms—SGM‑Consensus, REVEL, FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—classify the change as pathogenic. Rosetta reports an uncertain outcome and is not included in the consensus groups. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized predicts pathogenic; the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates likely pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, also predicts pathogenic. Taken together, the evidence overwhelmingly points to a pathogenic effect, and this conclusion is not contradicted by the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.049374 | Structured | 0.221408 | Uncertain | 0.948 | 0.266 | 0.000 | -8.820 | Likely Pathogenic | 0.963 | Likely Pathogenic | Likely Pathogenic | 2.92 | Destabilizing | 0.1 | 1.88 | Ambiguous | 2.40 | Destabilizing | 1.94 | Destabilizing | 0.617 | Likely Pathogenic | -4.77 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 2.82 | Benign | 0.00 | Affected | 0.0962 | 0.0480 | 0 | -1 | -5.2 | -12.05 | |||||||||||||||||||||||||||||
| c.1672C>G | H558D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant H558D is not reported in ClinVar and is absent from gnomAD. Prediction tools cluster into two groups: the single benign prediction comes from SIFT, while the remaining tools—REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus—indicate pathogenicity. High‑accuracy assessments further support a deleterious effect: the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is “Likely Pathogenic”; AlphaMissense‑Optimized and Foldetta are “Uncertain.” No evidence suggests a benign outcome. Consequently, the variant is most likely pathogenic based on the preponderance of predictions, and this assessment does not contradict the ClinVar status, which currently contains no entry for H558D. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.033407 | Structured | 0.011039 | Uncertain | 0.897 | 0.200 | 0.000 | -14.948 | Likely Pathogenic | 0.823 | Likely Pathogenic | Ambiguous | 0.54 | Ambiguous | 0.1 | 1.88 | Ambiguous | 1.21 | Ambiguous | 1.03 | Destabilizing | 0.635 | Likely Pathogenic | -5.90 | Deleterious | 0.959 | Probably Damaging | 0.905 | Possibly Damaging | -1.26 | Pathogenic | 0.09 | Tolerated | 0.2233 | 0.0908 | 1 | -1 | -0.3 | -22.05 | |||||||||||||||||||||||||||||
| c.1694T>G | L565R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L565R is not reported in ClinVar (ClinVar status: not listed) but is present in the gnomAD database (gnomAD ID: 6‑33440746‑T‑G). Prediction tools that agree on a benign effect include only FATHMM, whereas the remaining tools—REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, Foldetta, and the SGM Consensus—consistently predict a pathogenic impact. The Rosetta stability assessment is inconclusive and is therefore treated as unavailable. High‑accuracy methods all support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. Based on the collective predictions, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar evidence (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.034884 | Structured | 0.045819 | Uncertain | 0.922 | 0.205 | 0.000 | 6-33440746-T-G | -16.070 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 4.71 | Destabilizing | 0.1 | 1.88 | Ambiguous | 3.30 | Destabilizing | 2.66 | Destabilizing | 0.547 | Likely Pathogenic | -5.97 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | 2.74 | Benign | 0.00 | Affected | 3.37 | 35 | 0.1373 | 0.0615 | -2 | -3 | -8.3 | 43.03 | ||||||||||||||||||||||||||
| c.1820T>A | L607H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L607H is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on pathogenicity include REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized; no tool predicts a benign effect. Uncertain predictions come from FoldX, Rosetta, and Foldetta. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, while Foldetta’s stability analysis is inconclusive. Taken together, the overwhelming majority of evidence indicates that the variant is most likely pathogenic, and this conclusion does not contradict any existing ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.048328 | Structured | 0.194229 | Uncertain | 0.869 | 0.250 | 0.000 | -14.775 | Likely Pathogenic | 0.981 | Likely Pathogenic | Likely Pathogenic | 0.76 | Ambiguous | 0.1 | 1.88 | Ambiguous | 1.32 | Ambiguous | 1.38 | Destabilizing | 0.906 | Likely Pathogenic | -6.97 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.48 | Pathogenic | 0.01 | Affected | 0.1199 | 0.0541 | -2 | -3 | -7.0 | 23.98 | |||||||||||||||||||||||||||||
| c.1954T>C | F652L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F652L is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions are made by REVEL and FATHMM, whereas the remaining evaluated algorithms (premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) all indicate pathogenicity. Stability‑based assessments from FoldX and Rosetta are inconclusive, and Foldetta likewise yields an uncertain result. High‑accuracy methods give a clearer picture: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) labels the variant as likely pathogenic, and Foldetta remains uncertain. Overall, the preponderance of evidence points to a pathogenic effect for F652L. This conclusion is consistent with the absence of ClinVar annotation, so there is no contradiction with existing clinical data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.098513 | Structured | 0.356594 | Uncertain | 0.966 | 0.338 | 0.000 | -9.026 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 1.31 | Ambiguous | 0.1 | 1.88 | Ambiguous | 1.60 | Ambiguous | 1.35 | Destabilizing | 0.467 | Likely Benign | -5.65 | Deleterious | 0.997 | Probably Damaging | 0.619 | Possibly Damaging | 3.09 | Benign | 0.00 | Affected | 0.1750 | 0.2926 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1956T>A | F652L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F652L is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions are made by REVEL and FATHMM, whereas the remaining evaluated algorithms (premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) all indicate pathogenicity. Stability‑based assessments from FoldX and Rosetta are inconclusive, and Foldetta likewise yields an uncertain result. High‑accuracy methods give a clearer picture: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) labels the variant as likely pathogenic, and Foldetta remains uncertain. Overall, the preponderance of evidence points to a pathogenic effect for F652L. This conclusion is consistent with the absence of ClinVar annotation, so there is no contradiction with existing clinical data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.098513 | Structured | 0.356594 | Uncertain | 0.966 | 0.338 | 0.000 | -9.026 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 1.31 | Ambiguous | 0.1 | 1.88 | Ambiguous | 1.60 | Ambiguous | 1.35 | Destabilizing | 0.306 | Likely Benign | -5.65 | Deleterious | 0.997 | Probably Damaging | 0.619 | Possibly Damaging | 3.09 | Benign | 0.00 | Affected | 0.1750 | 0.2926 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1956T>G | F652L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F652L is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL and FATHMM, whereas the majority of other in silico predictors (premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) indicate a pathogenic impact. FoldX and Rosetta give uncertain results, and Foldetta likewise reports an uncertain stability change. High‑accuracy assessments show AlphaMissense‑Optimized classifying the variant as pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) labeling it Likely Pathogenic, and Foldetta remaining uncertain. Overall, the preponderance of evidence from multiple pathogenic‑predicting tools and the high‑accuracy methods points to a likely pathogenic effect for F652L, with no conflict from ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.098513 | Structured | 0.356594 | Uncertain | 0.966 | 0.338 | 0.000 | -9.026 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 1.31 | Ambiguous | 0.1 | 1.88 | Ambiguous | 1.60 | Ambiguous | 1.35 | Destabilizing | 0.305 | Likely Benign | -5.65 | Deleterious | 0.997 | Probably Damaging | 0.619 | Possibly Damaging | 3.09 | Benign | 0.00 | Affected | 0.1750 | 0.2926 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.2006A>G | N669S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N669S is not reported in ClinVar (ClinVar ID None) but is present in gnomAD (ID 6‑33441265‑A‑G). Functional prediction tools that agree on a benign effect include REVEL, premPS, SIFT, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive (2 benign vs 2 pathogenic), and Foldetta (combining FoldX‑MD and Rosetta) is also inconclusive. No folding‑stability metrics (FoldX, Rosetta, Foldetta) provide definitive evidence. Overall, the majority of predictions lean toward a benign impact, and this is consistent with the absence of a ClinVar pathogenic classification. Thus, the variant is most likely benign, with no contradiction to ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.142424 | Structured | 0.086615 | Uncertain | 0.872 | 0.380 | 0.000 | 6-33441265-A-G | 3 | 1.86e-6 | -8.369 | Likely Pathogenic | 0.187 | Likely Benign | Likely Benign | 0.55 | Ambiguous | 0.1 | 1.88 | Ambiguous | 1.22 | Ambiguous | 0.35 | Likely Benign | 0.210 | Likely Benign | -4.02 | Deleterious | 0.999 | Probably Damaging | 0.960 | Probably Damaging | 3.52 | Benign | 0.14 | Tolerated | 3.39 | 27 | 0.3521 | 0.4480 | 1 | 1 | 2.7 | -27.03 | |||||||||||||||||||||||||
| c.809A>T | E270V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E270V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that assess sequence conservation and structural impact uniformly classify the substitution as pathogenic: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all return pathogenic scores. No tool predicts a benign effect. Uncertain results are reported only by FoldX, Rosetta, Foldetta, and premPS, which are not considered evidence for or against pathogenicity. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is pathogenic; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is “Likely Pathogenic”; Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, is inconclusive. Overall, the consensus of available predictions indicates that E270V is most likely pathogenic, and this conclusion does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.144935 | Structured | 0.382573 | Uncertain | 0.938 | 0.231 | 0.125 | -15.969 | Likely Pathogenic | 0.996 | Likely Pathogenic | Likely Pathogenic | 1.38 | Ambiguous | 0.6 | 1.88 | Ambiguous | 1.63 | Ambiguous | -0.52 | Ambiguous | 0.574 | Likely Pathogenic | -6.43 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 1.53 | Pathogenic | 0.00 | Affected | 0.0672 | 0.4498 | -2 | -2 | 7.7 | -29.98 | |||||||||||||||||||||||||||||
| c.1231A>C | I411L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I411L is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, and FATHMM, whereas a majority (premPS, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, and AlphaMissense‑Default) predict a pathogenic impact. Tools with uncertain or mixed outputs are FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments are inconclusive: AlphaMissense‑Optimized is uncertain; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is a 2‑vs‑2 tie, yielding no definitive call; and Foldetta also reports an uncertain stability change. Overall, the majority of standard predictors lean toward pathogenicity, and this conclusion does not contradict the lack of ClinVar annotation. Thus, the variant is most likely pathogenic based on current computational evidence, though high‑accuracy tools remain inconclusive. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.116183 | Structured | 0.339366 | Uncertain | 0.927 | 0.198 | 0.000 | -10.723 | Likely Pathogenic | 0.819 | Likely Pathogenic | Ambiguous | 1.02 | Ambiguous | 0.3 | 1.89 | Ambiguous | 1.46 | Ambiguous | 1.17 | Destabilizing | 0.285 | Likely Benign | -1.84 | Neutral | 0.908 | Possibly Damaging | 0.943 | Probably Damaging | 3.36 | Benign | 0.01 | Affected | 0.0876 | 0.3539 | 2 | 2 | -0.7 | 0.00 | ||||||||||||||||||||||||||||||
| c.1404G>A | M468I 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant M468I is listed in ClinVar with an uncertain significance (ClinVar ID 3657719.0) and is present in gnomAD (6‑33438436‑G‑A). Functional prediction tools cluster into two groups: benign predictions come from premPS, PROVEAN, and SIFT, while pathogenic predictions arise from REVEL, FoldX, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, and AlphaMissense‑Default. Two tools report uncertainty: AlphaMissense‑Optimized and Rosetta. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is pathogenic. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is inconclusive, SGM Consensus is pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. Overall, the preponderance of evidence indicates a pathogenic impact for M468I, which does not contradict the ClinVar uncertain status but suggests a likely pathogenic classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.284882 | Structured | 0.339253 | Uncertain | 0.932 | 0.257 | 0.000 | Uncertain | 1 | 6-33438436-G-A | 1 | 6.20e-7 | -8.583 | Likely Pathogenic | 0.907 | Likely Pathogenic | Ambiguous | 2.53 | Destabilizing | 0.2 | 1.89 | Ambiguous | 2.21 | Destabilizing | 0.37 | Likely Benign | 0.508 | Likely Pathogenic | -1.06 | Neutral | 0.748 | Possibly Damaging | 0.886 | Possibly Damaging | -1.10 | Pathogenic | 0.07 | Tolerated | 3.37 | 31 | 0.1369 | 0.3354 | 1 | 2 | 2.6 | -18.03 | ||||||||||||||||||||||
| c.1404G>C | M468I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M468I is not reported in ClinVar and has no entry in gnomAD. Prediction tools that agree on a benign effect include premPS, PROVEAN, and SIFT, whereas the majority of algorithms—SGM‑Consensus, REVEL, FoldX, Foldetta, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, and AlphaMissense‑Default—classify the change as pathogenic. Two methods report uncertainty: Rosetta and AlphaMissense‑Optimized. High‑accuracy assessments further support a deleterious outcome: the SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts pathogenicity, and Foldetta, which integrates FoldX‑MD and Rosetta outputs, also indicates a pathogenic effect. AlphaMissense‑Optimized remains inconclusive. Overall, the preponderance of evidence points to a pathogenic impact for M468I, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.284882 | Structured | 0.339253 | Uncertain | 0.932 | 0.257 | 0.000 | -8.583 | Likely Pathogenic | 0.907 | Likely Pathogenic | Ambiguous | 2.53 | Destabilizing | 0.2 | 1.89 | Ambiguous | 2.21 | Destabilizing | 0.37 | Likely Benign | 0.508 | Likely Pathogenic | -1.06 | Neutral | 0.748 | Possibly Damaging | 0.886 | Possibly Damaging | -1.10 | Pathogenic | 0.07 | Tolerated | 3.37 | 31 | 0.1369 | 0.3354 | 1 | 2 | 2.6 | -18.03 | |||||||||||||||||||||||||||
| c.1404G>T | M468I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M468I is not reported in ClinVar and has no entry in gnomAD. Prediction tools that agree on a benign effect include premPS, PROVEAN, and SIFT, whereas the majority of algorithms—SGM‑Consensus, REVEL, FoldX, Foldetta, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, and AlphaMissense‑Default—classify the change as pathogenic. Two methods report uncertainty: Rosetta and AlphaMissense‑Optimized. High‑accuracy assessments further support a deleterious outcome: the SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts pathogenicity, and Foldetta, which integrates FoldX‑MD and Rosetta outputs, also indicates a pathogenic effect; AlphaMissense‑Optimized remains inconclusive. Overall, the preponderance of evidence points to a pathogenic impact for M468I, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.284882 | Structured | 0.339253 | Uncertain | 0.932 | 0.257 | 0.000 | -8.583 | Likely Pathogenic | 0.907 | Likely Pathogenic | Ambiguous | 2.53 | Destabilizing | 0.2 | 1.89 | Ambiguous | 2.21 | Destabilizing | 0.37 | Likely Benign | 0.510 | Likely Pathogenic | -1.06 | Neutral | 0.748 | Possibly Damaging | 0.886 | Possibly Damaging | -1.10 | Pathogenic | 0.07 | Tolerated | 3.37 | 31 | 0.1369 | 0.3354 | 1 | 2 | 2.6 | -18.03 | |||||||||||||||||||||||||||
| c.1579G>T | D527Y 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant D527Y is listed in ClinVar with an uncertain significance (ClinVar ID 1698369.0) and is not reported in gnomAD. Functional prediction tools cluster into two groups: the single benign prediction from premPS versus a consensus of pathogenic predictions from the remaining 12 tools (REVEL, SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized). High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is inconclusive. Protein‑stability calculations from FoldX and Rosetta are also uncertain. Overall, the preponderance of evidence indicates that D527Y is most likely pathogenic, which does not contradict the current ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.139895 | Structured | 0.021908 | Uncertain | 0.913 | 0.408 | 0.000 | Uncertain | 1 | -15.386 | Likely Pathogenic | 0.978 | Likely Pathogenic | Likely Pathogenic | -0.77 | Ambiguous | 0.2 | 1.89 | Ambiguous | 0.56 | Ambiguous | -0.14 | Likely Benign | 0.905 | Likely Pathogenic | -8.79 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | -2.41 | Pathogenic | 0.00 | Affected | 3.37 | 35 | 0.0554 | 0.4229 | -4 | -3 | 2.2 | 48.09 | 270.9 | -45.7 | 0.1 | 0.1 | -0.1 | 0.0 | X | Potentially Pathogenic | Asp527 is located on an α-α loop between the two α-helices (res. Gly502-Tyr518 and Ala533-Val560). In the WT simulations, the carboxylate group of the Asp527 side chain forms hydrogen bonds with the backbone atoms of loop residues (e.g., Ile529, Lys530) facing the membrane surface. In the variant simulations, Tyr527 is a bulkier residue that faces away from the loop and stacks with Phe646 in a nearby α-helix (res. Ser614-Ser668). Regardless, no negative structural effects are observed during the variant simulations. However, due to its location near the SynGAP-membrane interface, the effect of the residue swap cannot be fully addressed using the SynGAP solvent-only simulations. | ||||||||||||||||
| c.1846G>C | D616H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D616H missense variant is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, and FATHMM. Those that agree on a pathogenic effect comprise SGM‑Consensus, FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. Two tools give uncertain results—Rosetta and AlphaMissense‑Optimized—so their outputs are treated as unavailable for inference. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is uncertain, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta) is Pathogenic. Overall, the majority of evidence points to a pathogenic effect. The variant’s predicted pathogenicity does not contradict ClinVar status, as no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.129801 | Structured | 0.166689 | Uncertain | 0.867 | 0.252 | 0.000 | -9.815 | Likely Pathogenic | 0.904 | Likely Pathogenic | Ambiguous | 2.13 | Destabilizing | 0.2 | 1.89 | Ambiguous | 2.01 | Destabilizing | 0.45 | Likely Benign | 0.316 | Likely Benign | -5.57 | Deleterious | 0.999 | Probably Damaging | 0.952 | Probably Damaging | 3.30 | Benign | 0.03 | Affected | 0.1330 | 0.4273 | 1 | -1 | 0.3 | 22.05 | |||||||||||||||||||||||||||||
| c.2017C>A | L673I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L673I is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include SGM‑Consensus (Likely Benign), REVEL, premPS, PROVEAN, polyPhen‑2 HumVar, SIFT, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Only polyPhen‑2 HumDiv predicts a pathogenic outcome. The remaining tools (FoldX, Rosetta, Foldetta, ESM1b) return uncertain or inconclusive results and are treated as unavailable evidence. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely benign, and Foldetta as uncertain. Taken together, the majority of reliable predictors indicate a benign effect, and there is no conflict with ClinVar status (which has no entry). Therefore, the variant is most likely benign. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.060549 | Structured | 0.104692 | Uncertain | 0.545 | 0.369 | 0.000 | -7.823 | In-Between | 0.099 | Likely Benign | Likely Benign | 1.51 | Ambiguous | 0.2 | 1.89 | Ambiguous | 1.70 | Ambiguous | -0.02 | Likely Benign | 0.036 | Likely Benign | -0.14 | Neutral | 0.535 | Possibly Damaging | 0.112 | Benign | 3.37 | Benign | 0.47 | Tolerated | 0.0910 | 0.3689 | 2 | 2 | 0.7 | 0.00 | |||||||||||||||||||||||||||||
| c.713A>C | E238A 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant E238A is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: FATHMM predicts benign, whereas the remaining evaluated algorithms (SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) all predict pathogenicity. Four methods (FoldX, Rosetta, Foldetta, premPS) return uncertain results and are treated as unavailable. High‑accuracy assessments further support a damaging outcome: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta is uncertain. Overall, the preponderance of evidence indicates that E238A is most likely pathogenic, and this conclusion does not contradict the current ClinVar status, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.194234 | Structured | 0.332638 | Uncertain | 0.796 | 0.326 | 0.000 | -13.252 | Likely Pathogenic | 0.990 | Likely Pathogenic | Likely Pathogenic | 0.91 | Ambiguous | 0.3 | 1.89 | Ambiguous | 1.40 | Ambiguous | 0.57 | Ambiguous | 0.879 | Likely Pathogenic | -5.44 | Deleterious | 0.970 | Probably Damaging | 0.681 | Possibly Damaging | 5.44 | Benign | 0.04 | Affected | 0.4164 | 0.5610 | 0 | -1 | 5.3 | -58.04 | |||||||||||||||||||||||||||||
| c.754C>A | P252T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P252T is not reported in ClinVar and is absent from gnomAD. Prediction tools that classify it as benign include polyPhen‑2 HumDiv, polyPhen‑2 HumVar and FATHMM. Those that predict a pathogenic effect are REVEL, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized and the SGM‑Consensus score (Likely Pathogenic). Four tools (FoldX, Rosetta, Foldetta, premPS) give uncertain results and are treated as unavailable. High‑accuracy methods give the following: AlphaMissense‑Optimized predicts pathogenic; the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates Likely Pathogenic; Foldetta’s stability assessment is uncertain. Overall, the majority of evidence points to a pathogenic effect, and this is consistent with the lack of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.461924 | Structured | 0.211606 | Uncertain | 0.753 | 0.304 | 0.250 | -11.824 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 1.24 | Ambiguous | 0.1 | 1.89 | Ambiguous | 1.57 | Ambiguous | 0.71 | Ambiguous | 0.828 | Likely Pathogenic | -7.35 | Deleterious | 0.384 | Benign | 0.177 | Benign | 5.78 | Benign | 0.01 | Affected | 0.1514 | 0.4842 | 0 | -1 | 0.9 | 3.99 | |||||||||||||||||||||||||||||
| c.1187G>C | G396A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G396A is not reported in ClinVar and is absent from gnomAD. Functional prediction tools uniformly indicate a benign effect: REVEL, premPS, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, PROVEAN, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the change as benign. No tool predicts pathogenicity. High‑accuracy assessments corroborate this: AlphaMissense‑Optimized predicts benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also yields a benign verdict, while Foldetta (combining FoldX‑MD and Rosetta outputs) is inconclusive. Consequently, the variant is most likely benign based on the available predictions, and this assessment does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.414856 | Structured | 0.394626 | Uncertain | 0.640 | 0.584 | 0.500 | -3.655 | Likely Benign | 0.103 | Likely Benign | Likely Benign | 1.74 | Ambiguous | 0.7 | 1.90 | Ambiguous | 1.82 | Ambiguous | 0.41 | Likely Benign | 0.274 | Likely Benign | -1.28 | Neutral | 0.062 | Benign | 0.024 | Benign | 3.93 | Benign | 0.84 | Tolerated | 0.3846 | 0.5255 | 1 | 0 | 2.2 | 14.03 | |||||||||||||||||||||||||||||
| c.1405G>A | A469T 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant A469T is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from premPS, PROVEAN, SIFT, and AlphaMissense‑Optimized; pathogenic predictions arise from REVEL, FoldX, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN), and the Foldetta stability assessment (combining FoldX‑MD and Rosetta). The high‑accuracy subset shows AlphaMissense‑Optimized as benign, whereas SGM Consensus and Foldetta both predict pathogenic. Overall, the majority of evidence supports a pathogenic effect, and this conclusion does not conflict with the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.278302 | Structured | 0.343926 | Uncertain | 0.910 | 0.276 | 0.000 | Uncertain | 1 | -9.540 | Likely Pathogenic | 0.723 | Likely Pathogenic | Likely Benign | 2.26 | Destabilizing | 0.1 | 1.90 | Ambiguous | 2.08 | Destabilizing | 0.34 | Likely Benign | 0.527 | Likely Pathogenic | -1.46 | Neutral | 0.994 | Probably Damaging | 0.986 | Probably Damaging | -1.21 | Pathogenic | 0.42 | Tolerated | 0.1005 | 0.5884 | 1 | 0 | -2.5 | 30.03 | |||||||||||||||||||||||||||
| c.1426T>G | F476V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 F476V variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, and FATHMM. Those that agree on a pathogenic effect are FoldX, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, and Foldetta. Tools with uncertain or mixed outputs are Rosetta, premPS, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive, and Foldetta predicts a pathogenic impact. Overall, the majority of evidence points toward a pathogenic effect. The variant is most likely pathogenic based on current predictions, and this assessment does not contradict any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.257454 | Structured | 0.397815 | Uncertain | 0.821 | 0.250 | 0.000 | -12.329 | Likely Pathogenic | 0.918 | Likely Pathogenic | Ambiguous | 3.01 | Destabilizing | 0.2 | 1.90 | Ambiguous | 2.46 | Destabilizing | 0.64 | Ambiguous | 0.353 | Likely Benign | -1.63 | Neutral | 0.996 | Probably Damaging | 0.993 | Probably Damaging | 3.49 | Benign | 0.53 | Tolerated | 0.1478 | 0.2251 | -1 | -1 | 1.4 | -48.04 | ||||||||||||||||||||||||||||||
| c.1444C>A | L482I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L482I is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign calls come from PROVEAN and AlphaMissense‑Optimized, whereas the majority of other in silico predictors (REVEL, polyPhen‑2 HumDiv/HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default) indicate pathogenicity. Uncertain results are reported by FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments give a consistent picture: AlphaMissense‑Optimized predicts a benign effect, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) classifies the variant as likely pathogenic, and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, yields an uncertain outcome. Overall, the preponderance of pathogenic predictions suggests that the variant is most likely pathogenic, and this assessment does not contradict the absence of a ClinVar entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.254060 | Structured | 0.426236 | Uncertain | 0.795 | 0.248 | 0.000 | -11.116 | Likely Pathogenic | 0.760 | Likely Pathogenic | Likely Benign | 1.29 | Ambiguous | 0.1 | 1.90 | Ambiguous | 1.60 | Ambiguous | 0.71 | Ambiguous | 0.600 | Likely Pathogenic | -1.97 | Neutral | 0.994 | Probably Damaging | 0.994 | Probably Damaging | -1.31 | Pathogenic | 0.05 | Affected | 0.0677 | 0.2368 | 2 | 2 | 0.7 | 0.00 | |||||||||||||||||||||||||||||
| c.1474A>G | K492E 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant K492E is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Prediction tools that classify the variant as benign include only FATHMM. The remaining tools—REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus—predict it to be pathogenic or likely pathogenic. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized scores it as pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) reports it as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is inconclusive. Overall, the preponderance of evidence indicates that the variant is most likely pathogenic, which contradicts its current ClinVar status of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.061840 | Structured | 0.327121 | Uncertain | 0.947 | 0.192 | 0.000 | Conflicting | 2 | -16.175 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 1.53 | Ambiguous | 0.1 | 1.90 | Ambiguous | 1.72 | Ambiguous | 1.42 | Destabilizing | 0.510 | Likely Pathogenic | -3.98 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 2.99 | Benign | 0.01 | Affected | 3.37 | 35 | 0.2968 | 0.0650 | 1 | 0 | 0.4 | 0.94 | |||||||||||||||||||||||||
| c.1780T>G | F594V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F594V lies within the GAP domain. ClinVar has no entry for this variant, and it is not reported in gnomAD. Prediction tools that assess pathogenicity unanimously favor a deleterious effect: SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict pathogenicity; only Rosetta is uncertain. No tool predicts a benign outcome. High‑accuracy methods reinforce this view: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; and Foldetta (combining FoldX‑MD and Rosetta) is pathogenic. Based on the collective predictions, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which simply lacks an entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.009187 | Structured | 0.120166 | Uncertain | 0.946 | 0.147 | 0.000 | -12.648 | Likely Pathogenic | 0.993 | Likely Pathogenic | Likely Pathogenic | 2.91 | Destabilizing | 0.2 | 1.90 | Ambiguous | 2.41 | Destabilizing | 1.80 | Destabilizing | 0.931 | Likely Pathogenic | -6.97 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | -1.91 | Pathogenic | 0.01 | Affected | 0.1813 | 0.1575 | -1 | -1 | 1.4 | -48.04 | |||||||||||||||||||||||||||||
| c.1867C>A | L623I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L623I is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: benign calls come only from REVEL and PROVEAN, while pathogenic calls are made by FoldX, premPS, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus (which is derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). Uncertain results are reported by Rosetta and AlphaMissense‑Optimized. High‑accuracy assessments reinforce the pathogenic prediction: AlphaMissense‑Optimized is inconclusive, the SGM‑Consensus is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenicity. Overall, the preponderance of evidence indicates that the variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.175930 | Structured | 0.060667 | Uncertain | 0.962 | 0.211 | 0.000 | -11.952 | Likely Pathogenic | 0.911 | Likely Pathogenic | Ambiguous | 3.15 | Destabilizing | 0.5 | 1.90 | Ambiguous | 2.53 | Destabilizing | 1.13 | Destabilizing | 0.398 | Likely Benign | -1.99 | Neutral | 0.999 | Probably Damaging | 0.997 | Probably Damaging | 1.63 | Pathogenic | 0.02 | Affected | 0.1109 | 0.3767 | 2 | 2 | 0.7 | 0.00 | |||||||||||||||||||||||||||||
| c.1172G>C | G391A 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant G391A is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic outcome are FoldX, polyPhen‑2 HumDiv, and FATHMM. Predictions that are inconclusive are Rosetta and Foldetta. The high‑accuracy consensus from AlphaMissense‑Optimized is benign, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is also likely benign, and Foldetta remains uncertain. Overall, the majority of evidence points to a benign effect, and this is consistent with the lack of a ClinVar pathogenic classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.637480 | Disordered | 0.409509 | Uncertain | 0.279 | 0.741 | 0.750 | -5.712 | Likely Benign | 0.120 | Likely Benign | Likely Benign | 2.02 | Destabilizing | 0.5 | 1.92 | Ambiguous | 1.97 | Ambiguous | 0.11 | Likely Benign | 0.442 | Likely Benign | -0.76 | Neutral | 0.633 | Possibly Damaging | 0.219 | Benign | 1.33 | Pathogenic | 0.19 | Tolerated | 0.3828 | 0.4870 | 1 | 0 | 2.2 | 14.03 | |||||||||||||||||||||||||||||
| c.2187C>A | N729K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N729K has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus call (Likely Benign). Only AlphaMissense‑Default predicts a pathogenic outcome. Tools with uncertain or mixed results are Foldetta (protein‑folding stability) and Rosetta. High‑accuracy assessments: AlphaMissense‑Optimized reports a benign effect; the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates likely benign; Foldetta’s stability prediction is inconclusive. Overall, the majority of evidence points to a benign impact for the variant, and this conclusion does not contradict the current ClinVar status, which contains no report for this change. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | 0.750527 | Disordered | 0.426547 | Uncertain | 0.651 | 0.583 | 0.625 | -5.101 | Likely Benign | 0.648 | Likely Pathogenic | Likely Benign | -0.03 | Likely Benign | 0.1 | 1.92 | Ambiguous | 0.95 | Ambiguous | 0.12 | Likely Benign | 0.036 | Likely Benign | -1.39 | Neutral | 0.109 | Benign | 0.033 | Benign | 3.51 | Benign | 0.47 | Tolerated | 0.1948 | 0.3612 | 1 | 0 | -0.4 | 14.07 | ||||||||||||||||||||||||||||||
| c.2187C>G | N729K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N729K has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus call (Likely Benign). Only AlphaMissense‑Default predicts a pathogenic outcome. Tools with uncertain or mixed results are Foldetta (protein‑folding stability) and Rosetta. High‑accuracy assessments: AlphaMissense‑Optimized reports a benign effect; the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates likely benign; Foldetta’s stability prediction is inconclusive. Overall, the majority of evidence points to a benign impact for the variant, and this conclusion does not contradict the current ClinVar status, which contains no report for this change. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | 0.750527 | Disordered | 0.426547 | Uncertain | 0.651 | 0.583 | 0.625 | -5.101 | Likely Benign | 0.648 | Likely Pathogenic | Likely Benign | -0.03 | Likely Benign | 0.1 | 1.92 | Ambiguous | 0.95 | Ambiguous | 0.12 | Likely Benign | 0.036 | Likely Benign | -1.39 | Neutral | 0.109 | Benign | 0.033 | Benign | 3.51 | Benign | 0.47 | Tolerated | 0.1948 | 0.3612 | 1 | 0 | -0.4 | 14.07 | ||||||||||||||||||||||||||||||
| c.761A>C | K254T 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant K254T is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: pathogenic calls come from REVEL, PROVEAN, both polyPhen‑2 HumDiv and HumVar, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Pathogenic). Benign predictions are limited to SIFT and FATHMM. Uncertain results are reported by FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments reinforce the pathogenic signal: AlphaMissense‑Optimized predicts pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates Likely Pathogenic, while Foldetta’s stability analysis is inconclusive. Overall, the majority of evidence points to a pathogenic effect for K254T, and this conclusion does not conflict with ClinVar status, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.513880 | Disordered | 0.207751 | Uncertain | 0.799 | 0.285 | 0.375 | -13.793 | Likely Pathogenic | 0.991 | Likely Pathogenic | Likely Pathogenic | 0.51 | Ambiguous | 0.0 | 1.92 | Ambiguous | 1.22 | Ambiguous | 0.62 | Ambiguous | 0.883 | Likely Pathogenic | -5.14 | Deleterious | 0.970 | Probably Damaging | 0.749 | Possibly Damaging | 5.88 | Benign | 0.06 | Tolerated | 0.1825 | 0.2562 | 0 | -1 | 3.2 | -27.07 | |||||||||||||||||||||||||||||
| c.1388A>G | D463G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D463G missense variant is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, and FATHMM, while pathogenic predictions arise from PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, and ESM1b. Five tools favor pathogenicity versus three favor benign, with the remaining five (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Optimized) yielding uncertain results. High‑accuracy assessments are mixed: AlphaMissense‑Optimized is inconclusive; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as likely pathogenic; Foldetta, which integrates FoldX‑MD and Rosetta outputs, also remains inconclusive. Overall, the preponderance of evidence points to a pathogenic effect, and this conclusion is not contradicted by ClinVar, which has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.260850 | Structured | 0.305622 | Uncertain | 0.940 | 0.176 | 0.000 | -10.713 | Likely Pathogenic | 0.921 | Likely Pathogenic | Ambiguous | 0.66 | Ambiguous | 0.1 | 1.93 | Ambiguous | 1.30 | Ambiguous | 0.54 | Ambiguous | 0.422 | Likely Benign | -6.36 | Deleterious | 0.994 | Probably Damaging | 0.824 | Possibly Damaging | 3.32 | Benign | 0.09 | Tolerated | 0.3751 | 0.4898 | 1 | -1 | 3.1 | -58.04 | |||||||||||||||||||||||||||||
| c.1159G>A | G387S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 G387S missense variant is not reported in ClinVar (ClinVar status: none) but is present in gnomAD (variant ID 6‑33438064‑G‑A). Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are FoldX and FATHMM; Rosetta is uncertain. The high‑accuracy consensus (SGM‑Consensus) is derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN and is therefore likely benign. AlphaMissense‑Optimized itself predicts benign. Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, predicts a pathogenic effect. Overall, the majority of evidence points to a benign impact, and this assessment does not contradict any ClinVar classification (none available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.642678 | Disordered | 0.422910 | Uncertain | 0.293 | 0.861 | 0.750 | 6-33438064-G-A | -4.674 | Likely Benign | 0.089 | Likely Benign | Likely Benign | 2.37 | Destabilizing | 0.5 | 1.94 | Ambiguous | 2.16 | Destabilizing | 0.12 | Likely Benign | 0.359 | Likely Benign | -0.12 | Neutral | 0.000 | Benign | 0.002 | Benign | 1.33 | Pathogenic | 0.13 | Tolerated | 4.32 | 3 | 0.3051 | 0.4724 | 0 | 1 | -0.4 | 30.03 | ||||||||||||||||||||||||||
| c.1256A>G | E419G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E419G is listed in ClinVar with an uncertain significance (ClinVar ID 2004834.0) and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL and FATHMM, while pathogenic predictions are made by PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as likely pathogenic. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenic, SGM‑Consensus confirms likely pathogenic, and the Foldetta stability analysis is inconclusive. No evidence from FoldX, Rosetta, or premPS is available. Overall, the preponderance of predictions indicates that E419G is most likely pathogenic, which contrasts with the current ClinVar designation of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.102787 | Structured | 0.371949 | Uncertain | 0.961 | 0.261 | 0.000 | Uncertain | 1 | -10.589 | Likely Pathogenic | 0.956 | Likely Pathogenic | Likely Pathogenic | 1.41 | Ambiguous | 0.0 | 1.94 | Ambiguous | 1.68 | Ambiguous | 0.83 | Ambiguous | 0.469 | Likely Benign | -6.42 | Deleterious | 1.000 | Probably Damaging | 0.997 | Probably Damaging | 3.31 | Benign | 0.02 | Affected | 3.37 | 29 | 0.2992 | 0.5728 | 0 | -2 | 3.1 | -72.06 | 165.3 | 110.8 | 0.0 | 0.0 | -0.1 | 0.0 | X | Potentially Pathogenic | The carboxylate group of Glu419, located on an α helix (res. Met414-Glu436), forms a salt bridge with the side chain of either Arg716 or Lys418 from an opposing helix (res. Pro713-Arg726). The backbone amide group of Glu419 does not form H-bonds, resulting in a slight bend in the α helix. Thus, although glycine is known as an “α helix breaker,” the residue swap does not disrupt the continuity or integrity of the α helix. However, because Gly419 cannot form a salt bridge with the guanidinium group of the Arg716 side chain, the C2-GAP domain tertiary structure could be compromised during folding. | ||||||||||||||||
| c.1460A>C | N487T 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant N487T is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL and FATHMM, whereas the majority of tools predict a pathogenic impact: PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is classified as “Likely Pathogenic.” High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts pathogenic, SGM‑Consensus (majority vote) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is inconclusive. No evidence from FoldX, Rosetta, or premPS is available to alter this assessment. Overall, the preponderance of computational evidence indicates that the variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.209395 | Structured | 0.338511 | Uncertain | 0.890 | 0.243 | 0.125 | -12.618 | Likely Pathogenic | 0.981 | Likely Pathogenic | Likely Pathogenic | 1.92 | Ambiguous | 0.1 | 1.94 | Ambiguous | 1.93 | Ambiguous | 0.68 | Ambiguous | 0.481 | Likely Benign | -5.97 | Deleterious | 0.987 | Probably Damaging | 0.980 | Probably Damaging | 2.78 | Benign | 0.05 | Affected | 0.1200 | 0.3441 | 0 | 0 | 2.8 | -13.00 | |||||||||||||||||||||||||||||
| c.1701G>C | E567D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E567D missense variant is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, SIFT, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Default. Uncertain or unavailable results come from FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized classifying the variant as benign, while the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates a likely pathogenic outcome; Foldetta’s stability prediction is uncertain. Overall, the majority of evidence points toward a pathogenic impact, and this conclusion does not contradict the ClinVar status, which is currently unreported. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.021816 | Structured | 0.051008 | Uncertain | 0.916 | 0.234 | 0.000 | -8.276 | Likely Pathogenic | 0.575 | Likely Pathogenic | Likely Benign | 0.87 | Ambiguous | 0.1 | 1.94 | Ambiguous | 1.41 | Ambiguous | 0.70 | Ambiguous | 0.184 | Likely Benign | -2.72 | Deleterious | 0.996 | Probably Damaging | 0.989 | Probably Damaging | 3.41 | Benign | 0.21 | Tolerated | 0.1602 | 0.3426 | 3 | 2 | 0.0 | -14.03 | |||||||||||||||||||||||||||||
| c.1701G>T | E567D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E567D missense variant is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, SIFT, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic impact are SGM‑Consensus, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and AlphaMissense‑Default. Uncertain or unavailable results come from FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized classifying the variant as benign, while the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates a likely pathogenic outcome; Foldetta’s stability prediction is inconclusive. Overall, the majority of pathogenic predictions outweigh the benign ones, suggesting the variant is most likely pathogenic. This conclusion is not contradicted by ClinVar status, as no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.021816 | Structured | 0.051008 | Uncertain | 0.916 | 0.234 | 0.000 | -8.276 | Likely Pathogenic | 0.575 | Likely Pathogenic | Likely Benign | 0.87 | Ambiguous | 0.1 | 1.94 | Ambiguous | 1.41 | Ambiguous | 0.70 | Ambiguous | 0.184 | Likely Benign | -2.72 | Deleterious | 0.996 | Probably Damaging | 0.989 | Probably Damaging | 3.41 | Benign | 0.21 | Tolerated | 0.1602 | 0.3426 | 3 | 2 | 0.0 | -14.03 | |||||||||||||||||||||||||||||
| c.1718G>A | R573Q 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant R573Q is reported in ClinVar as Pathogenic (ClinVar ID 1176819.0) and is not present in gnomAD. Functional prediction tools largely agree on a deleterious effect: pathogenic predictions come from SGM‑Consensus, REVEL, FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, and AlphaMissense‑Default, while only SIFT predicts a benign outcome. Two tools give inconclusive results: Rosetta (Uncertain) and AlphaMissense‑Optimized (Uncertain). High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized remains uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is Pathogenic. Overall, the preponderance of evidence indicates the variant is most likely pathogenic, consistent with its ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.134866 | Structured | 0.032433 | Uncertain | 0.934 | 0.235 | 0.000 | Likely Pathogenic | 1 | -9.900 | Likely Pathogenic | 0.923 | Likely Pathogenic | Ambiguous | 2.28 | Destabilizing | 0.8 | 1.94 | Ambiguous | 2.11 | Destabilizing | 1.08 | Destabilizing | 0.733 | Likely Pathogenic | -3.16 | Deleterious | 1.000 | Probably Damaging | 0.995 | Probably Damaging | -1.31 | Pathogenic | 0.12 | Tolerated | 3.37 | 35 | 0.2390 | 0.1651 | 1 | 1 | 1.0 | -28.06 | 230.1 | 49.9 | 0.0 | 0.0 | -0.6 | 0.0 | X | X | Potentially Pathogenic | The guanidinium group of Arg573, located in an α-helix (res. Arg563-Glu578), forms a salt bridge with the carboxylate groups of Glu582 and/or Asp586 from a nearby α-helix (res. Glu582-Met603) in the WT simulations. Additionally, the Arg573 side chain stacks planarly with the aromatic phenol ring of Tyr665 and hydrogen bonds with the hydroxyl group of Ser668 from another α-helix (res. Ser641-Ser668). In the variant simulations, although the carboxamide group of the Gln573 side chain can hydrogen bond with the carboxylate group of Glu582 or the hydroxyl group of Ser668, these interactions are not as coordinated, stable, or strong as those of the positively charged Arg573. Consequently, the integrity of the opposing α-helix end (res. Glu582-Met603) is weakened. Overall, the residue swap has the potential to substantially affect the tertiary structure assembly during the protein folding process. | |||||||||||||||
| c.1754C>G | A585G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A585G is not reported in ClinVar and is absent from gnomAD. Prediction tools cluster into two groups: benign predictions come from REVEL, PROVEAN, SIFT, ESM1b, and AlphaMissense‑Optimized, while pathogenic predictions arise from polyPhen‑2 (HumDiv and HumVar), FATHMM, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as benign, whereas the SGM Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is inconclusive due to a 2‑to‑2 split, and Foldetta reports an uncertain stability change. No evidence from these high‑confidence tools supports pathogenicity. Overall, the balance of evidence favors a benign effect for A585G, and this conclusion is consistent with the lack of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.060549 | Structured | 0.055884 | Uncertain | 0.880 | 0.244 | 0.000 | -3.879 | Likely Benign | 0.629 | Likely Pathogenic | Likely Benign | 1.62 | Ambiguous | 0.0 | 1.94 | Ambiguous | 1.78 | Ambiguous | 0.59 | Ambiguous | 0.384 | Likely Benign | -1.16 | Neutral | 0.999 | Probably Damaging | 0.995 | Probably Damaging | -1.33 | Pathogenic | 0.24 | Tolerated | 0.1724 | 0.2299 | 1 | 0 | -2.2 | -14.03 | ||||||||||||||||||||||||||||||
| c.2048T>G | I683S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I683S is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are limited to FATHMM, whereas the majority of tools predict a pathogenic impact: SGM‑Consensus, REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. High‑accuracy methods give a consistent pathogenic signal: AlphaMissense‑Optimized is uncertain, SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is Pathogenic. No prediction or folding stability result is missing or inconclusive; all available evidence points to a deleterious effect. Based on the collective predictions, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which currently has no classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.200174 | Structured | 0.143268 | Uncertain | 0.848 | 0.314 | 0.000 | -11.443 | Likely Pathogenic | 0.950 | Likely Pathogenic | Ambiguous | 2.53 | Destabilizing | 0.2 | 1.94 | Ambiguous | 2.24 | Destabilizing | 1.35 | Destabilizing | 0.552 | Likely Pathogenic | -5.88 | Deleterious | 1.000 | Probably Damaging | 0.989 | Probably Damaging | 3.29 | Benign | 0.05 | Affected | 0.1936 | 0.1320 | -1 | -2 | -5.3 | -26.08 | |||||||||||||||||||||||||||||
| c.814C>G | R272G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R272G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: benign predictions are limited to REVEL, whereas pathogenic predictions are made by FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. Uncertain results come from Rosetta and AlphaMissense‑Optimized. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is inconclusive, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) reports “Likely Pathogenic,” and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts a destabilizing, pathogenic effect. Overall, the preponderance of evidence indicates the variant is most likely pathogenic, and this assessment is consistent with the absence of a ClinVar entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.071867 | Structured | 0.425620 | Uncertain | 0.925 | 0.215 | 0.125 | -11.540 | Likely Pathogenic | 0.831 | Likely Pathogenic | Ambiguous | 3.40 | Destabilizing | 0.0 | 1.94 | Ambiguous | 2.67 | Destabilizing | 1.06 | Destabilizing | 0.479 | Likely Benign | -4.57 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | 1.75 | Pathogenic | 0.01 | Affected | 0.3082 | 0.3062 | -3 | -2 | 4.1 | -99.14 | |||||||||||||||||||||||||||||
| c.1130T>C | M377T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M377T is reported in gnomAD (variant ID 6‑33438035‑T‑C) but has no ClinVar entry. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Only SIFT predicts a pathogenic outcome. Uncertain results are reported for FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as Likely Benign, and Foldetta as Uncertain. Overall, the majority of evidence points to a benign effect for M377T, and this conclusion does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.675549 | Disordered | 0.431183 | Uncertain | 0.324 | 0.884 | 0.625 | 6-33438035-T-C | 5 | 1.17e-5 | -1.881 | Likely Benign | 0.090 | Likely Benign | Likely Benign | 0.90 | Ambiguous | 0.4 | 1.95 | Ambiguous | 1.43 | Ambiguous | 0.59 | Ambiguous | 0.245 | Likely Benign | -0.65 | Neutral | 0.000 | Benign | 0.002 | Benign | 5.47 | Benign | 0.05 | Affected | 4.32 | 12 | 0.3064 | 0.3267 | -1 | -1 | -2.6 | -30.09 | ||||||||||||||||||||||||
| c.1694T>A | L565Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L565Q is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only FATHMM, whereas the remaining tools—SGM‑Consensus, REVEL, FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—consistently predict a pathogenic or likely pathogenic impact; Rosetta is uncertain. High‑accuracy methods give the following results: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates likely pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. No prediction or stability result is missing or inconclusive. Based on the overwhelming agreement among pathogenic predictors and the high‑accuracy tools, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.034884 | Structured | 0.045819 | Uncertain | 0.922 | 0.205 | 0.000 | -13.912 | Likely Pathogenic | 0.996 | Likely Pathogenic | Likely Pathogenic | 2.86 | Destabilizing | 0.1 | 1.95 | Ambiguous | 2.41 | Destabilizing | 2.39 | Destabilizing | 0.545 | Likely Pathogenic | -5.97 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 2.76 | Benign | 0.00 | Affected | 0.1149 | 0.0972 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.1847A>T | D616V 2D  3DClick to see structure in 3D Viewer AISynGAP1 D616V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign calls come from REVEL, premPS, and FATHMM, while pathogenic calls are made by FoldX, Foldetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and the SGM‑Consensus. Uncertain results are reported by Rosetta and AlphaMissense‑Optimized. High‑accuracy assessments give a pathogenic signal: AlphaMissense‑Optimized is inconclusive, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. Overall, the majority of evidence, including the high‑accuracy tools, supports a pathogenic effect for D616V. This conclusion is not contradicted by ClinVar, which has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.129801 | Structured | 0.166689 | Uncertain | 0.867 | 0.252 | 0.000 | -13.992 | Likely Pathogenic | 0.919 | Likely Pathogenic | Ambiguous | 2.41 | Destabilizing | 0.2 | 1.95 | Ambiguous | 2.18 | Destabilizing | 0.36 | Likely Benign | 0.268 | Likely Benign | -7.36 | Deleterious | 0.972 | Probably Damaging | 0.682 | Possibly Damaging | 3.26 | Benign | 0.00 | Affected | 0.0699 | 0.4393 | -2 | -3 | 7.7 | -15.96 | |||||||||||||||||||||||||||||
| c.2065C>T | L689F 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L689F is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include REVEL and FATHMM, whereas a majority of tools predict a pathogenic impact: FoldX, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus. Two tools (Rosetta and premPS) yield uncertain results. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM Consensus is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. No prediction or stability result is missing or inconclusive. Based on the preponderance of pathogenic predictions, the variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.042364 | Structured | 0.227227 | Uncertain | 0.963 | 0.248 | 0.000 | -9.817 | Likely Pathogenic | 0.978 | Likely Pathogenic | Likely Pathogenic | 2.45 | Destabilizing | 0.2 | 1.95 | Ambiguous | 2.20 | Destabilizing | 0.67 | Ambiguous | 0.286 | Likely Benign | -3.98 | Deleterious | 0.999 | Probably Damaging | 0.860 | Possibly Damaging | 3.18 | Benign | 0.00 | Affected | 0.0608 | 0.2891 | 2 | 0 | -1.0 | 34.02 | |||||||||||||||||||||||||||||
| c.599T>G | L200W 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L200W is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL and FATHMM, whereas the majority of tools (SGM‑Consensus, FoldX, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, and Foldetta) predict a pathogenic impact. Two tools (AlphaMissense‑Optimized and Rosetta) provide inconclusive results and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as Likely Pathogenic (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN), and Foldetta as Pathogenic, the latter integrating stability predictions from FoldX‑MD and Rosetta. Overall, the preponderance of evidence indicates that the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.366687 | Structured | 0.428168 | Uncertain | 0.687 | 0.453 | 0.125 | -11.550 | Likely Pathogenic | 0.824 | Likely Pathogenic | Ambiguous | 2.90 | Destabilizing | 1.0 | 1.95 | Ambiguous | 2.43 | Destabilizing | 1.20 | Destabilizing | 0.200 | Likely Benign | -2.93 | Deleterious | 0.999 | Probably Damaging | 0.970 | Probably Damaging | 3.98 | Benign | 0.02 | Affected | 0.0627 | 0.2893 | -2 | -2 | -4.7 | 73.05 | |||||||||||||||||||||||||||||
| c.1241T>C | M414T 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant M414T is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, and FATHMM, whereas pathogenic predictions arise from SGM‑Consensus, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and Foldetta; Rosetta remains uncertain. High‑accuracy methods all support a deleterious effect: AlphaMissense‑Optimized scores the variant as pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates likely pathogenic, and Foldetta predicts a destabilizing, pathogenic outcome. Consequently, the consensus of the most reliable predictors classifies M414T as pathogenic, with no conflict with the ClinVar status, which simply lacks an entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.081712 | Structured | 0.329108 | Uncertain | 0.914 | 0.217 | 0.000 | -8.698 | Likely Pathogenic | 0.975 | Likely Pathogenic | Likely Pathogenic | 2.04 | Destabilizing | 0.1 | 1.96 | Ambiguous | 2.00 | Destabilizing | 1.27 | Destabilizing | 0.406 | Likely Benign | -5.00 | Deleterious | 0.997 | Probably Damaging | 0.972 | Probably Damaging | 3.41 | Benign | 0.08 | Tolerated | 0.1771 | 0.1976 | -1 | -1 | -2.6 | -30.09 | |||||||||||||||||||||||||||||
| c.1886T>C | V629A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 V629A missense variant is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, SIFT, and FATHMM. Those that predict a pathogenic effect comprise SGM‑Consensus, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and AlphaMissense‑Default. Two tools give uncertain results: Rosetta and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as Likely Pathogenic (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN), and Foldetta as Pathogenic (combining FoldX‑MD and Rosetta outputs). Overall, the majority of predictions (8 pathogenic vs. 3 benign) indicate that V629A is most likely pathogenic, and this conclusion does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.040537 | Structured | 0.034796 | Uncertain | 0.970 | 0.236 | 0.000 | -8.652 | Likely Pathogenic | 0.926 | Likely Pathogenic | Ambiguous | 2.24 | Destabilizing | 0.1 | 1.96 | Ambiguous | 2.10 | Destabilizing | 1.58 | Destabilizing | 0.492 | Likely Benign | -3.58 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.18 | Benign | 0.11 | Tolerated | 0.2518 | 0.2124 | 0 | 0 | -2.4 | -28.05 | |||||||||||||||||||||||||||||
| c.1947G>A | M649I 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant M649I has no ClinVar entry and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, polyPhen‑2 HumVar, and FATHMM, whereas the majority of tools (SGM‑Consensus, FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic impact; Rosetta is inconclusive and is not counted. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as pathogenic. Taken together, the preponderance of evidence supports a pathogenic classification for M649I, and this conclusion does not conflict with ClinVar status, which is currently unavailable. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.051831 | Structured | 0.360413 | Uncertain | 0.962 | 0.345 | 0.000 | -9.361 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 2.42 | Destabilizing | 0.2 | 1.96 | Ambiguous | 2.19 | Destabilizing | 1.01 | Destabilizing | 0.449 | Likely Benign | -3.99 | Deleterious | 0.672 | Possibly Damaging | 0.093 | Benign | 3.40 | Benign | 0.02 | Affected | 3.38 | 27 | 0.1215 | 0.2980 | 2 | 1 | 2.6 | -18.03 | 243.7 | 21.5 | 0.0 | 0.1 | 0.0 | 0.1 | X | Potentially Benign | The thioether side chain of Met649, located on an α helix (res. Ser641-Glu666), bridges Phe652, Phe648, and Phe639 in an inter-helix hydrophobic cavity in the WT simulations. In the variant simulations, the sec-butyl side chain of Ile649 maintains hydrophobic interactions with nearby residues, with no significant effects on the protein structure.However, methionine is known as a bridging motif for aromatic residues, and these Met-aromatic interactions are lost in the variant. Indeed, in the second variant simulation,the bridging of Phe652, Phe648 and Phe639 is completely lost. In reality, the effect could be more severe on the structure during the protein folding. | ||||||||||||||||||
| c.1947G>C | M649I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M649I is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that indicate a benign effect include REVEL, polyPhen‑2 HumVar, and FATHMM, whereas the majority of other in silico predictors (FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) report a pathogenic outcome; Rosetta is inconclusive. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is “Likely Pathogenic,” and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenicity. Overall, the preponderance of evidence points to a pathogenic effect for M649I, which is consistent with the ClinVar “Uncertain” classification rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.051831 | Structured | 0.360413 | Uncertain | 0.962 | 0.345 | 0.000 | Uncertain | 1 | -9.361 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 2.42 | Destabilizing | 0.2 | 1.96 | Ambiguous | 2.19 | Destabilizing | 1.01 | Destabilizing | 0.449 | Likely Benign | -3.99 | Deleterious | 0.672 | Possibly Damaging | 0.093 | Benign | 3.40 | Benign | 0.02 | Affected | 3.38 | 27 | 0.1215 | 0.2980 | 2 | 1 | 2.6 | -18.03 | 243.7 | 21.5 | 0.0 | 0.1 | 0.0 | 0.1 | X | Potentially Benign | The thioether side chain of Met649, located on an α helix (res. Ser641-Glu666), bridges Phe652, Phe648, and Phe639 in an inter-helix hydrophobic cavity in the WT simulations. In the variant simulations, the sec-butyl side chain of Ile649 maintains hydrophobic interactions with nearby residues, with no significant effects on the protein structure.However, methionine is known as a bridging motif for aromatic residues, and these Met-aromatic interactions are lost in the variant. Indeed, in the second variant simulation,the bridging of Phe652, Phe648 and Phe639 is completely lost. In reality, the effect could be more severe on the structure during the protein folding. | ||||||||||||||||
| c.1947G>T | M649I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M649I is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign calls come from REVEL, polyPhen‑2 HumVar, and FATHMM, whereas pathogenic calls are made by FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, votes strongly for pathogenicity (3/4 pathogenic). High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenic, the SGM‑Consensus (majority vote) is likely pathogenic, and Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, also predicts pathogenic. Rosetta alone is uncertain and is treated as unavailable. Overall, the majority of evidence points to a pathogenic impact, and this conclusion does not contradict any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.051831 | Structured | 0.360413 | Uncertain | 0.962 | 0.345 | 0.000 | -9.361 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 2.42 | Destabilizing | 0.2 | 1.96 | Ambiguous | 2.19 | Destabilizing | 1.01 | Destabilizing | 0.449 | Likely Benign | -3.99 | Deleterious | 0.672 | Possibly Damaging | 0.093 | Benign | 3.40 | Benign | 0.02 | Affected | 3.38 | 27 | 0.1215 | 0.2980 | 2 | 1 | 2.6 | -18.03 | 243.7 | 21.5 | 0.0 | 0.1 | 0.0 | 0.1 | X | Potentially Benign | The thioether side chain of Met649, located on an α helix (res. Ser641-Glu666), bridges Phe652, Phe648, and Phe639 in an inter-helix hydrophobic cavity in the WT simulations. In the variant simulations, the sec-butyl side chain of Ile649 maintains hydrophobic interactions with nearby residues, with no significant effects on the protein structure.However, methionine is known as a bridging motif for aromatic residues, and these Met-aromatic interactions are lost in the variant. Indeed, in the second variant simulation,the bridging of Phe652, Phe648 and Phe639 is completely lost. In reality, the effect could be more severe on the structure during the protein folding. | ||||||||||||||||||
| c.630C>A | H210Q 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant H210Q is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show mixed results: benign calls come from REVEL, FoldX, polyPhen‑2 (HumDiv and HumVar) and FATHMM, whereas pathogenic calls are made by premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default and AlphaMissense‑Optimized. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM and PROVEAN, favors a pathogenic outcome (3/4). High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts pathogenic, and the SGM‑Consensus (majority vote) also predicts pathogenic. Foldetta, a protein‑folding stability predictor that integrates FoldX‑MD and Rosetta, is inconclusive and therefore not used as evidence. Overall, the majority of reliable predictors indicate a pathogenic effect for H210Q, and this conclusion does not conflict with any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.144935 | Structured | 0.390904 | Uncertain | 0.872 | 0.298 | 0.125 | -12.639 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 0.26 | Likely Benign | 0.3 | 1.96 | Ambiguous | 1.11 | Ambiguous | 1.20 | Destabilizing | 0.258 | Likely Benign | -6.84 | Deleterious | 0.141 | Benign | 0.064 | Benign | 3.10 | Benign | 0.00 | Affected | 0.1016 | 0.2852 | 3 | 0 | -0.3 | -9.01 | |||||||||||||||||||||||||||||
| c.630C>G | H210Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant H210Q is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign calls from REVEL, FoldX, polyPhen‑2 (HumDiv and HumVar) and FATHMM, whereas pathogenic calls come from premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default and AlphaMissense‑Optimized. When predictions are grouped, five tools favor a benign effect and six favor a pathogenic effect. High‑accuracy assessments further support a deleterious outcome: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates pathogenic, and the Foldetta stability analysis is inconclusive. No evidence from ClinVar contradicts these findings. Therefore, the variant is most likely pathogenic based on the aggregate computational evidence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.144935 | Structured | 0.390904 | Uncertain | 0.872 | 0.298 | 0.125 | -12.639 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 0.26 | Likely Benign | 0.3 | 1.96 | Ambiguous | 1.11 | Ambiguous | 1.20 | Destabilizing | 0.258 | Likely Benign | -6.84 | Deleterious | 0.141 | Benign | 0.064 | Benign | 3.10 | Benign | 0.00 | Affected | 0.1016 | 0.2852 | 3 | 0 | -0.3 | -9.01 | |||||||||||||||||||||||||||||
| c.755C>A | P252H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P252H missense variant has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include only FATHMM, while the remaining evaluated algorithms (SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) uniformly predict a pathogenic impact. High‑accuracy assessments further support this: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is “Likely Pathogenic”; and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is uncertain. No other high‑confidence stability predictions are available. Overall, the preponderance of evidence indicates that P252H is most likely pathogenic, and this conclusion is not contradicted by ClinVar status, which is currently absent. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.461924 | Structured | 0.211606 | Uncertain | 0.753 | 0.304 | 0.250 | -11.991 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 1.06 | Ambiguous | 0.2 | 1.96 | Ambiguous | 1.51 | Ambiguous | 0.82 | Ambiguous | 0.845 | Likely Pathogenic | -8.27 | Deleterious | 0.999 | Probably Damaging | 0.936 | Probably Damaging | 5.76 | Benign | 0.00 | Affected | 0.1592 | 0.4128 | 0 | -2 | -1.6 | 40.02 | |||||||||||||||||||||||||||||
| c.1049T>C | V350A 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant V350A is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN), premPS, PROVEAN, ESM1b, and FATHMM. Stability‑based methods FoldX, Rosetta, and Foldetta give uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as pathogenic, and Foldetta as unavailable. Overall, the predictions are split, with a slight edge toward pathogenicity from the consensus and high‑accuracy tools. Therefore, the variant is most likely pathogenic based on the current computational evidence, and this assessment does not contradict any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.179055 | Structured | 0.353227 | Uncertain | 0.931 | 0.371 | 0.000 | -8.323 | Likely Pathogenic | 0.280 | Likely Benign | Likely Benign | 1.42 | Ambiguous | 0.0 | 1.97 | Ambiguous | 1.70 | Ambiguous | 2.07 | Destabilizing | 0.096 | Likely Benign | -2.73 | Deleterious | 0.435 | Benign | 0.115 | Benign | 1.64 | Pathogenic | 0.55 | Tolerated | 0.3207 | 0.2967 | 0 | 0 | -2.4 | -28.05 | |||||||||||||||||||||||||||||
| c.1073T>C | F358S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 F358S variant has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, SIFT, and FATHMM, whereas a majority of algorithms predict a pathogenic outcome: FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote). Rosetta’s assessment is uncertain and is not taken as evidence. High‑accuracy methods give a consistent pathogenic signal: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. Based on the preponderance of pathogenic predictions and the agreement of the high‑accuracy tools, the variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.222385 | Structured | 0.407113 | Uncertain | 0.912 | 0.441 | 0.250 | -9.316 | Likely Pathogenic | 0.977 | Likely Pathogenic | Likely Pathogenic | 2.32 | Destabilizing | 0.2 | 1.97 | Ambiguous | 2.15 | Destabilizing | 1.14 | Destabilizing | 0.493 | Likely Benign | -6.48 | Deleterious | 0.998 | Probably Damaging | 0.986 | Probably Damaging | 4.07 | Benign | 0.20 | Tolerated | 0.3891 | 0.1333 | -3 | -2 | -3.6 | -60.10 | |||||||||||||||||||||||||||||
| c.1342G>T | A448S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 A448S missense variant is not reported in ClinVar (status: None) and has no entry in gnomAD. Prediction tools that indicate a benign effect include REVEL, SIFT, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect comprise SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Default. Uncertain or inconclusive results are reported for FoldX, Rosetta, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely pathogenic (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN), and Foldetta as uncertain. Overall, the majority of predictions lean toward pathogenicity, and this conclusion does not contradict any ClinVar annotation because none exists. Thus, the variant is most likely pathogenic based on the available computational evidence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.225814 | Structured | 0.292774 | Uncertain | 0.973 | 0.257 | 0.000 | -9.213 | Likely Pathogenic | 0.590 | Likely Pathogenic | Likely Benign | 1.18 | Ambiguous | 0.1 | 1.97 | Ambiguous | 1.58 | Ambiguous | 0.55 | Ambiguous | 0.310 | Likely Benign | -2.96 | Deleterious | 0.965 | Probably Damaging | 0.972 | Probably Damaging | 3.27 | Benign | 0.06 | Tolerated | 0.2420 | 0.3471 | 1 | 1 | -2.6 | 16.00 | |||||||||||||||||||||||||||||
| c.1753G>A | A585T 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant A585T is reported in gnomAD (ID 6‑33440805‑G‑A) but has no ClinVar entry. Functional prediction tools cluster into two groups: benign predictions come from REVEL, premPS, PROVEAN, and SIFT, while pathogenic predictions arise from polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, and AlphaMissense‑Default. Four tools (FoldX, Rosetta, Foldetta, AlphaMissense‑Optimized) give uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the balance of evidence favors a pathogenic effect for A585T. This conclusion is not contradicted by ClinVar, which contains no classification for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.060549 | Structured | 0.055884 | Uncertain | 0.880 | 0.244 | 0.000 | 6-33440805-G-A | 13 | 8.05e-6 | -10.063 | Likely Pathogenic | 0.876 | Likely Pathogenic | Ambiguous | 1.66 | Ambiguous | 0.2 | 1.97 | Ambiguous | 1.82 | Ambiguous | 0.23 | Likely Benign | 0.465 | Likely Benign | -1.73 | Neutral | 1.000 | Probably Damaging | 0.994 | Probably Damaging | -1.30 | Pathogenic | 0.26 | Tolerated | 3.37 | 35 | 0.1132 | 0.4212 | 0 | 1 | -2.5 | 30.03 | ||||||||||||||||||||||||
| c.725G>C | W242S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant W242S, located in the PH domain, is not reported in ClinVar and is absent from gnomAD. Prediction tools that assess pathogenicity unanimously classify the variant as deleterious: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict a pathogenic effect. No tool predicts a benign outcome. High‑accuracy assessments further support a harmful impact: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is “Likely Pathogenic,” while Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, yields an uncertain result. Overall, the consensus of available predictions indicates that W242S is most likely pathogenic, and this conclusion does not contradict any ClinVar annotation (none is present). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.328603 | Structured | 0.352582 | Uncertain | 0.847 | 0.341 | 0.000 | -14.674 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 1.96 | Ambiguous | 0.2 | 1.97 | Ambiguous | 1.97 | Ambiguous | 0.83 | Ambiguous | 0.797 | Likely Pathogenic | -12.71 | Deleterious | 0.900 | Possibly Damaging | 0.535 | Possibly Damaging | 1.52 | Pathogenic | 0.00 | Affected | 0.4424 | 0.1418 | -2 | -3 | 0.1 | -99.14 | |||||||||||||||||||||||||||||
| c.1482A>G | I494M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 I494M missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include FoldX, PROVEAN, SIFT, and AlphaMissense‑Optimized. Those that agree on a pathogenic effect are REVEL, polyPhen‑2 (HumDiv and HumVar), ESM1b, and FATHMM. The remaining tools—Rosetta, Foldetta, premPS, and AlphaMissense‑Default—return uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the balance of evidence leans toward pathogenicity, with a majority of tools predicting a deleterious effect. This conclusion does not contradict the ClinVar status, which currently has no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.155435 | Structured | 0.353330 | Uncertain | 0.941 | 0.157 | 0.000 | -8.223 | Likely Pathogenic | 0.356 | Ambiguous | Likely Benign | 0.35 | Likely Benign | 0.2 | 1.98 | Ambiguous | 1.17 | Ambiguous | 0.98 | Ambiguous | 0.542 | Likely Pathogenic | -2.25 | Neutral | 1.000 | Probably Damaging | 0.997 | Probably Damaging | -1.15 | Pathogenic | 0.23 | Tolerated | 0.0676 | 0.1789 | 2 | 1 | -2.6 | 18.03 | ||||||||||||||||||||||||||||||
| c.655T>G | C219G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C219G is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect are limited to FATHMM, whereas the remaining 11 tools—including REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus—consistently predict a pathogenic impact. High‑accuracy assessments further support this view: AlphaMissense‑Optimized returns a pathogenic score, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates pathogenicity, and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, is inconclusive. Taken together, the overwhelming majority of evidence points to a pathogenic effect for C219G. This conclusion aligns with the absence of a ClinVar entry and does not contradict any existing clinical annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.254060 | Structured | 0.426845 | Uncertain | 0.903 | 0.279 | 0.000 | -15.072 | Likely Pathogenic | 0.996 | Likely Pathogenic | Likely Pathogenic | 0.72 | Ambiguous | 0.3 | 1.98 | Ambiguous | 1.35 | Ambiguous | 1.18 | Destabilizing | 0.900 | Likely Pathogenic | -10.09 | Deleterious | 0.900 | Possibly Damaging | 0.461 | Possibly Damaging | 5.83 | Benign | 0.02 | Affected | 0.2159 | 0.2169 | -3 | -3 | -2.9 | -46.09 | |||||||||||||||||||||||||||||
| c.1234T>A | L412M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L412M has no ClinVar entry and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, PROVEAN, and FATHMM, while pathogenic predictions arise from polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. Five tools give uncertain or inconclusive results (FoldX, Rosetta, Foldetta, premPS, AlphaMissense‑Optimized). High‑accuracy assessments are likewise inconclusive: AlphaMissense‑Optimized is uncertain; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is a 2‑vs‑2 tie; and Foldetta is uncertain. Consequently, the overall evidence leans toward a pathogenic effect, with no ClinVar record to contradict this assessment. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.122885 | Structured | 0.331108 | Uncertain | 0.937 | 0.196 | 0.000 | -9.342 | Likely Pathogenic | 0.944 | Likely Pathogenic | Ambiguous | 0.75 | Ambiguous | 0.0 | 1.99 | Ambiguous | 1.37 | Ambiguous | 0.86 | Ambiguous | 0.246 | Likely Benign | -1.84 | Neutral | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.25 | Benign | 0.05 | Affected | 0.0746 | 0.3482 | 4 | 2 | -1.9 | 18.03 | ||||||||||||||||||||||||||||||
| c.1416G>C | E472D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E472D is not reported in ClinVar and has no gnomAD entry. Consensus from multiple in‑silico predictors shows a split: benign calls come from REVEL and SIFT, whereas the majority of tools—SGM‑Consensus, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized—predict pathogenicity. FoldX, Rosetta, and Foldetta provide uncertain or inconclusive stability estimates. High‑accuracy methods reinforce the pathogenic signal: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (derived from the four high‑confidence predictors) is pathogenic, and Foldetta remains uncertain. Overall, the preponderance of evidence points to a pathogenic effect for E472D, and this assessment does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.264545 | Structured | 0.359300 | Uncertain | 0.878 | 0.231 | 0.000 | -9.798 | Likely Pathogenic | 0.992 | Likely Pathogenic | Likely Pathogenic | 1.27 | Ambiguous | 0.3 | 1.99 | Ambiguous | 1.63 | Ambiguous | 1.00 | Destabilizing | 0.304 | Likely Benign | -2.75 | Deleterious | 0.989 | Probably Damaging | 0.979 | Probably Damaging | 2.43 | Pathogenic | 0.06 | Tolerated | 0.1975 | 0.3918 | 3 | 2 | 0.0 | -14.03 | |||||||||||||||||||||||||||||
| c.1416G>T | E472D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E472D is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL and SIFT, whereas a majority of tools predict a pathogenic impact: polyPhen‑2 (HumDiv and HumVar), FATHMM, ESM1b, PROVEAN, AlphaMissense‑Default, AlphaMissense‑Optimized, premPS, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Stability‑based methods (FoldX, Rosetta, Foldetta) yield uncertain results and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus as Likely Pathogenic, and Foldetta as uncertain. Overall, the preponderance of evidence from multiple independent predictors indicates that the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.264545 | Structured | 0.359300 | Uncertain | 0.878 | 0.231 | 0.000 | -9.798 | Likely Pathogenic | 0.992 | Likely Pathogenic | Likely Pathogenic | 1.27 | Ambiguous | 0.3 | 1.99 | Ambiguous | 1.63 | Ambiguous | 1.00 | Destabilizing | 0.303 | Likely Benign | -2.75 | Deleterious | 0.989 | Probably Damaging | 0.979 | Probably Damaging | 2.43 | Pathogenic | 0.06 | Tolerated | 0.1975 | 0.3918 | 3 | 2 | 0.0 | -14.03 | |||||||||||||||||||||||||||||
| c.1462A>G | T488A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant T488A is not reported in ClinVar and is absent from gnomAD. Prediction tools cluster into two groups: benign predictions come from REVEL and FATHMM, while the majority of tools (FoldX, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus) predict pathogenicity; Rosetta and premPS are uncertain. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. Taken together, the preponderance of evidence indicates that T488A is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.206376 | Structured | 0.332663 | Uncertain | 0.928 | 0.233 | 0.125 | -11.341 | Likely Pathogenic | 0.963 | Likely Pathogenic | Likely Pathogenic | 2.12 | Destabilizing | 0.2 | 1.99 | Ambiguous | 2.06 | Destabilizing | 0.57 | Ambiguous | 0.497 | Likely Benign | -4.64 | Deleterious | 0.996 | Probably Damaging | 0.989 | Probably Damaging | 3.24 | Benign | 0.01 | Affected | 0.2871 | 0.2872 | 1 | 0 | 2.5 | -30.03 | |||||||||||||||||||||||||||||
| c.1529T>C | I510T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I510T is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only AlphaMissense‑Optimized. All other evaluated algorithms—SGM‑Consensus, REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and Foldetta—classify the variant as pathogenic, while Rosetta remains uncertain. High‑accuracy assessments further support a pathogenic interpretation: AlphaMissense‑Optimized predicts benign, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts likely pathogenic, and Foldetta predicts pathogenic. Consequently, the variant is most likely pathogenic based on the collective predictions, and this assessment does not contradict any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.025762 | Structured | 0.250630 | Uncertain | 0.945 | 0.273 | 0.000 | -9.993 | Likely Pathogenic | 0.701 | Likely Pathogenic | Likely Benign | 3.08 | Destabilizing | 0.2 | 1.99 | Ambiguous | 2.54 | Destabilizing | 1.95 | Destabilizing | 0.914 | Likely Pathogenic | -3.63 | Deleterious | 1.000 | Probably Damaging | 0.997 | Probably Damaging | -1.43 | Pathogenic | 0.00 | Affected | 0.0960 | 0.0440 | 0 | -1 | -5.2 | -12.05 | |||||||||||||||||||||||||||||
| c.603T>A | D201E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D201E missense variant (ClinVar ID 3004688.0) is classified as **Benign** in ClinVar and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote). Only polyPhen‑2 HumDiv predicts a pathogenic outcome, while Rosetta, Foldetta, and AlphaMissense‑Default are uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as **Benign**, the SGM‑Consensus as **Likely Benign**, and Foldetta as **Uncertain**. Taken together, the overwhelming majority of evidence points to a benign impact, and this conclusion aligns with the ClinVar designation. Thus, the variant is most likely benign, with no contradiction to its ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | PH | 0.366687 | Structured | 0.428570 | Uncertain | 0.698 | 0.447 | 0.125 | Benign | 1 | -2.640 | Likely Benign | 0.406 | Ambiguous | Likely Benign | 0.42 | Likely Benign | 0.2 | 1.99 | Ambiguous | 1.21 | Ambiguous | 0.23 | Likely Benign | 0.165 | Likely Benign | -0.69 | Neutral | 0.633 | Possibly Damaging | 0.108 | Benign | 4.30 | Benign | 1.00 | Tolerated | 3.46 | 9 | 0.1069 | 0.5505 | 3 | 2 | 0.0 | 14.03 | 258.7 | -24.8 | 0.9 | 0.1 | -0.3 | 0.2 | X | Uncertain | Asp201, an acidic residue located in the N-terminal loop before the first anti-parallel β sheet strand (res. Ile205-Pro208), is replaced by another acidic residue, glutamate. The carboxylate groups of both Asp201 and Glu201 side chains form hydrogen bonds with the hydroxyl group of Ser221 in the simulations. Due to its shorter side chain, Asp201 can also hydrogen bond with the backbone amide groups of neighboring loop residues Ser204 and Asp203. However, since the model ends abruptly at the N-terminus, no definite conclusions can be drawn from the simulations. | ||||||||||||||||
| c.603T>G | D201E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D201E missense variant (gnomAD ID 6‑33435245‑T‑G) is listed in ClinVar with an uncertain significance. Across a broad panel of in silico predictors, the majority indicate a benign effect: REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, and AlphaMissense‑Optimized all score benign. Only polyPhen‑2 HumDiv predicts pathogenicity, while Rosetta, Foldetta, and AlphaMissense‑Default remain inconclusive. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, resolves to “likely benign.” High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized is benign, SGM‑Consensus is likely benign, and Foldetta is inconclusive. Taken together, the preponderance of evidence points to a benign impact, which does not conflict with the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | PH | 0.366687 | Structured | 0.428570 | Uncertain | 0.698 | 0.447 | 0.125 | Conflicting | 2 | 6-33435245-T-G | 20 | 1.24e-5 | -2.640 | Likely Benign | 0.406 | Ambiguous | Likely Benign | 0.42 | Likely Benign | 0.2 | 1.99 | Ambiguous | 1.21 | Ambiguous | 0.23 | Likely Benign | 0.165 | Likely Benign | -0.69 | Neutral | 0.633 | Possibly Damaging | 0.108 | Benign | 4.30 | Benign | 1.00 | Tolerated | 3.46 | 9 | 0.1069 | 0.5505 | 3 | 2 | 0.0 | 14.03 | 258.7 | -24.8 | 0.9 | 0.1 | -0.3 | 0.2 | X | Uncertain | Asp201, an acidic residue located in the N-terminal loop before the first anti-parallel β sheet strand (res. Ile205-Pro208), is replaced by another acidic residue, glutamate. The carboxylate groups of both Asp201 and Glu201 side chains form hydrogen bonds with the hydroxyl group of Ser221 in the simulations. Due to its shorter side chain, Asp201 can also hydrogen bond with the backbone amide groups of neighboring loop residues Ser204 and Asp203. However, since the model ends abruptly at the N-terminus, no definite conclusions can be drawn from the simulations. | |||||||||||||
| c.806T>G | I269R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I269R is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: the benign group contains only SIFT, whereas the pathogenic group includes SGM‑Consensus (Likely Pathogenic), REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. FoldX and Rosetta give uncertain results, and Foldetta is also uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, SGM‑Consensus as Likely Pathogenic, and Foldetta as unavailable. Based on the preponderance of pathogenic predictions, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.216401 | Structured | 0.343787 | Uncertain | 0.937 | 0.244 | 0.125 | -14.567 | Likely Pathogenic | 0.994 | Likely Pathogenic | Likely Pathogenic | 1.34 | Ambiguous | 0.1 | 1.99 | Ambiguous | 1.67 | Ambiguous | 1.50 | Destabilizing | 0.765 | Likely Pathogenic | -5.23 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | 1.69 | Pathogenic | 0.07 | Tolerated | 0.1108 | 0.0870 | -2 | -3 | -9.0 | 43.03 | |||||||||||||||||||||||||||||
| c.1180A>C | K394Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 K394Q missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. Those that predict a pathogenic outcome are SIFT and Rosetta. The remaining tools—Foldetta, premPS, ESM1b, and AlphaMissense‑Default—return uncertain results and are treated as unavailable evidence. High‑accuracy assessments show AlphaMissense‑Optimized classifying the variant as benign, while the SGM Consensus (derived from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also leans toward benign, with two benign votes and two uncertain votes. Foldetta’s stability prediction is uncertain and thus not considered. Overall, the majority of reliable predictions indicate a benign effect, and this conclusion does not contradict any existing ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.505461 | Disordered | 0.399336 | Uncertain | 0.387 | 0.634 | 0.625 | -7.261 | In-Between | 0.468 | Ambiguous | Likely Benign | 0.15 | Likely Benign | 0.0 | 2.00 | Destabilizing | 1.08 | Ambiguous | 0.64 | Ambiguous | 0.330 | Likely Benign | -2.46 | Neutral | 0.001 | Benign | 0.009 | Benign | 4.61 | Benign | 0.01 | Affected | 0.5365 | 0.2106 | Weaken | 1 | 1 | 0.4 | -0.04 | |||||||||||||||||||||||||||||
| c.1469C>G | A490G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A490G is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only AlphaMissense‑Optimized. Those that predict a pathogenic effect comprise REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and the SGM‑Consensus (Likely Pathogenic). Predictions that are uncertain or inconclusive are FoldX, Foldetta, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points to a pathogenic impact for A490G. This conclusion does not contradict ClinVar status, as no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.120615 | Structured | 0.322979 | Uncertain | 0.938 | 0.210 | 0.125 | -9.767 | Likely Pathogenic | 0.384 | Ambiguous | Likely Benign | 1.24 | Ambiguous | 0.0 | 2.00 | Destabilizing | 1.62 | Ambiguous | 1.13 | Destabilizing | 0.744 | Likely Pathogenic | -3.44 | Deleterious | 0.999 | Probably Damaging | 0.995 | Probably Damaging | -1.46 | Pathogenic | 0.01 | Affected | 0.2051 | 0.2228 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.1570T>G | C524G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense change at residue 524 (C524G) is not reported in ClinVar (ClinVar ID None) and has no entry in gnomAD (gnomAD ID None). Prediction tools that assess pathogenicity unanimously classify the variant as deleterious: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict pathogenicity, while FoldX and Foldetta are uncertain. No tool predicts a benign effect. High‑accuracy assessments further support a damaging outcome: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is “Likely Pathogenic”; Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, remains uncertain. Overall, the evidence strongly favors a pathogenic impact for C524G, and this conclusion does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.134866 | Structured | 0.024729 | Uncertain | 0.916 | 0.385 | 0.125 | -13.597 | Likely Pathogenic | 0.985 | Likely Pathogenic | Likely Pathogenic | 1.07 | Ambiguous | 0.2 | 2.00 | Destabilizing | 1.54 | Ambiguous | 1.66 | Destabilizing | 0.924 | Likely Pathogenic | -11.93 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.40 | Pathogenic | 0.00 | Affected | 0.3579 | 0.2745 | -3 | -3 | -2.9 | -46.09 | |||||||||||||||||||||||||||||
| c.1832T>G | M611R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M611R is not reported in ClinVar (ClinVar ID None) and has no entries in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are AlphaMissense‑Optimized, polyPhen2_HumVar, and SIFT. Tools that agree on a pathogenic effect include SGM‑Consensus, REVEL, Rosetta, premPS, PROVEAN, polyPhen2_HumDiv, ESM1b, FATHMM, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized predicts a benign outcome, whereas the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is reported as uncertain. No prediction or folding result is missing; all available data are considered. Based on the overall distribution of predictions, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.236433 | Structured | 0.210791 | Uncertain | 0.870 | 0.253 | 0.000 | -11.050 | Likely Pathogenic | 0.642 | Likely Pathogenic | Likely Benign | 1.80 | Ambiguous | 0.8 | 2.00 | Destabilizing | 1.90 | Ambiguous | 1.58 | Destabilizing | 0.644 | Likely Pathogenic | -4.10 | Deleterious | 0.779 | Possibly Damaging | 0.159 | Benign | -1.21 | Pathogenic | 0.21 | Tolerated | 0.1399 | 0.0837 | 0 | -1 | -6.4 | 24.99 | |||||||||||||||||||||||||||||
| c.976C>G | H326D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 H326D missense variant is not reported in ClinVar (ClinVar ID None) and has no entries in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only SIFT, which scores the variant as benign. All other evaluated algorithms—REVEL, Rosetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized—predict a pathogenic outcome. Tools with inconclusive results are FoldX, Foldetta, and premPS, which are listed as uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also yields a pathogenic verdict. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, reports an uncertain result. Overall, the preponderance of evidence indicates the variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.342579 | Structured | 0.418150 | Uncertain | 0.944 | 0.455 | 0.000 | -13.000 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 0.87 | Ambiguous | 0.6 | 2.00 | Destabilizing | 1.44 | Ambiguous | 0.96 | Ambiguous | 0.561 | Likely Pathogenic | -7.67 | Deleterious | 0.997 | Probably Damaging | 0.994 | Probably Damaging | 2.04 | Pathogenic | 0.10 | Tolerated | 0.2612 | 0.1611 | 1 | -1 | -0.3 | -22.05 | |||||||||||||||||||||||||||||
| c.1193C>A | P398Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P398Q has no ClinVar record (ClinVar ID None) and is not reported in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only FATHMM, while the majority of tools (REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, AlphaMissense‑Default) predict a pathogenic impact. Two tools report uncertainty: ESM1b and AlphaMissense‑Optimized. High‑accuracy assessments further support pathogenicity: the SGM Consensus (derived from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) yields a pathogenic verdict, and Foldetta, which integrates FoldX‑MD and Rosetta outputs, also predicts pathogenicity. No prediction or stability result is missing or inconclusive. Consequently, the variant is most likely pathogenic based on the collective evidence, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.436924 | Structured | 0.401041 | Uncertain | 0.891 | 0.525 | 0.250 | -7.402 | In-Between | 0.792 | Likely Pathogenic | Ambiguous | 2.13 | Destabilizing | 0.1 | 2.01 | Destabilizing | 2.07 | Destabilizing | 1.01 | Destabilizing | 0.719 | Likely Pathogenic | -5.57 | Deleterious | 0.996 | Probably Damaging | 0.724 | Possibly Damaging | 5.48 | Benign | 0.00 | Affected | 0.1522 | 0.5176 | 0 | -1 | -1.9 | 31.01 | ||||||||||||||||||||||||||||||
| c.1687A>G | R563G 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant R563G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a predominance of pathogenic calls: Rosetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and AlphaMissense‑Default all predict deleterious effects, while the consensus SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates likely pathogenicity. Benign predictions come from REVEL, SIFT, and FATHMM. High‑accuracy assessments give an uncertain result from AlphaMissense‑Optimized, a pathogenic outcome from the SGM‑Consensus, and an uncertain outcome from Foldetta (which integrates FoldX‑MD and Rosetta stability scores). Overall, the balance of evidence favors a pathogenic interpretation, and this conclusion does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.039760 | Structured | 0.031987 | Uncertain | 0.876 | 0.209 | 0.000 | -9.549 | Likely Pathogenic | 0.848 | Likely Pathogenic | Ambiguous | 1.41 | Ambiguous | 0.0 | 2.01 | Destabilizing | 1.71 | Ambiguous | 0.89 | Ambiguous | 0.253 | Likely Benign | -5.68 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.42 | Benign | 0.13 | Tolerated | 0.3082 | 0.1969 | -3 | -2 | 4.1 | -99.14 | |||||||||||||||||||||||||||||
| c.1936C>G | L646V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L646V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split assessment: benign predictions come from REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, AlphaMissense‑Optimized, and FATHMM, whereas pathogenic predictions arise from FoldX, Rosetta, premPS, SIFT, and ESM1b. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, resolves to “Likely Benign” (3 benign vs 1 pathogenic votes). High‑accuracy assessments further indicate AlphaMissense‑Optimized as benign, SGM‑Consensus as likely benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as pathogenic. Overall, the majority of tools, including the high‑accuracy methods, lean toward a benign effect, and this conclusion does not contradict any ClinVar annotation (none exists). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.048328 | Structured | 0.303751 | Uncertain | 0.941 | 0.344 | 0.000 | -8.995 | Likely Pathogenic | 0.332 | Likely Benign | Likely Benign | 3.82 | Destabilizing | 0.2 | 2.01 | Destabilizing | 2.92 | Destabilizing | 1.41 | Destabilizing | 0.125 | Likely Benign | -1.40 | Neutral | 0.040 | Benign | 0.021 | Benign | 3.21 | Benign | 0.03 | Affected | 0.2200 | 0.3767 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.1987T>C | F663L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F663L is not reported in ClinVar and has no entries in gnomAD. Prediction tools that agree on a benign effect are limited to FATHMM, which scores the variant as benign. All other evaluated algorithms—SGM‑Consensus, REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—consistently predict a pathogenic impact. High‑accuracy assessments further support this: AlphaMissense‑Optimized classifies the variant as pathogenic, the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also predicts pathogenicity, while Foldetta’s stability analysis is inconclusive and therefore unavailable. FoldX and Foldetta are treated as unavailable due to uncertain outputs. Based on the overwhelming agreement among pathogenic predictors and the lack of benign evidence, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.056825 | Structured | 0.093963 | Uncertain | 0.944 | 0.355 | 0.000 | -11.996 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.91 | Ambiguous | 0.1 | 2.01 | Destabilizing | 1.46 | Ambiguous | 1.51 | Destabilizing | 0.540 | Likely Pathogenic | -5.99 | Deleterious | 0.999 | Probably Damaging | 0.976 | Probably Damaging | 3.07 | Benign | 0.01 | Affected | 0.1973 | 0.2936 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1989T>A | F663L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F663L is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL and FATHMM, whereas the majority of other in silico predictors (PolyPhen‑2 HumDiv/HumVar, SIFT, ESM1b, PROVEAN, AlphaMissense‑Default, AlphaMissense‑Optimized, SGM Consensus, premPS, and Rosetta) indicate a pathogenic impact. FoldX‑MD and Rosetta‑based stability calculations are inconclusive, yielding an uncertain Foldetta result. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, SGM Consensus as likely pathogenic, and Foldetta as uncertain. Taken together, the preponderance of evidence from consensus and high‑accuracy tools points to a pathogenic effect for F663L. This conclusion is not contradicted by ClinVar status, as no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.056825 | Structured | 0.093963 | Uncertain | 0.944 | 0.355 | 0.000 | -11.996 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.91 | Ambiguous | 0.1 | 2.01 | Destabilizing | 1.46 | Ambiguous | 1.51 | Destabilizing | 0.423 | Likely Benign | -5.99 | Deleterious | 0.999 | Probably Damaging | 0.976 | Probably Damaging | 3.07 | Benign | 0.01 | Affected | 0.1973 | 0.2936 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1989T>G | F663L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F663L is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL and FATHMM, whereas the majority of other in silico predictors (PolyPhen‑2 HumDiv/HumVar, SIFT, ESM1b, PROVEAN, AlphaMissense‑Default, AlphaMissense‑Optimized, premPS, and the SGM Consensus) indicate a pathogenic impact. FoldX‑MD and Rosetta give conflicting stability results, with FoldX uncertain and Rosetta pathogenic; Foldetta, which integrates both, is also uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus as likely pathogenic, and Foldetta as inconclusive. Overall, the preponderance of evidence points to a pathogenic effect for F663L. This conclusion is not contradicted by ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.056825 | Structured | 0.093963 | Uncertain | 0.944 | 0.355 | 0.000 | -11.996 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.91 | Ambiguous | 0.1 | 2.01 | Destabilizing | 1.46 | Ambiguous | 1.51 | Destabilizing | 0.423 | Likely Benign | -5.99 | Deleterious | 0.999 | Probably Damaging | 0.976 | Probably Damaging | 3.07 | Benign | 0.01 | Affected | 0.1973 | 0.2936 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.710C>G | A237G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A237G is not reported in ClinVar and is absent from gnomAD. Prediction tools that indicate a benign effect include ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, polyPhen‑2 HumVar, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). Tools that predict a pathogenic effect are REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 HumDiv, and SIFT. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus as likely benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Given the balance of evidence, the majority of high‑confidence predictions lean toward a benign impact, and there is no ClinVar annotation to contradict this assessment. Thus, the variant is most likely benign. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | PH | 0.200174 | Structured | 0.334699 | Uncertain | 0.719 | 0.352 | 0.000 | -6.647 | Likely Benign | 0.316 | Likely Benign | Likely Benign | 1.00 | Ambiguous | 0.0 | 2.01 | Destabilizing | 1.51 | Ambiguous | 1.14 | Destabilizing | 0.538 | Likely Pathogenic | -2.91 | Deleterious | 0.900 | Possibly Damaging | 0.430 | Benign | 5.81 | Benign | 0.05 | Affected | 0.1809 | 0.2910 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.950T>G | L317R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L317R is not reported in ClinVar (ClinVar ID = None) and is absent from gnomAD (gnomAD ID = None). Prediction tools that assess pathogenicity unanimously classify the variant as deleterious: SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool predicts a benign effect. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized predicts Pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts Pathogenic. Based on the collective evidence, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status because the variant is not yet catalogued there. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.106997 | Structured | 0.394031 | Uncertain | 0.874 | 0.240 | 0.125 | -14.185 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 3.33 | Destabilizing | 0.2 | 2.01 | Destabilizing | 2.67 | Destabilizing | 1.63 | Destabilizing | 0.569 | Likely Pathogenic | -5.52 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.75 | Pathogenic | 0.00 | Affected | 0.1501 | 0.0893 | -3 | -2 | -8.3 | 43.03 | |||||||||||||||||||||||||||||
| c.1182A>C | K394N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 K394N missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, FATHMM, and polyPhen‑2 HumVar. Those that agree on a pathogenic effect are Rosetta, PROVEAN, polyPhen‑2 HumDiv, SIFT, and AlphaMissense‑Default. Predictions that are uncertain or inconclusive are Foldetta, premPS, ESM1b, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, Foldetta as uncertain, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic. Overall, the majority of available predictions lean toward pathogenicity, and this conclusion does not contradict the ClinVar status, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.505461 | Disordered | 0.399336 | Uncertain | 0.387 | 0.634 | 0.625 | -7.408 | In-Between | 0.861 | Likely Pathogenic | Ambiguous | 0.08 | Likely Benign | 0.1 | 2.02 | Destabilizing | 1.05 | Ambiguous | 0.66 | Ambiguous | 0.299 | Likely Benign | -3.17 | Deleterious | 0.535 | Possibly Damaging | 0.188 | Benign | 4.60 | Benign | 0.01 | Affected | 0.4353 | 0.2654 | 1 | 0 | 0.4 | -14.07 | ||||||||||||||||||||||||||||||
| c.1182A>T | K394N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 K394N missense variant has no ClinVar entry and is not reported in gnomAD. Functional prediction tools show a mixed signal: benign calls come from REVEL, FoldX, FATHMM, and polyPhen‑2 HumVar, while pathogenic calls come from Rosetta, PROVEAN, polyPhen‑2 HumDiv, SIFT, and AlphaMissense‑Default. Four tools (Foldetta, premPS, ESM1b, AlphaMissense‑Optimized) return uncertain results. High‑accuracy assessments further clarify the picture: AlphaMissense‑Optimized remains uncertain; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) resolves as pathogenic; and Foldetta is uncertain. Taken together, the majority of evidence—including the high‑accuracy consensus—points to a pathogenic effect. This conclusion is not contradicted by ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.505461 | Disordered | 0.399336 | Uncertain | 0.387 | 0.634 | 0.625 | -7.408 | In-Between | 0.861 | Likely Pathogenic | Ambiguous | 0.08 | Likely Benign | 0.1 | 2.02 | Destabilizing | 1.05 | Ambiguous | 0.66 | Ambiguous | 0.299 | Likely Benign | -3.17 | Deleterious | 0.535 | Possibly Damaging | 0.188 | Benign | 4.60 | Benign | 0.01 | Affected | 0.4353 | 0.2654 | 1 | 0 | 0.4 | -14.07 | ||||||||||||||||||||||||||||||
| c.1307A>G | E436G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E436G missense variant is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only FATHMM, while the majority of algorithms (SGM‑Consensus, REVEL, Rosetta, PROVEAN, polyPhen‑2 HumDiv/HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic impact; FoldX, premPS, and Foldetta are inconclusive. High‑accuracy assessments further support a deleterious outcome: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta’s stability prediction is uncertain. Based on the preponderance of pathogenic predictions and the lack of benign consensus, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.239899 | Structured | 0.321046 | Uncertain | 0.934 | 0.289 | 0.000 | -12.355 | Likely Pathogenic | 0.966 | Likely Pathogenic | Likely Pathogenic | 1.50 | Ambiguous | 0.1 | 2.02 | Destabilizing | 1.76 | Ambiguous | 0.58 | Ambiguous | 0.802 | Likely Pathogenic | -6.63 | Deleterious | 1.000 | Probably Damaging | 0.993 | Probably Damaging | 4.61 | Benign | 0.03 | Affected | 0.2910 | 0.4841 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.1378A>G | K460E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K460E is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, SIFT, and FATHMM. In contrast, a majority of tools predict a pathogenic impact: AlphaMissense‑Default, AlphaMissense‑Optimized, ESM1b, PROVEAN, polyPhen‑2 (HumDiv and HumVar), Rosetta, and premPS all indicate pathogenicity, while the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also reports “Likely Pathogenic.” High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, SGM‑Consensus as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. No prediction or folding‑stability result is missing or inconclusive. Overall, the preponderance of evidence points to a pathogenic effect for K460E, and this conclusion does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.155435 | Structured | 0.289547 | Uncertain | 0.938 | 0.150 | 0.125 | -13.304 | Likely Pathogenic | 0.982 | Likely Pathogenic | Likely Pathogenic | 1.68 | Ambiguous | 0.0 | 2.02 | Destabilizing | 1.85 | Ambiguous | 1.05 | Destabilizing | 0.398 | Likely Benign | -3.40 | Deleterious | 0.999 | Probably Damaging | 0.991 | Probably Damaging | 3.34 | Benign | 0.09 | Tolerated | 0.3966 | 0.1022 | 0 | 1 | 0.4 | 0.94 | |||||||||||||||||||||||||||||
| c.1789T>G | F597V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F597V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools uniformly indicate a deleterious effect. Benign predictions: none. Pathogenic predictions: SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments corroborate this: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. All available evidence points to a pathogenic effect. Therefore, the variant is most likely pathogenic, and this conclusion is consistent with the absence of a ClinVar entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.010926 | Structured | 0.142961 | Uncertain | 0.944 | 0.151 | 0.000 | -13.883 | Likely Pathogenic | 0.981 | Likely Pathogenic | Likely Pathogenic | 3.75 | Destabilizing | 0.7 | 2.02 | Destabilizing | 2.89 | Destabilizing | 1.60 | Destabilizing | 0.939 | Likely Pathogenic | -6.97 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | -2.16 | Pathogenic | 0.01 | Affected | 0.2237 | 0.1583 | -1 | -1 | 1.4 | -48.04 | |||||||||||||||||||||||||||||
| c.1967A>G | E656G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E656G missense variant has no ClinVar entry and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: pathogenic calls come from REVEL, Rosetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Pathogenic). Only premPS and FATHMM predict a benign outcome, while FoldX and Foldetta are inconclusive. High‑accuracy assessments reinforce the pathogenic signal: AlphaMissense‑Optimized is pathogenic; the SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also indicates pathogenicity; Foldetta remains uncertain. Overall, the preponderance of evidence points to a pathogenic effect for E656G, and this conclusion is not contradicted by ClinVar status, which currently lacks an entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.032017 | Structured | 0.242242 | Uncertain | 0.963 | 0.264 | 0.000 | -14.112 | Likely Pathogenic | 0.978 | Likely Pathogenic | Likely Pathogenic | 0.96 | Ambiguous | 0.1 | 2.02 | Destabilizing | 1.49 | Ambiguous | 0.22 | Likely Benign | 0.534 | Likely Pathogenic | -6.28 | Deleterious | 1.000 | Probably Damaging | 0.941 | Probably Damaging | 3.44 | Benign | 0.05 | Affected | 0.3216 | 0.5208 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.2036T>G | F679C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F679C has no ClinVar entry and is not reported in gnomAD. Prediction tools largely agree on a deleterious effect: FATHMM is the sole benign predictor, whereas SGM‑Consensus, REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the variant as pathogenic. FoldX and Foldetta are uncertain and are treated as unavailable evidence. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates likely pathogenic, and Foldetta remains inconclusive. Overall, the preponderance of evidence points to a pathogenic impact for F679C. This prediction is not contradicted by ClinVar status, which currently lacks any classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.222385 | Structured | 0.129316 | Uncertain | 0.700 | 0.320 | 0.000 | -10.269 | Likely Pathogenic | 0.958 | Likely Pathogenic | Likely Pathogenic | 1.65 | Ambiguous | 0.3 | 2.02 | Destabilizing | 1.84 | Ambiguous | 1.17 | Destabilizing | 0.532 | Likely Pathogenic | -7.86 | Deleterious | 0.999 | Probably Damaging | 0.993 | Probably Damaging | 3.40 | Benign | 0.00 | Affected | 0.2344 | 0.0949 | -4 | -2 | -0.3 | -44.04 | |||||||||||||||||||||||||||||
| c.746C>G | A249G 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant A249G is not reported in ClinVar and is absent from gnomAD. Prediction tools largely agree on a deleterious effect: pathogenic calls come from SGM‑Consensus (Likely Pathogenic), REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. Only FATHMM predicts a benign outcome. Uncertain results are reported by FoldX, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show SGM‑Consensus as Likely Pathogenic, while AlphaMissense‑Optimized and Foldetta remain inconclusive. Overall, the preponderance of evidence supports a pathogenic classification for A249G, and this conclusion does not conflict with ClinVar, which contains no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.505461 | Disordered | 0.255452 | Uncertain | 0.810 | 0.336 | 0.125 | -9.678 | Likely Pathogenic | 0.947 | Likely Pathogenic | Ambiguous | 1.04 | Ambiguous | 0.2 | 2.02 | Destabilizing | 1.53 | Ambiguous | 1.19 | Destabilizing | 0.607 | Likely Pathogenic | -2.97 | Deleterious | 0.990 | Probably Damaging | 0.760 | Possibly Damaging | 5.59 | Benign | 0.04 | Affected | 0.1667 | 0.2675 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.1108G>T | G370C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G370C has no ClinVar entry and is not reported in gnomAD. Functional prediction tools fall into two groups: benign predictions come from premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, AlphaMissense‑Default, and AlphaMissense‑Optimized; pathogenic predictions come from REVEL, FoldX, Rosetta, Foldetta, and FATHMM. ESM1b is uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as pathogenic. No prediction or stability result is missing. Overall, the majority of tools predict a benign effect, and the high‑accuracy consensus also leans benign, while only one high‑accuracy method (Foldetta) suggests pathogenicity. Thus, the variant is most likely benign based on the available predictions, and this assessment does not contradict any ClinVar status, as none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.461924 | Structured | 0.434325 | Uncertain | 0.359 | 0.720 | 0.500 | -7.071 | In-Between | 0.119 | Likely Benign | Likely Benign | 3.01 | Destabilizing | 2.1 | 2.03 | Destabilizing | 2.52 | Destabilizing | 0.29 | Likely Benign | 0.511 | Likely Pathogenic | -1.00 | Neutral | 0.353 | Benign | 0.010 | Benign | 1.32 | Pathogenic | 0.06 | Tolerated | 0.1245 | 0.4412 | -3 | -3 | 2.9 | 46.09 | ||||||||||||||||||||||||||||||
| c.1169G>C | G390A 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant G390A is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, premPS, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are SIFT, FATHMM, and Rosetta. The high‑accuracy assessment shows AlphaMissense‑Optimized as benign, the SGM Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as likely benign, and Foldetta (combining FoldX‑MD and Rosetta stability outputs) as pathogenic. FoldX alone is inconclusive. Overall, the majority of evidence points to a benign impact, and this conclusion does not contradict any ClinVar annotation because no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.626927 | Disordered | 0.413274 | Uncertain | 0.304 | 0.763 | 0.875 | -6.768 | Likely Benign | 0.100 | Likely Benign | Likely Benign | 1.96 | Ambiguous | 0.4 | 2.03 | Destabilizing | 2.00 | Destabilizing | 0.02 | Likely Benign | 0.402 | Likely Benign | -0.55 | Neutral | 0.143 | Benign | 0.028 | Benign | 1.33 | Pathogenic | 0.05 | Affected | 0.4130 | 0.5053 | 1 | 0 | 2.2 | 14.03 | |||||||||||||||||||||||||||||
| c.1627C>G | L543V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L543V is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only REVEL, which scores the variant as benign. All other evaluated algorithms—FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN)—classify the variant as pathogenic. AlphaMissense‑Optimized is inconclusive (uncertain). High‑accuracy assessments further support pathogenicity: the SGM‑Consensus predicts “Likely Pathogenic,” and Foldetta, which integrates FoldX‑MD and Rosetta outputs, predicts a destabilizing, pathogenic effect. AlphaMissense‑Optimized remains uncertain. Based on the overwhelming majority of predictions, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.048328 | Structured | 0.020918 | Uncertain | 0.963 | 0.314 | 0.000 | -11.561 | Likely Pathogenic | 0.908 | Likely Pathogenic | Ambiguous | 3.09 | Destabilizing | 0.3 | 2.03 | Destabilizing | 2.56 | Destabilizing | 1.28 | Destabilizing | 0.398 | Likely Benign | -2.99 | Deleterious | 0.998 | Probably Damaging | 0.992 | Probably Damaging | 1.99 | Pathogenic | 0.01 | Affected | 0.1334 | 0.2028 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.2077C>G | H693D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant H693D is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that assess pathogenicity largely agree: SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict a pathogenic effect. Only FATHMM predicts a benign outcome. High‑accuracy methods reinforce the pathogenic signal: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta) is pathogenic. No predictions are missing or inconclusive. Based on the consensus of these tools, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.073402 | Structured | 0.323991 | Uncertain | 0.964 | 0.260 | 0.000 | -15.500 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 2.60 | Destabilizing | 0.1 | 2.03 | Destabilizing | 2.32 | Destabilizing | 1.62 | Destabilizing | 0.578 | Likely Pathogenic | -8.97 | Deleterious | 1.000 | Probably Damaging | 0.991 | Probably Damaging | 3.09 | Benign | 0.01 | Affected | 0.2166 | 0.1108 | 1 | -1 | -0.3 | -22.05 | |||||||||||||||||||||||||||||
| c.2084T>A | L695Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L695Q is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include SIFT, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic impact comprise REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, AlphaMissense‑Default, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). FoldX and Foldetta provide uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while the SGM‑Consensus, derived from a majority of high‑confidence predictors, indicates pathogenicity; Foldetta remains inconclusive. Overall, the majority of reliable tools predict a pathogenic effect, and this is consistent with the lack of ClinVar annotation (no contradiction). Thus, the variant is most likely pathogenic based on current computational predictions. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.118441 | Structured | 0.373419 | Uncertain | 0.942 | 0.258 | 0.000 | -12.192 | Likely Pathogenic | 0.706 | Likely Pathogenic | Likely Benign | 1.88 | Ambiguous | 0.1 | 2.03 | Destabilizing | 1.96 | Ambiguous | 1.71 | Destabilizing | 0.554 | Likely Pathogenic | -5.92 | Deleterious | 1.000 | Probably Damaging | 0.993 | Probably Damaging | 3.17 | Benign | 0.08 | Tolerated | 0.0869 | 0.0688 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.634T>A | S212T 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant S212T is not reported in ClinVar and has no entries in gnomAD. Prediction tools that agree on a benign effect include FoldX and FATHMM, whereas the majority of tools predict a pathogenic impact: REVEL, SGM‑Consensus, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. With most evidence pointing to deleterious effects and no conflicting ClinVar annotation, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.158265 | Structured | 0.381517 | Uncertain | 0.846 | 0.278 | 0.125 | -11.473 | Likely Pathogenic | 0.939 | Likely Pathogenic | Ambiguous | 0.32 | Likely Benign | 0.2 | 2.03 | Destabilizing | 1.18 | Ambiguous | 0.58 | Ambiguous | 0.767 | Likely Pathogenic | -2.58 | Deleterious | 0.956 | Probably Damaging | 0.931 | Probably Damaging | 5.79 | Benign | 0.01 | Affected | 0.1077 | 0.6284 | 1 | 1 | 0.1 | 14.03 | |||||||||||||||||||||||||||||
| c.1636T>C | C546R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C546R is not reported in ClinVar (ClinVar ID = None) and is absent from gnomAD (gnomAD ID = None). Prediction tools that agree on a benign effect include only SIFT, whereas the majority of tools predict a pathogenic impact: SGM‑Consensus (Likely Pathogenic), REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. FoldX and Foldetta return uncertain results and are therefore not considered evidence for either side. High‑accuracy assessments show AlphaMissense‑Optimized as Pathogenic, SGM‑Consensus as Likely Pathogenic, and Foldetta as Uncertain. Based on the preponderance of pathogenic predictions, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status because no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.027463 | Structured | 0.009041 | Uncertain | 0.960 | 0.288 | 0.000 | -15.237 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | -0.70 | Ambiguous | 0.3 | 2.04 | Destabilizing | 0.67 | Ambiguous | 1.70 | Destabilizing | 0.894 | Likely Pathogenic | -9.73 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | -1.26 | Pathogenic | 0.06 | Tolerated | 0.1808 | 0.1685 | -4 | -3 | -7.0 | 53.05 | |||||||||||||||||||||||||||||
| c.1705T>C | F569L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F569L is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect are none; all available predictors that provide a verdict classify the variant as pathogenic, with the exception of FoldX and Foldetta, whose results are uncertain and therefore treated as unavailable. The high‑accuracy predictors give the following: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates likely pathogenic; Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, is uncertain and thus not considered evidence. Based on the overwhelming consensus of pathogenic predictions and the lack of contrary evidence, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.024393 | Structured | 0.054289 | Uncertain | 0.941 | 0.242 | 0.000 | -9.784 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.86 | Ambiguous | 0.1 | 2.04 | Destabilizing | 1.45 | Ambiguous | 1.28 | Destabilizing | 0.804 | Likely Pathogenic | -5.98 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | -1.13 | Pathogenic | 0.05 | Affected | 0.1977 | 0.2476 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1707T>A | F569L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F569L is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect are none; all available predictors that provide a verdict classify the variant as pathogenic, with the exception of FoldX and Foldetta, whose results are uncertain and therefore treated as unavailable. The high‑accuracy predictors give the following: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates likely pathogenic; Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, is uncertain and thus not considered evidence. Based on the overwhelming consensus of pathogenic predictions and the lack of contrary evidence, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.024393 | Structured | 0.054289 | Uncertain | 0.941 | 0.242 | 0.000 | -9.784 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.86 | Ambiguous | 0.1 | 2.04 | Destabilizing | 1.45 | Ambiguous | 1.28 | Destabilizing | 0.675 | Likely Pathogenic | -5.98 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | -1.13 | Pathogenic | 0.05 | Affected | 0.1977 | 0.2476 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1707T>G | F569L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F569L is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect are none; all available predictors that provide a verdict classify the variant as pathogenic, with the exception of FoldX and Foldetta, whose outputs are uncertain and therefore treated as unavailable. The high‑accuracy predictors give the following results: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates likely pathogenic; and Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta, is uncertain. Based on the collective evidence, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which currently contains no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.024393 | Structured | 0.054289 | Uncertain | 0.941 | 0.242 | 0.000 | -9.784 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.86 | Ambiguous | 0.1 | 2.04 | Destabilizing | 1.45 | Ambiguous | 1.28 | Destabilizing | 0.677 | Likely Pathogenic | -5.98 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | -1.13 | Pathogenic | 0.05 | Affected | 0.1977 | 0.2476 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.2018T>G | L673R 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L673R is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign calls come from REVEL, polyPhen‑2 HumVar, FATHMM, and AlphaMissense‑Optimized; pathogenic calls come from Rosetta, premPS, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, and the SGM Consensus (majority vote). FoldX and Foldetta are inconclusive. High‑accuracy assessments give a benign result from AlphaMissense‑Optimized, a likely pathogenic verdict from the SGM Consensus, and an uncertain outcome from Foldetta. Overall, the majority of evidence points toward pathogenicity, and this conclusion does not conflict with the ClinVar status, which currently has no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.060549 | Structured | 0.104692 | Uncertain | 0.545 | 0.369 | 0.000 | -14.474 | Likely Pathogenic | 0.714 | Likely Pathogenic | Likely Benign | 1.10 | Ambiguous | 0.2 | 2.04 | Destabilizing | 1.57 | Ambiguous | 1.23 | Destabilizing | 0.178 | Likely Benign | -3.81 | Deleterious | 0.991 | Probably Damaging | 0.433 | Benign | 3.36 | Benign | 0.03 | Affected | 0.1348 | 0.1015 | -3 | -2 | -8.3 | 43.03 | |||||||||||||||||||||||||||||
| c.2150T>A | L717H 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L717H occurs in the GAP domain. ClinVar has no entry for this variant, and it is not present in gnomAD. Prediction tools that agree on a benign effect are REVEL and FATHMM; all other evaluated algorithms (FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic impact, and the SGM‑Consensus score is “Likely Pathogenic.” High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. No predictions or stability results are missing or inconclusive. Based on the overwhelming majority of computational evidence, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.239899 | Structured | 0.429342 | Uncertain | 0.969 | 0.397 | 0.000 | -11.107 | Likely Pathogenic | 0.977 | Likely Pathogenic | Likely Pathogenic | 2.27 | Destabilizing | 0.1 | 2.04 | Destabilizing | 2.16 | Destabilizing | 1.05 | Destabilizing | 0.317 | Likely Benign | -5.23 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.29 | Benign | 0.00 | Affected | 0.0918 | 0.0558 | -2 | -3 | -7.0 | 23.98 | |||||||||||||||||||||||||||||
| c.973C>A | L325M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L325M is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, PROVEAN, ESM1b, and AlphaMissense‑Optimized. Those that agree on a pathogenic effect are Rosetta, polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, and premPS. Two tools give uncertain results: Foldetta and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as benign, and Foldetta as uncertain. No prediction or folding stability result is missing or inconclusive. Overall, the majority of tools (six pathogenic vs five benign) lean toward pathogenicity, but the high‑accuracy methods and several benign predictions introduce uncertainty. Thus, the variant is most likely pathogenic based on the collective evidence, and this assessment is not contradicted by any ClinVar entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.352862 | Structured | 0.424577 | Uncertain | 0.955 | 0.436 | 0.000 | -6.725 | Likely Benign | 0.445 | Ambiguous | Likely Benign | 0.22 | Likely Benign | 0.0 | 2.04 | Destabilizing | 1.13 | Ambiguous | 1.05 | Destabilizing | 0.306 | Likely Benign | -0.86 | Neutral | 0.997 | Probably Damaging | 0.939 | Probably Damaging | 1.36 | Pathogenic | 0.01 | Affected | 0.0934 | 0.3942 | 4 | 2 | -1.9 | 18.03 | ||||||||||||||||||||||||||||||
| c.1142G>C | G381A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G381A is reported in gnomAD (variant ID 6-33438047‑G‑C) but has no ClinVar entry. Functional prediction tools fall into two groups: benign predictions come from premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and polyPhen‑2 HumVar; pathogenic predictions come from REVEL, FoldX, Rosetta, polyPhen‑2 HumDiv, FATHMM, and the SGM‑Consensus score. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely benign, while Foldetta (combining FoldX‑MD and Rosetta stability outputs) indicates a pathogenic effect. No prediction or stability result is missing or inconclusive. Overall, the majority of tools suggest a benign effect, and the high‑accuracy consensus leans toward benign, though Foldetta’s pathogenic signal introduces uncertainty. The variant is most likely benign based on the collective predictions, and this assessment does not contradict any ClinVar status because none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.724957 | Disordered | 0.431692 | Uncertain | 0.301 | 0.951 | 0.750 | 6-33438047-G-C | 1 | 6.23e-7 | -6.266 | Likely Benign | 0.103 | Likely Benign | Likely Benign | 3.97 | Destabilizing | 0.7 | 2.05 | Destabilizing | 3.01 | Destabilizing | 0.06 | Likely Benign | 0.507 | Likely Pathogenic | -0.63 | Neutral | 0.718 | Possibly Damaging | 0.332 | Benign | 1.33 | Pathogenic | 0.52 | Tolerated | 4.32 | 9 | 0.3809 | 0.4770 | 0 | 1 | 2.2 | 14.03 | ||||||||||||||||||||||||
| c.1294T>C | C432R 2D  3DClick to see structure in 3D Viewer AIClinVar has no entry for this SynGAP1 missense variant, and it is absent from gnomAD. Prediction tools that agree on a benign effect include only FATHMM. The majority of algorithms predict a pathogenic impact: REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Pathogenic). FoldX and Foldetta report uncertain results. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus is likely pathogenic, while Foldetta remains inconclusive. Overall, the consensus of the available predictions indicates that the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which currently has no classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.111485 | Structured | 0.362533 | Uncertain | 0.960 | 0.285 | 0.000 | -13.858 | Likely Pathogenic | 0.996 | Likely Pathogenic | Likely Pathogenic | 1.03 | Ambiguous | 0.2 | 2.05 | Destabilizing | 1.54 | Ambiguous | 1.73 | Destabilizing | 0.690 | Likely Pathogenic | -11.46 | Deleterious | 0.999 | Probably Damaging | 0.993 | Probably Damaging | 3.45 | Benign | 0.01 | Affected | 0.1359 | 0.1766 | -4 | -3 | -7.0 | 53.05 | |||||||||||||||||||||||||||||
| c.1448T>C | I483T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I483T is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect are REVEL and FATHMM, whereas the remaining tools—FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—consistently predict a pathogenic impact. High‑accuracy assessments further support this: AlphaMissense‑Optimized classifies the variant as pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) reports it as Likely Pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, also predicts a pathogenic effect. Based on the preponderance of pathogenic predictions and the absence of any benign consensus, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.206376 | Structured | 0.415850 | Uncertain | 0.798 | 0.254 | 0.000 | -10.692 | Likely Pathogenic | 0.992 | Likely Pathogenic | Likely Pathogenic | 2.32 | Destabilizing | 0.0 | 2.05 | Destabilizing | 2.19 | Destabilizing | 1.77 | Destabilizing | 0.474 | Likely Benign | -4.24 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.15 | Benign | 0.05 | Affected | 0.0888 | 0.0840 | 0 | -1 | -5.2 | -12.05 | |||||||||||||||||||||||||||||
| c.1661T>A | V554E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V554E is not reported in ClinVar and is absent from gnomAD. Prediction tools largely agree on a deleterious effect: all evaluated algorithms except FATHMM classify the variant as pathogenic (REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default). FATHMM is the sole benign prediction. High‑accuracy assessments reinforce the pathogenic signal: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenic. No predictions are missing or inconclusive. Based on the overwhelming consensus of pathogenic predictions, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.020522 | Structured | 0.007349 | Uncertain | 0.955 | 0.226 | 0.000 | -12.813 | Likely Pathogenic | 0.992 | Likely Pathogenic | Likely Pathogenic | 2.51 | Destabilizing | 0.1 | 2.05 | Destabilizing | 2.28 | Destabilizing | 2.42 | Destabilizing | 0.590 | Likely Pathogenic | -5.96 | Deleterious | 1.000 | Probably Damaging | 0.994 | Probably Damaging | 3.21 | Benign | 0.00 | Affected | 0.0862 | 0.1180 | -2 | -2 | -7.7 | 29.98 | |||||||||||||||||||||||||||||
| c.1765A>T | I589F 2D  AIThe SynGAP1 missense variant I589F is not reported in ClinVar (ClinVar status: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that assess pathogenicity all agree that the variant is deleterious: SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify it as pathogenic. No tool predicts a benign effect. High‑accuracy methods reinforce this consensus: AlphaMissense‑Optimized predicts pathogenicity; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also predicts pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, indicates a pathogenic impact. All available predictions are concordant and supportive. Therefore, the variant is most likely pathogenic based on current computational evidence, and this assessment does not contradict any ClinVar annotation (none exists). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.018415 | Structured | 0.084536 | Uncertain | 0.927 | 0.214 | 0.000 | -15.300 | Likely Pathogenic | 0.993 | Likely Pathogenic | Likely Pathogenic | 7.38 | Destabilizing | 3.4 | 2.05 | Destabilizing | 4.72 | Destabilizing | 1.04 | Destabilizing | 0.905 | Likely Pathogenic | -3.98 | Deleterious | 0.999 | Probably Damaging | 0.993 | Probably Damaging | -1.98 | Pathogenic | 0.00 | Affected | 0.0888 | 0.2574 | 1 | 0 | -1.7 | 34.02 | |||||||||||||||||||||||||||||
| c.1879G>T | A627S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 A627S missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are Rosetta, PROVEAN, and ESM1b. The remaining tools—FoldX, Foldetta, and premPS—return uncertain or inconclusive results and are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive due to a 2‑vs‑2 split, and Foldetta is uncertain. Overall, the majority of available predictions lean toward a benign impact. Thus, the variant is most likely benign, and this conclusion does not contradict the ClinVar status, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.100716 | Structured | 0.037862 | Uncertain | 0.970 | 0.210 | 0.000 | -10.782 | Likely Pathogenic | 0.329 | Likely Benign | Likely Benign | 1.11 | Ambiguous | 0.2 | 2.05 | Destabilizing | 1.58 | Ambiguous | 0.71 | Ambiguous | 0.316 | Likely Benign | -2.94 | Deleterious | 0.411 | Benign | 0.387 | Benign | 2.78 | Benign | 0.11 | Tolerated | 0.2266 | 0.4224 | 1 | 1 | -2.6 | 16.00 | ||||||||||||||||||||||||||||||
| c.1979T>C | M660T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M660T is catalogued in gnomAD (ID 6‑33441238‑T‑C) but has no ClinVar entry, so its clinical status is currently unreported. In silico prediction tools largely agree that the substitution is deleterious: pathogenic predictions come from SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Only FATHMM predicts a benign effect. High‑accuracy assessments reinforce the pathogenic view: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. No prediction or folding result is missing or inconclusive. Based on the consensus of these tools, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.047319 | Structured | 0.134270 | Uncertain | 0.944 | 0.289 | 0.000 | 6-33441238-T-C | 1 | 6.20e-7 | -9.791 | Likely Pathogenic | 0.989 | Likely Pathogenic | Likely Pathogenic | 3.62 | Destabilizing | 0.1 | 2.05 | Destabilizing | 2.84 | Destabilizing | 1.91 | Destabilizing | 0.561 | Likely Pathogenic | -5.99 | Deleterious | 0.967 | Probably Damaging | 0.633 | Possibly Damaging | 3.36 | Benign | 0.02 | Affected | 3.38 | 28 | 0.1811 | 0.1630 | -1 | -1 | -2.6 | -30.09 | ||||||||||||||||||||||||
| c.841T>C | Y281H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant Y281H is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: SIFT classifies it as benign, whereas the remaining 13 tools (SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default) predict pathogenicity. High‑accuracy methods provide a clearer picture: AlphaMissense‑Optimized is uncertain, SGM‑Consensus indicates likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic folding instability. With the overwhelming consensus from most predictors and the high‑accuracy tools supporting a damaging effect, the variant is most likely pathogenic. This conclusion is not contradicted by ClinVar, which contains no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.098513 | Structured | 0.337647 | Uncertain | 0.927 | 0.254 | 0.000 | -8.522 | Likely Pathogenic | 0.884 | Likely Pathogenic | Ambiguous | 2.65 | Destabilizing | 0.2 | 2.05 | Destabilizing | 2.35 | Destabilizing | 1.63 | Destabilizing | 0.717 | Likely Pathogenic | -3.82 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 0.99 | Pathogenic | 0.16 | Tolerated | 0.2651 | 0.1503 | 0 | 2 | -1.9 | -26.03 | |||||||||||||||||||||||||||||
| c.1316T>A | L439Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L439Q is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that agree on a benign effect include only FATHMM, whereas all other evaluated algorithms—SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—classify the variant as pathogenic. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. No prediction or folding stability result is missing or inconclusive. Based on the overwhelming consensus of pathogenic predictions, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.222385 | Structured | 0.281542 | Uncertain | 0.942 | 0.265 | 0.000 | -11.162 | Likely Pathogenic | 0.972 | Likely Pathogenic | Likely Pathogenic | 2.50 | Destabilizing | 0.1 | 2.06 | Destabilizing | 2.28 | Destabilizing | 1.35 | Destabilizing | 0.575 | Likely Pathogenic | -5.15 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.21 | Benign | 0.01 | Affected | 0.1065 | 0.0488 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.1382C>G | A461G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A461G has no ClinVar entry and is not reported in gnomAD. Prediction tools cluster into three groups: benign predictions come from REVEL, FATHMM, and AlphaMissense‑Optimized; pathogenic predictions arise from Rosetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and ESM1b; the remaining tools (FoldX, premPS, AlphaMissense‑Default, Foldetta) give uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points toward a pathogenic effect, with no ClinVar record to contradict this assessment. Thus, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.179055 | Structured | 0.292531 | Uncertain | 0.936 | 0.151 | 0.125 | -11.360 | Likely Pathogenic | 0.556 | Ambiguous | Likely Benign | 1.46 | Ambiguous | 0.1 | 2.06 | Destabilizing | 1.76 | Ambiguous | 0.83 | Ambiguous | 0.276 | Likely Benign | -3.79 | Deleterious | 0.991 | Probably Damaging | 0.628 | Possibly Damaging | 3.32 | Benign | 0.00 | Affected | 0.1988 | 0.3317 | 1 | 0 | -2.2 | -14.03 | ||||||||||||||||||||||||||||||
| c.1484A>G | E495G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E495G missense variant is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33438516‑A‑G). Among the available in‑silico predictors, the following tools uniformly indicate a pathogenic effect: REVEL, Rosetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus (which itself is a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). No tool in the dataset predicts a benign outcome; predictions that are uncertain (FoldX, Foldetta, premPS, AlphaMissense‑Optimized) are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as “Uncertain,” SGM‑Consensus as “Likely Pathogenic,” and Foldetta as “Uncertain.” Overall, the preponderance of pathogenic predictions strongly suggests that the variant is most likely pathogenic, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.164327 | Structured | 0.364496 | Uncertain | 0.933 | 0.161 | 0.000 | Uncertain | 1 | 6-33438516-A-G | 1 | 6.20e-7 | -9.400 | Likely Pathogenic | 0.923 | Likely Pathogenic | Ambiguous | 1.21 | Ambiguous | 0.0 | 2.06 | Destabilizing | 1.64 | Ambiguous | 0.78 | Ambiguous | 0.867 | Likely Pathogenic | -6.70 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | -1.46 | Pathogenic | 0.02 | Affected | 3.37 | 35 | 0.2177 | 0.4784 | -2 | 0 | 3.1 | -72.06 | ||||||||||||||||||||||
| c.1690G>A | E564K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E564K is not reported in ClinVar and is present in the gnomAD database (variant ID 6‑33440742‑G‑A). Functional prediction tools largely agree on a deleterious effect: pathogenic calls are made by REVEL, Rosetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized, while only SIFT predicts a benign outcome. Uncertain results are reported by FoldX, Foldetta, and premPS. High‑accuracy assessments reinforce the pathogenic prediction: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic, and Foldetta remains uncertain. Overall, the majority of evidence supports a pathogenic classification, and this conclusion does not contradict the ClinVar status, which currently contains no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.023534 | Structured | 0.038418 | Uncertain | 0.891 | 0.208 | 0.000 | 6-33440742-G-A | -15.834 | Likely Pathogenic | 0.989 | Likely Pathogenic | Likely Pathogenic | 0.76 | Ambiguous | 0.1 | 2.06 | Destabilizing | 1.41 | Ambiguous | 0.89 | Ambiguous | 0.854 | Likely Pathogenic | -3.95 | Deleterious | 0.997 | Probably Damaging | 0.987 | Probably Damaging | -1.35 | Pathogenic | 0.10 | Tolerated | 3.37 | 35 | 0.1988 | 0.5280 | 1 | 0 | -0.4 | -0.94 | ||||||||||||||||||||||||||
| c.1928A>G | E643G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E643G is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include polyPhen‑2 HumVar, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic impact comprise SGM‑Consensus, REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, and AlphaMissense‑Default; FoldX and Foldetta are uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic. Foldetta’s stability prediction is uncertain. Overall, the majority of reliable tools predict pathogenicity, and this conclusion does not contradict the lack of ClinVar annotation. Thus, the variant is most likely pathogenic based on current computational evidence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.033407 | Structured | 0.215915 | Uncertain | 0.871 | 0.315 | 0.000 | -12.503 | Likely Pathogenic | 0.707 | Likely Pathogenic | Likely Benign | 1.45 | Ambiguous | 0.3 | 2.06 | Destabilizing | 1.76 | Ambiguous | 1.01 | Destabilizing | 0.520 | Likely Pathogenic | -6.81 | Deleterious | 0.983 | Probably Damaging | 0.390 | Benign | 2.94 | Benign | 0.00 | Affected | 0.2821 | 0.5319 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.2051A>T | D684V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant D684V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: pathogenic calls come from REVEL, FoldX, Rosetta, Foldetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized, while only premPS and FATHMM predict a benign outcome. High‑accuracy assessments reinforce the pathogenic interpretation: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta) is pathogenic. No evidence suggests a benign effect, and the lack of ClinVar annotation means there is no conflicting clinical classification. Therefore, the variant is most likely pathogenic, with no contradiction to ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.254060 | Structured | 0.153798 | Uncertain | 0.870 | 0.282 | 0.000 | -16.128 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 3.86 | Destabilizing | 1.1 | 2.06 | Destabilizing | 2.96 | Destabilizing | 0.07 | Likely Benign | 0.601 | Likely Pathogenic | -8.98 | Deleterious | 0.901 | Possibly Damaging | 0.480 | Possibly Damaging | 3.44 | Benign | 0.00 | Affected | 0.0775 | 0.6209 | -2 | -3 | 7.7 | -15.96 | |||||||||||||||||||||||||||||
| c.1220A>C | Q407P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant Q407P is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include only FATHMM, whereas the remaining tools (SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) uniformly predict a pathogenic impact; premPS is uncertain and is not counted. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. Based on the consensus of these predictions, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.109221 | Structured | 0.382522 | Uncertain | 0.916 | 0.271 | 0.000 | -13.578 | Likely Pathogenic | 0.994 | Likely Pathogenic | Likely Pathogenic | 3.04 | Destabilizing | 0.5 | 2.07 | Destabilizing | 2.56 | Destabilizing | 0.88 | Ambiguous | 0.515 | Likely Pathogenic | -5.40 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.88 | Benign | 0.02 | Affected | 0.1786 | 0.4352 | 0 | -1 | 1.9 | -31.01 | |||||||||||||||||||||||||||||
| c.1262C>A | A421E 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant A421E is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign calls come from REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and FATHMM, whereas pathogenic calls are made by ESM1b, PROVEAN, AlphaMissense‑Default, AlphaMissense‑Optimized, Rosetta, premPS, and the SGM‑Consensus score (Likely Pathogenic). Stability‑based methods give mixed results: FoldX is uncertain, Foldetta is uncertain, and Rosetta alone predicts pathogenic. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta remains uncertain. Overall, the majority of evidence, including the high‑accuracy tools, points to a pathogenic impact for A421E, and this conclusion does not conflict with the absence of a ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.067594 | Structured | 0.404927 | Uncertain | 0.965 | 0.257 | 0.000 | -11.993 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.63 | Ambiguous | 0.2 | 2.07 | Destabilizing | 1.35 | Ambiguous | 1.30 | Destabilizing | 0.233 | Likely Benign | -4.24 | Deleterious | 0.368 | Benign | 0.144 | Benign | 3.44 | Benign | 0.14 | Tolerated | 0.1288 | 0.1830 | 0 | -1 | -5.3 | 58.04 | |||||||||||||||||||||||||||||
| c.1550T>A | L517Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L517Q is not reported in ClinVar and is absent from gnomAD, indicating no known population frequency data. Prediction tools that assess pathogenicity unanimously classify the variant as deleterious: SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool predicts a benign effect, so the benign group is empty. High‑accuracy methods reinforce this view: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. All available evidence points to a pathogenic impact. Thus, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status because the variant is not yet catalogued there. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.118441 | Structured | 0.147645 | Uncertain | 0.938 | 0.296 | 0.000 | -11.660 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 2.17 | Destabilizing | 0.1 | 2.07 | Destabilizing | 2.12 | Destabilizing | 1.87 | Destabilizing | 0.946 | Likely Pathogenic | -5.71 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.50 | Pathogenic | 0.00 | Affected | 0.1031 | 0.0888 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.1847A>C | D616A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D616A missense variant is not reported in ClinVar and has no entries in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, premPS, SIFT, FATHMM, and polyPhen‑2 HumVar, while pathogenic predictions arise from SGM‑Consensus (Likely Pathogenic), Rosetta, PROVEAN, polyPhen‑2 HumDiv, ESM1b, and AlphaMissense‑Default. The high‑accuracy AlphaMissense‑Optimized model classifies the variant as benign, whereas the SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, indicates pathogenicity. Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is inconclusive. Overall, the majority of evidence points toward a pathogenic effect, and this assessment does not conflict with the absence of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.129801 | Structured | 0.166689 | Uncertain | 0.867 | 0.252 | 0.000 | -11.386 | Likely Pathogenic | 0.664 | Likely Pathogenic | Likely Benign | 1.76 | Ambiguous | 0.2 | 2.07 | Destabilizing | 1.92 | Ambiguous | 0.41 | Likely Benign | 0.126 | Likely Benign | -6.13 | Deleterious | 0.539 | Possibly Damaging | 0.122 | Benign | 3.32 | Benign | 0.10 | Tolerated | 0.3543 | 0.4207 | 0 | -2 | 5.3 | -44.01 | |||||||||||||||||||||||||||||
| c.1861C>G | R621G 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant R621G is reported in gnomAD (ID 6‑33440913‑C‑G) but has no ClinVar entry. In silico predictors largely agree on a deleterious effect: pathogenic calls come from REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The only benign prediction is from FATHMM; FoldX and Foldetta give uncertain results. High‑accuracy assessments reinforce the pathogenic signal: AlphaMissense‑Optimized predicts pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic, and Foldetta remains uncertain. Overall, the preponderance of evidence indicates the variant is most likely pathogenic, and this conclusion is not contradicted by ClinVar data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.222385 | Structured | 0.084420 | Uncertain | 0.945 | 0.216 | 0.000 | 6-33440913-C-G | 1 | 6.20e-7 | -16.611 | Likely Pathogenic | 0.993 | Likely Pathogenic | Likely Pathogenic | 1.18 | Ambiguous | 0.3 | 2.07 | Destabilizing | 1.63 | Ambiguous | 1.17 | Destabilizing | 0.558 | Likely Pathogenic | -6.97 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 2.84 | Benign | 0.01 | Affected | 3.37 | 35 | 0.3109 | 0.2651 | -2 | -3 | 4.1 | -99.14 | ||||||||||||||||||||||||
| c.1880C>G | A627G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A627G is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. Tools that agree on a pathogenic effect comprise SGM‑Consensus, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. FoldX and Foldetta are inconclusive. High‑accuracy assessments show AlphaMissense‑Optimized predicts benign, while the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts pathogenic. Foldetta remains uncertain. Overall, the majority of evidence points to a pathogenic impact, and this conclusion does not contradict the lack of ClinVar annotation. Thus, the variant is most likely pathogenic based on current predictions. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.100716 | Structured | 0.037862 | Uncertain | 0.970 | 0.210 | 0.000 | -11.716 | Likely Pathogenic | 0.640 | Likely Pathogenic | Likely Benign | 1.38 | Ambiguous | 0.1 | 2.07 | Destabilizing | 1.73 | Ambiguous | 1.25 | Destabilizing | 0.458 | Likely Benign | -3.96 | Deleterious | 0.997 | Probably Damaging | 0.876 | Possibly Damaging | 2.51 | Benign | 0.01 | Affected | 0.1921 | 0.2990 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.1213C>A | R405S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R405S is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: benign predictions come from REVEL and FATHMM, whereas pathogenic predictions are made by FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as Likely Pathogenic. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized predicts Pathogenic; the SGM‑Consensus itself is Likely Pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, also predicts Pathogenic. Taken together, the overwhelming majority of evidence indicates that R405S is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.250310 | Structured | 0.404888 | Uncertain | 0.949 | 0.315 | 0.000 | -11.326 | Likely Pathogenic | 0.977 | Likely Pathogenic | Likely Pathogenic | 2.66 | Destabilizing | 0.5 | 2.08 | Destabilizing | 2.37 | Destabilizing | 1.22 | Destabilizing | 0.405 | Likely Benign | -5.40 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.67 | Benign | 0.05 | Affected | 0.2829 | 0.4646 | 0 | -1 | 3.7 | -69.11 | |||||||||||||||||||||||||||||
| c.1942T>C | F648L 2D  AISynGAP1 missense variant F648L is listed in ClinVar with an uncertain significance (ClinVar ID 3383902.0) and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, and FATHMM, whereas the remaining tools—FoldX, Rosetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, AlphaMissense‑Optimized, and ESM1b—consistently predict pathogenicity. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also indicates likely pathogenic. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized scores pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts a destabilizing, pathogenic change. Taken together, the preponderance of evidence points to a pathogenic impact for F648L, which contradicts the current ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.051831 | Structured | 0.346782 | Uncertain | 0.943 | 0.339 | 0.000 | Uncertain | 1 | -9.296 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 2.71 | Destabilizing | 0.8 | 2.08 | Destabilizing | 2.40 | Destabilizing | 1.04 | Destabilizing | 0.468 | Likely Benign | -5.98 | Deleterious | 0.999 | Probably Damaging | 0.976 | Probably Damaging | 3.45 | Benign | 0.08 | Tolerated | 0.1961 | 0.3126 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||
| c.1944C>A | F648L 2D  AIThe SynGAP1 missense variant F648L is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, and FATHMM, while the remaining 12 tools—including FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—predict a pathogenic effect. High‑accuracy assessments reinforce this trend: AlphaMissense‑Optimized is pathogenic; Foldetta, which integrates FoldX‑MD and Rosetta outputs, is pathogenic; and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is Likely Pathogenic. Because the majority of evidence points to a deleterious impact and there is no ClinVar annotation to contradict this, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.051831 | Structured | 0.346782 | Uncertain | 0.943 | 0.339 | 0.000 | -9.296 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 2.71 | Destabilizing | 0.8 | 2.08 | Destabilizing | 2.40 | Destabilizing | 1.04 | Destabilizing | 0.319 | Likely Benign | -5.98 | Deleterious | 0.999 | Probably Damaging | 0.976 | Probably Damaging | 3.45 | Benign | 0.08 | Tolerated | 0.1961 | 0.3126 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1944C>G | F648L 2D  AIThe SynGAP1 missense variant F648L is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, SIFT, and FATHMM. In contrast, the majority of tools predict a pathogenic impact: FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), AlphaMissense‑Default, AlphaMissense‑Optimized, and ESM1b all indicate pathogenicity, and the SGM Consensus score is “Likely Pathogenic.” High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. No predictions are missing or inconclusive. Based on the preponderance of evidence, the variant is most likely pathogenic, and this conclusion is consistent with the absence of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.051831 | Structured | 0.346782 | Uncertain | 0.943 | 0.339 | 0.000 | -9.296 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 2.71 | Destabilizing | 0.8 | 2.08 | Destabilizing | 2.40 | Destabilizing | 1.04 | Destabilizing | 0.319 | Likely Benign | -5.98 | Deleterious | 0.999 | Probably Damaging | 0.976 | Probably Damaging | 3.45 | Benign | 0.08 | Tolerated | 0.1961 | 0.3126 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.2048T>A | I683N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I683N is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default all predict pathogenicity, whereas only FATHMM predicts a benign outcome. High‑accuracy assessments reinforce the pathogenic signal: AlphaMissense‑Optimized returns a pathogenic score, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is labeled Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta stability outputs) also indicates pathogenicity. No predictions are missing or inconclusive. Consequently, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation (none available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.200174 | Structured | 0.143268 | Uncertain | 0.848 | 0.314 | 0.000 | -12.120 | Likely Pathogenic | 0.974 | Likely Pathogenic | Likely Pathogenic | 2.18 | Destabilizing | 0.1 | 2.08 | Destabilizing | 2.13 | Destabilizing | 1.46 | Destabilizing | 0.546 | Likely Pathogenic | -6.87 | Deleterious | 1.000 | Probably Damaging | 0.992 | Probably Damaging | 3.26 | Benign | 0.01 | Affected | 0.0978 | 0.1133 | -2 | -3 | -8.0 | 0.94 | |||||||||||||||||||||||||||||
| c.2096T>A | V699E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 V699E missense variant is not reported in ClinVar and is absent from gnomAD. Prediction tools that classify it as benign include REVEL and FATHMM, whereas the majority of other in‑silico predictors (FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, and the SGM‑Consensus score) indicate a pathogenic effect. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as Pathogenic. Based on the preponderance of pathogenic predictions and the high‑accuracy tools’ results, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.069024 | Structured | 0.432975 | Uncertain | 0.935 | 0.315 | 0.000 | -12.647 | Likely Pathogenic | 0.813 | Likely Pathogenic | Ambiguous | 2.41 | Destabilizing | 0.1 | 2.08 | Destabilizing | 2.25 | Destabilizing | 1.82 | Destabilizing | 0.468 | Likely Benign | -4.56 | Deleterious | 1.000 | Probably Damaging | 0.986 | Probably Damaging | 3.42 | Benign | 0.02 | Affected | 0.1092 | 0.1326 | -2 | -2 | -7.7 | 29.98 | |||||||||||||||||||||||||||||
| c.850C>G | L284V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 L284V missense variant has no ClinVar entry and is not present in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, and AlphaMissense‑Optimized. Those that predict a pathogenic impact are Rosetta, premPS, polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM. Predictions that are uncertain or inconclusive are FoldX, AlphaMissense‑Default, and Foldetta. High‑accuracy methods give a benign consensus: AlphaMissense‑Optimized is benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also favors benign, while Foldetta remains uncertain. Overall, the balance of evidence leans toward a pathogenic effect, and this assessment does not contradict the ClinVar status, which currently has no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.094817 | Structured | 0.371601 | Uncertain | 0.950 | 0.255 | 0.000 | -6.726 | Likely Benign | 0.347 | Ambiguous | Likely Benign | 1.65 | Ambiguous | 0.2 | 2.08 | Destabilizing | 1.87 | Ambiguous | 1.38 | Destabilizing | 0.248 | Likely Benign | -2.32 | Neutral | 0.999 | Probably Damaging | 0.994 | Probably Damaging | 2.18 | Pathogenic | 0.03 | Affected | 0.1259 | 0.2456 | 2 | 1 | 0.4 | -14.03 | ||||||||||||||||||||||||||||||
| c.893C>G | P298R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P298R has no ClinVar entry and is not reported in gnomAD. Computational predictors fall into two consensus groups: benign predictions come from REVEL, FoldX, PROVEAN, SIFT, and AlphaMissense‑Optimized; pathogenic predictions come from SGM‑Consensus, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, and AlphaMissense‑Default. Uncertain results are reported only by Foldetta and premPS. High‑accuracy tools give a mixed picture: AlphaMissense‑Optimized predicts benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts likely pathogenic, and Foldetta reports an uncertain stability change. Overall, the majority of evidence points toward a pathogenic effect, and this assessment is not contradicted by ClinVar, which currently has no classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.328603 | Structured | 0.268765 | Uncertain | 0.860 | 0.283 | 0.500 | -11.427 | Likely Pathogenic | 0.733 | Likely Pathogenic | Likely Benign | 0.45 | Likely Benign | 0.0 | 2.08 | Destabilizing | 1.27 | Ambiguous | 0.61 | Ambiguous | 0.280 | Likely Benign | -1.83 | Neutral | 0.997 | Probably Damaging | 0.952 | Probably Damaging | 2.03 | Pathogenic | 0.09 | Tolerated | 0.1299 | 0.3872 | 0 | -2 | -2.9 | 59.07 | |||||||||||||||||||||||||||||
| c.2174T>A | L725Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L725Q is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect are limited to REVEL, which scores the variant as benign. The majority of tools predict a pathogenic impact: premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, Rosetta, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Uncertain or inconclusive results come from FoldX, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as likely pathogenic, and Foldetta as uncertain. Taken together, the preponderance of evidence indicates that the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar annotation because none exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | 0.557691 | Disordered | 0.455613 | Uncertain | 0.911 | 0.491 | 0.625 | -13.952 | Likely Pathogenic | 0.888 | Likely Pathogenic | Ambiguous | 1.55 | Ambiguous | 0.1 | 2.09 | Destabilizing | 1.82 | Ambiguous | 1.88 | Destabilizing | 0.319 | Likely Benign | -5.43 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 1.28 | Pathogenic | 0.00 | Affected | 0.1198 | 0.1203 | -2 | -2 | -7.3 | 14.97 | ||||||||||||||||||||||||||||||
| c.1405G>T | A469S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A469S is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include PROVEAN, SIFT, and AlphaMissense‑Optimized. Tools that agree on a pathogenic effect are SGM‑Consensus, REVEL, Rosetta, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, and AlphaMissense‑Default. FoldX, Foldetta, and premPS give uncertain or inconclusive results. High‑accuracy methods give the following: AlphaMissense‑Optimized predicts a benign change; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, predicts pathogenic; Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, is uncertain. Overall, the majority of evidence points to a pathogenic effect, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.278302 | Structured | 0.343926 | Uncertain | 0.910 | 0.276 | 0.000 | -9.051 | Likely Pathogenic | 0.627 | Likely Pathogenic | Likely Benign | 0.86 | Ambiguous | 0.0 | 2.10 | Destabilizing | 1.48 | Ambiguous | 0.81 | Ambiguous | 0.558 | Likely Pathogenic | -1.53 | Neutral | 0.953 | Possibly Damaging | 0.985 | Probably Damaging | -1.30 | Pathogenic | 0.41 | Tolerated | 0.2113 | 0.4505 | 1 | 1 | -2.6 | 16.00 | |||||||||||||||||||||||||||||
| c.1769G>A | S590N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S590N is not reported in ClinVar and has no entries in gnomAD. Prediction tools that agree on a benign effect include REVEL and FATHMM, while the majority of other in silico predictors (PolyPhen‑2 HumDiv/HumVar, SIFT, ESM1b, PROVEAN, premPS, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus) indicate a pathogenic or likely pathogenic impact. FoldX and Foldetta, which assess protein‑folding stability, return uncertain results and are therefore not considered evidence for either outcome. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus as likely pathogenic, and Foldetta as uncertain. Overall, the preponderance of pathogenic predictions strongly suggests that S590N is most likely pathogenic, a conclusion that is consistent with the absence of ClinVar annotation and gnomAD data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.022667 | Structured | 0.088943 | Uncertain | 0.918 | 0.199 | 0.000 | -15.149 | Likely Pathogenic | 0.957 | Likely Pathogenic | Likely Pathogenic | 1.27 | Ambiguous | 0.1 | 2.10 | Destabilizing | 1.69 | Ambiguous | 1.48 | Destabilizing | 0.411 | Likely Benign | -2.96 | Deleterious | 0.921 | Possibly Damaging | 0.598 | Possibly Damaging | 3.14 | Benign | 0.01 | Affected | 0.1254 | 0.4202 | 1 | 1 | -2.7 | 27.03 | |||||||||||||||||||||||||||||
| c.2017C>G | L673V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L673V is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include SGM‑Consensus (Likely Benign), REVEL, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic outcome are FoldX, Rosetta, Foldetta, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as Likely Benign, and Foldetta as pathogenic, yielding one pathogenic versus two benign predictions. Overall, the majority of evidence points to a benign impact, and this conclusion does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.060549 | Structured | 0.104692 | Uncertain | 0.545 | 0.369 | 0.000 | -8.132 | Likely Pathogenic | 0.099 | Likely Benign | Likely Benign | 2.18 | Destabilizing | 0.3 | 2.10 | Destabilizing | 2.14 | Destabilizing | 0.10 | Likely Benign | 0.017 | Likely Benign | -0.58 | Neutral | 0.016 | Benign | 0.003 | Benign | 3.36 | Benign | 0.28 | Tolerated | 0.1448 | 0.3777 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.806T>C | I269T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I269T is not reported in ClinVar (no ClinVar entry) but is present in gnomAD (variant ID 6‑33437711‑T‑C). Among general in‑silico predictors, only SIFT classifies the change as benign, whereas the remaining tools that provide a definitive call (REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default) all predict a pathogenic effect. High‑accuracy assessments give a more nuanced view: AlphaMissense‑Optimized is uncertain; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates a likely pathogenic outcome; and Foldetta, which integrates FoldX‑MD and Rosetta stability predictions, also reports a pathogenic effect. Based on the preponderance of pathogenic predictions and the high‑accuracy consensus, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.216401 | Structured | 0.343787 | Uncertain | 0.937 | 0.244 | 0.125 | 6-33437711-T-C | 2 | 1.24e-6 | -9.376 | Likely Pathogenic | 0.887 | Likely Pathogenic | Ambiguous | 1.97 | Ambiguous | 0.1 | 2.10 | Destabilizing | 2.04 | Destabilizing | 1.38 | Destabilizing | 0.727 | Likely Pathogenic | -3.70 | Deleterious | 0.997 | Probably Damaging | 0.994 | Probably Damaging | 1.72 | Pathogenic | 0.09 | Tolerated | 3.38 | 19 | 0.0833 | 0.0808 | -1 | 0 | -5.2 | -12.05 | ||||||||||||||||||||||||
| c.1507C>G | Q503E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant Q503E is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that agree on a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, FATHMM, and Rosetta. Uncertain or inconclusive results come from FoldX, Foldetta, premPS, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also predicts benign, while Foldetta remains uncertain. Overall, the majority of available predictions favor a benign impact, and this conclusion does not contradict the lack of ClinVar annotation. Thus, the variant is most likely benign based on current computational evidence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.040537 | Structured | 0.322935 | Uncertain | 0.848 | 0.168 | 0.000 | -7.909 | In-Between | 0.146 | Likely Benign | Likely Benign | 0.51 | Ambiguous | 0.1 | 2.11 | Destabilizing | 1.31 | Ambiguous | 0.85 | Ambiguous | 0.410 | Likely Benign | -2.21 | Neutral | 0.931 | Possibly Damaging | 0.500 | Possibly Damaging | -1.43 | Pathogenic | 0.17 | Tolerated | 0.1168 | 0.1508 | 2 | 2 | 0.0 | 0.98 | ||||||||||||||||||||||||||||||
| c.2152C>G | L718V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L718V is not reported in ClinVar and is absent from gnomAD. Prediction tools that indicate a benign effect include REVEL and AlphaMissense‑Optimized, whereas the remaining tools (FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default) all predict a pathogenic outcome. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized reports benign, the SGM Consensus (derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. Based on the preponderance of pathogenic predictions and the high‑accuracy consensus, the variant is most likely pathogenic; this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.298791 | Structured | 0.438417 | Uncertain | 0.966 | 0.385 | 0.000 | -11.585 | Likely Pathogenic | 0.693 | Likely Pathogenic | Likely Benign | 3.15 | Destabilizing | 0.1 | 2.11 | Destabilizing | 2.63 | Destabilizing | 1.14 | Destabilizing | 0.237 | Likely Benign | -2.83 | Deleterious | 0.998 | Probably Damaging | 0.992 | Probably Damaging | 1.36 | Pathogenic | 0.00 | Affected | 0.1485 | 0.2656 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.824C>G | P275R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P275R is not reported in ClinVar and is absent from gnomAD. Prediction tools that assess functional impact uniformly indicate a deleterious effect: REVEL, FoldX, Rosetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default all classify the change as pathogenic. No tool predicts a benign outcome. High‑accuracy assessments further support this view: the SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as Likely Pathogenic; Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, also predicts a pathogenic effect. AlphaMissense‑Optimized remains uncertain, but its result does not counter the overall consensus. Consequently, the variant is most likely pathogenic, and this conclusion is consistent with the absence of a ClinVar entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.059222 | Structured | 0.353469 | Uncertain | 0.811 | 0.208 | 0.250 | -13.557 | Likely Pathogenic | 0.915 | Likely Pathogenic | Ambiguous | 2.10 | Destabilizing | 0.6 | 2.11 | Destabilizing | 2.11 | Destabilizing | 0.82 | Ambiguous | 0.645 | Likely Pathogenic | -6.36 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.75 | Pathogenic | 0.01 | Affected | 0.1637 | 0.3039 | 0 | -2 | -2.9 | 59.07 | |||||||||||||||||||||||||||||
| c.1481T>C | I494T 2D  AIThe SynGAP1 missense variant I494T is not reported in ClinVar (ClinVar status: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include only AlphaMissense‑Optimized. All other evaluated tools—SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default—predict a pathogenic impact. High‑accuracy methods give the following results: AlphaMissense‑Optimized predicts benign; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts likely pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. No predictions are missing or inconclusive. Based on the overwhelming majority of pathogenic predictions, the variant is most likely pathogenic, and this assessment does not contradict the ClinVar status, which is currently unreported. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.155435 | Structured | 0.353330 | Uncertain | 0.941 | 0.157 | 0.000 | -10.033 | Likely Pathogenic | 0.754 | Likely Pathogenic | Likely Benign | 2.45 | Destabilizing | 0.7 | 2.12 | Destabilizing | 2.29 | Destabilizing | 1.73 | Destabilizing | 0.907 | Likely Pathogenic | -4.61 | Deleterious | 1.000 | Probably Damaging | 0.995 | Probably Damaging | -1.38 | Pathogenic | 0.01 | Affected | 0.1003 | 0.0640 | 0 | -1 | -5.2 | -12.05 | |||||||||||||||||||||||||||||
| c.1679T>G | V560G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 V560G missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only SIFT, whereas the majority of tools (REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default) predict a pathogenic impact. Uncertain results come from FoldX, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show that the SGM Consensus (derived from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is pathogenic, AlphaMissense‑Optimized is uncertain, and Foldetta is uncertain. Overall, the preponderance of evidence indicates that V560G is most likely pathogenic, and this conclusion does not contradict any ClinVar annotation because no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.021381 | Structured | 0.013872 | Uncertain | 0.853 | 0.204 | 0.000 | -12.485 | Likely Pathogenic | 0.799 | Likely Pathogenic | Ambiguous | 0.66 | Ambiguous | 0.1 | 2.12 | Destabilizing | 1.39 | Ambiguous | 1.80 | Destabilizing | 0.753 | Likely Pathogenic | -5.87 | Deleterious | 0.981 | Probably Damaging | 1.000 | Probably Damaging | -1.25 | Pathogenic | 0.19 | Tolerated | 0.1738 | 0.2036 | -1 | -3 | -4.6 | -42.08 | ||||||||||||||||||||||||||||||
| c.1709C>G | A570G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 A570G missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include SIFT and AlphaMissense‑Optimized. Those that predict a pathogenic effect are REVEL, Rosetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), and FATHMM. The remaining tools (FoldX, premPS, ESM1b, AlphaMissense‑Default) give uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) and Foldetta are unavailable due to mixed or uncertain outputs. Overall, the majority of evaluated tools (seven pathogenic vs. two benign) indicate that the variant is most likely pathogenic, and this assessment does not contradict ClinVar status because no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.046336 | Structured | 0.054494 | Uncertain | 0.932 | 0.263 | 0.000 | -7.509 | In-Between | 0.562 | Ambiguous | Likely Benign | 1.34 | Ambiguous | 0.1 | 2.12 | Destabilizing | 1.73 | Ambiguous | 0.99 | Ambiguous | 0.607 | Likely Pathogenic | -3.62 | Deleterious | 0.999 | Probably Damaging | 0.995 | Probably Damaging | -1.30 | Pathogenic | 0.09 | Tolerated | 0.1700 | 0.2499 | 1 | 0 | -2.2 | -14.03 | ||||||||||||||||||||||||||||||
| c.1789T>C | F597L 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant F597L is listed in ClinVar with an uncertain significance (ClinVar ID 3658115.0) and is not reported in gnomAD. Prediction tools that classify the variant as benign include only SIFT, whereas the remaining tools—SGM‑Consensus, REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized—predict it to be pathogenic. The high‑accuracy AlphaMissense‑Optimized score is pathogenic, and the SGM‑Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates likely pathogenic. Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is inconclusive. Overall, the preponderance of evidence points to a pathogenic effect for F597L, which is consistent with its ClinVar uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.010926 | Structured | 0.142961 | Uncertain | 0.944 | 0.151 | 0.000 | Uncertain | 1 | -10.173 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 0.74 | Ambiguous | 0.1 | 2.12 | Destabilizing | 1.43 | Ambiguous | 1.20 | Destabilizing | 0.929 | Likely Pathogenic | -5.97 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | -2.06 | Pathogenic | 0.13 | Tolerated | 0.2232 | 0.2596 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||
| c.1791C>A | F597L 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant F597L is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: the benign group contains only SIFT, while the pathogenic group includes SGM‑Consensus (Likely Pathogenic), REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. FoldX and Foldetta are inconclusive, providing no definitive evidence. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts a damaging effect, SGM‑Consensus concurs with a likely pathogenic classification, and Foldetta remains uncertain. Taken together, the overwhelming majority of predictions indicate a pathogenic impact, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.010926 | Structured | 0.142961 | Uncertain | 0.944 | 0.151 | 0.000 | -10.173 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 0.74 | Ambiguous | 0.1 | 2.12 | Destabilizing | 1.43 | Ambiguous | 1.20 | Destabilizing | 0.879 | Likely Pathogenic | -5.97 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | -2.06 | Pathogenic | 0.13 | Tolerated | 0.2232 | 0.2596 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1791C>G | F597L 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant F597L is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: the benign group contains only SIFT, while the pathogenic group includes SGM‑Consensus (Likely Pathogenic), REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. FoldX and Foldetta are inconclusive, providing no definitive evidence. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts a damaging effect, SGM‑Consensus concurs with a likely pathogenic classification, and Foldetta remains uncertain. Taken together, the overwhelming majority of predictions indicate a pathogenic impact, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.010926 | Structured | 0.142961 | Uncertain | 0.944 | 0.151 | 0.000 | -10.173 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 0.74 | Ambiguous | 0.1 | 2.12 | Destabilizing | 1.43 | Ambiguous | 1.20 | Destabilizing | 0.879 | Likely Pathogenic | -5.97 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | -2.06 | Pathogenic | 0.13 | Tolerated | 0.2232 | 0.2596 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1876A>T | I626F 2D  3DClick to see structure in 3D Viewer AISynGAP1 I626F is not reported in ClinVar and is absent from gnomAD. Prediction tools that classify the variant as benign include only FATHMM, whereas the remaining tools—SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS (uncertain), PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default—predict it to be pathogenic. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is uncertain, but the SGM‑Consensus (derived from a majority of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is pathogenic, and Foldetta, which integrates FoldX‑MD and Rosetta outputs, also predicts pathogenicity. Taken together, the overwhelming majority of evidence indicates that I626F is likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.109221 | Structured | 0.040732 | Uncertain | 0.970 | 0.223 | 0.000 | -14.483 | Likely Pathogenic | 0.952 | Likely Pathogenic | Ambiguous | 4.37 | Destabilizing | 0.3 | 2.12 | Destabilizing | 3.25 | Destabilizing | 0.66 | Ambiguous | 0.631 | Likely Pathogenic | -3.78 | Deleterious | 0.999 | Probably Damaging | 0.993 | Probably Damaging | 3.07 | Benign | 0.00 | Affected | 0.0481 | 0.1964 | 1 | 0 | -1.7 | 34.02 | |||||||||||||||||||||||||||||
| c.2050G>T | D684Y 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant D684Y is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that assess the variant’s effect largely agree on a deleterious outcome: SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict pathogenicity. Only premPS and FATHMM predict a benign effect. High‑accuracy methods reinforce the pathogenic signal: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. No prediction or folding stability result is missing or inconclusive. Based on the overwhelming consensus of pathogenic predictions, the variant is most likely pathogenic, and this assessment does not contradict the absence of ClinVar reporting. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.254060 | Structured | 0.153798 | Uncertain | 0.870 | 0.282 | 0.000 | -15.224 | Likely Pathogenic | 0.994 | Likely Pathogenic | Likely Pathogenic | 3.65 | Destabilizing | 1.5 | 2.12 | Destabilizing | 2.89 | Destabilizing | -0.06 | Likely Benign | 0.600 | Likely Pathogenic | -8.98 | Deleterious | 1.000 | Probably Damaging | 0.963 | Probably Damaging | 3.44 | Benign | 0.00 | Affected | 0.0575 | 0.6564 | -4 | -3 | 2.2 | 48.09 | |||||||||||||||||||||||||||||
| c.878G>A | R293H 2D  AISynGAP1 missense variant R293H is listed in ClinVar with an uncertain significance (ClinVar ID 3901513.0) and is not reported in gnomAD. Prediction tools that indicate a benign effect include REVEL and premPS, whereas the remaining 13 tools—FoldX, Rosetta, Foldetta, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus—predict a pathogenic outcome. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized scores the variant as pathogenic; the SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also reports pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta stability predictions, classifies the variant as pathogenic. Overall, the preponderance of evidence indicates that R293H is most likely pathogenic, a conclusion that does not contradict the current ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.335645 | Structured | 0.338192 | Uncertain | 0.924 | 0.269 | 0.125 | Uncertain | 1 | -13.009 | Likely Pathogenic | 0.973 | Likely Pathogenic | Likely Pathogenic | 4.45 | Destabilizing | 2.3 | 2.12 | Destabilizing | 3.29 | Destabilizing | 0.32 | Likely Benign | 0.438 | Likely Benign | -4.60 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 1.45 | Pathogenic | 0.04 | Affected | 0.3120 | 0.2573 | 2 | 0 | 1.3 | -19.05 | |||||||||||||||||||||||||||
| c.1262C>G | A421G 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant A421G is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, polyPhen‑2 HumVar, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are Rosetta, premPS, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). FoldX and Foldetta give uncertain results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM Consensus as likely pathogenic, and Foldetta as uncertain. Overall, the majority of tools and the SGM Consensus favor a pathogenic interpretation, while a minority suggest benign. Because there is no ClinVar entry, the predictions do not contradict existing clinical classification. The variant is most likely pathogenic based on the collective computational evidence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.067594 | Structured | 0.404927 | Uncertain | 0.965 | 0.257 | 0.000 | -9.699 | Likely Pathogenic | 0.757 | Likely Pathogenic | Likely Benign | 1.47 | Ambiguous | 0.1 | 2.13 | Destabilizing | 1.80 | Ambiguous | 1.19 | Destabilizing | 0.137 | Likely Benign | -3.59 | Deleterious | 0.536 | Possibly Damaging | 0.176 | Benign | 3.41 | Benign | 0.05 | Affected | 0.1692 | 0.2499 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.1466T>A | L489H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L489H is not reported in ClinVar (ClinVar status: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that assess pathogenicity all agree that the variant is deleterious: SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify it as pathogenic. No tool predicts a benign effect. High‑accuracy methods reinforce this consensus: AlphaMissense‑Optimized predicts pathogenicity; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also predicts pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, indicates a destabilizing, pathogenic effect. All available predictions are concordant and supportive. Therefore, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation (none exists). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.191378 | Structured | 0.326126 | Uncertain | 0.949 | 0.234 | 0.125 | -15.946 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 3.12 | Destabilizing | 0.2 | 2.13 | Destabilizing | 2.63 | Destabilizing | 1.99 | Destabilizing | 0.919 | Likely Pathogenic | -6.74 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.60 | Pathogenic | 0.00 | Affected | 0.1132 | 0.0871 | -2 | -3 | -7.0 | 23.98 | |||||||||||||||||||||||||||||
| c.1681T>A | F561I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F561I is not reported in ClinVar (ClinVar ID None) but is present in gnomAD (variant ID 6‑33440733‑T‑A). Prediction tools that agree on a benign effect include only SIFT. All other evaluated predictors—SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized—classify the variant as pathogenic. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also predicts pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. No prediction or folding stability result is missing or inconclusive. Based on the overwhelming consensus of pathogenic predictions, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.024826 | Structured | 0.018013 | Uncertain | 0.903 | 0.196 | 0.000 | 6-33440733-T-A | 1 | 6.32e-7 | -11.708 | Likely Pathogenic | 0.990 | Likely Pathogenic | Likely Pathogenic | 3.82 | Destabilizing | 0.1 | 2.13 | Destabilizing | 2.98 | Destabilizing | 1.46 | Destabilizing | 0.710 | Likely Pathogenic | -5.97 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | -1.06 | Pathogenic | 0.09 | Tolerated | 3.37 | 35 | 0.1825 | 0.1925 | 0 | 1 | 1.7 | -34.02 | ||||||||||||||||||||||||
| c.1751T>A | I584N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I584N is not reported in ClinVar (ClinVar status: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that assess pathogenicity all agree that the variant is deleterious: SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify it as pathogenic. No tool predicts a benign effect. High‑accuracy methods reinforce this consensus: AlphaMissense‑Optimized predicts pathogenicity; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also predicts pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, indicates a pathogenic impact. All available predictions are concordant and supportive. Based on these computational assessments, the variant is most likely pathogenic, and this conclusion does not contradict any existing ClinVar annotation (none is present). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.059222 | Structured | 0.046673 | Uncertain | 0.846 | 0.244 | 0.000 | -13.153 | Likely Pathogenic | 0.962 | Likely Pathogenic | Likely Pathogenic | 2.70 | Destabilizing | 0.1 | 2.13 | Destabilizing | 2.42 | Destabilizing | 2.08 | Destabilizing | 0.706 | Likely Pathogenic | -6.57 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | -1.18 | Pathogenic | 0.01 | Affected | 0.0693 | 0.0470 | -2 | -3 | -8.0 | 0.94 | |||||||||||||||||||||||||||||
| c.1847A>G | D616G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D616G missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, SIFT, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are Rosetta, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and ESM1b. Tools with uncertain or inconclusive results—FoldX, AlphaMissense‑Default, and Foldetta—are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the predictions are split evenly between benign and pathogenic, with no clear consensus. Thus, the variant is most likely of uncertain significance; it does not contradict any ClinVar status because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.129801 | Structured | 0.166689 | Uncertain | 0.867 | 0.252 | 0.000 | -10.310 | Likely Pathogenic | 0.547 | Ambiguous | Likely Benign | 1.48 | Ambiguous | 0.1 | 2.13 | Destabilizing | 1.81 | Ambiguous | 0.49 | Likely Benign | 0.144 | Likely Benign | -5.60 | Deleterious | 0.985 | Probably Damaging | 0.800 | Possibly Damaging | 3.37 | Benign | 0.06 | Tolerated | 0.3496 | 0.4386 | 1 | -1 | 3.1 | -58.04 | ||||||||||||||||||||||||||||||
| c.874G>T | A292S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A292S is not reported in ClinVar (ClinVar ID None) and has no entries in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL and FoldX, whereas a majority of tools predict a pathogenic impact: SGM‑Consensus (Likely Pathogenic), PolyPhen‑2 HumDiv, PolyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and PROVEAN. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as Likely Pathogenic (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN), and Foldetta as uncertain. No prediction or folding‑stability result is missing; uncertain outcomes are treated as unavailable. Overall, the balance of evidence favors a pathogenic classification. This conclusion does not contradict ClinVar status, as no ClinVar assertion exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.247041 | Structured | 0.362042 | Uncertain | 0.929 | 0.256 | 0.000 | -8.514 | Likely Pathogenic | 0.804 | Likely Pathogenic | Ambiguous | 0.40 | Likely Benign | 0.2 | 2.13 | Destabilizing | 1.27 | Ambiguous | 0.70 | Ambiguous | 0.364 | Likely Benign | -2.76 | Deleterious | 0.999 | Probably Damaging | 0.996 | Probably Damaging | 1.69 | Pathogenic | 0.03 | Affected | 0.2850 | 0.6389 | 1 | 1 | -2.6 | 16.00 | |||||||||||||||||||||||||||||
| c.1805T>C | I602T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I602T is not reported in ClinVar and is absent from gnomAD. Functional prediction tools uniformly indicate a deleterious effect: REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) all classify it as pathogenic or likely pathogenic. No tool reports a benign outcome. High‑accuracy assessments corroborate this trend: AlphaMissense‑Optimized is uncertain, SGM‑Consensus is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenicity. Consequently, the variant is most likely pathogenic according to the available computational evidence, and this assessment does not contradict any ClinVar annotation (none exists). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.010221 | Structured | 0.186541 | Uncertain | 0.963 | 0.171 | 0.000 | -12.238 | Likely Pathogenic | 0.948 | Likely Pathogenic | Ambiguous | 2.39 | Destabilizing | 0.1 | 2.14 | Destabilizing | 2.27 | Destabilizing | 1.94 | Destabilizing | 0.931 | Likely Pathogenic | -4.82 | Deleterious | 1.000 | Probably Damaging | 0.996 | Probably Damaging | -2.00 | Pathogenic | 0.00 | Affected | 0.0890 | 0.0989 | 0 | -1 | -5.2 | -12.05 | |||||||||||||||||||||||||||||
| c.1813C>A | P605T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P605T is not reported in ClinVar (ClinVar status: none) and is absent from gnomAD (gnomAD ID: none). Prediction tools that agree on a pathogenic effect include SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized; no tool predicts a benign outcome. High‑accuracy methods specifically show pathogenicity: AlphaMissense‑Optimized predicts pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also predicts pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. No predictions or stability results are missing or inconclusive. Based on the unanimous pathogenic predictions and the absence of any ClinVar or gnomAD evidence to the contrary, the variant is most likely pathogenic and does not contradict ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.023087 | Structured | 0.192737 | Uncertain | 0.929 | 0.231 | 0.000 | -11.533 | Likely Pathogenic | 0.989 | Likely Pathogenic | Likely Pathogenic | 2.87 | Destabilizing | 0.4 | 2.14 | Destabilizing | 2.51 | Destabilizing | 0.72 | Ambiguous | 0.801 | Likely Pathogenic | -7.96 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 0.69 | Pathogenic | 0.00 | Affected | 0.1626 | 0.5228 | 0 | -1 | 0.9 | 3.99 | |||||||||||||||||||||||||||||
| c.1822T>A | F608I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F608I is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that assess pathogenicity all converge on a deleterious effect: REVEL, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict pathogenic or likely pathogenic. The only tool with an inconclusive result is FoldX, which is listed as uncertain. No tool predicts a benign outcome. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, is pathogenic. Based on the collective evidence, the variant is most likely pathogenic, and this conclusion is consistent with the absence of any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.106997 | Structured | 0.197190 | Uncertain | 0.891 | 0.247 | 0.000 | -14.939 | Likely Pathogenic | 0.991 | Likely Pathogenic | Likely Pathogenic | 1.92 | Ambiguous | 0.1 | 2.14 | Destabilizing | 2.03 | Destabilizing | 1.24 | Destabilizing | 0.904 | Likely Pathogenic | -5.97 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | -1.62 | Pathogenic | 0.00 | Affected | 0.2115 | 0.2361 | 1 | 0 | 1.7 | -34.02 | |||||||||||||||||||||||||||||
| c.649G>A | E217K 2D  3DClick to see structure in 3D Viewer AISynGAP1 E217K is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Computational predictions cluster into two groups: benign calls from premPS, PROVEAN, polyPhen‑2 HumVar, SIFT, and FATHMM; pathogenic calls from REVEL, Rosetta, polyPhen‑2 HumDiv, AlphaMissense‑Default, and AlphaMissense‑Optimized. Three tools give uncertain results: FoldX, Foldetta, and ESM1b. High‑accuracy methods give conflicting outcomes: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) yields benign; Foldetta remains uncertain. Because the majority of standard tools are split evenly and the high‑accuracy predictions are discordant, the evidence does not decisively support either outcome. The variant is therefore most likely pathogenic based on the preponderance of pathogenic predictions, and this assessment does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.278302 | Structured | 0.404912 | Uncertain | 0.823 | 0.284 | 0.000 | -7.169 | In-Between | 0.990 | Likely Pathogenic | Likely Pathogenic | 0.52 | Ambiguous | 0.5 | 2.14 | Destabilizing | 1.33 | Ambiguous | 0.45 | Likely Benign | 0.563 | Likely Pathogenic | -2.38 | Neutral | 0.900 | Possibly Damaging | 0.307 | Benign | 5.95 | Benign | 0.13 | Tolerated | 0.2742 | 0.8216 | 0 | 1 | -0.4 | -0.94 | ||||||||||||||||||||||||||||||
| c.1046C>G | P349R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P349R is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only REVEL, whereas the majority of tools predict a pathogenic impact: PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and Rosetta. Uncertain or inconclusive results come from FoldX, Foldetta, premPS, and AlphaMissense‑Optimized. High‑accuracy assessments show that AlphaMissense‑Optimized is uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is Likely Pathogenic, and Foldetta remains uncertain. Overall, the preponderance of evidence points to a pathogenic effect for P349R. This conclusion is not contradicted by ClinVar status, as the variant has no ClinVar entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.167087 | Structured | 0.348607 | Uncertain | 0.947 | 0.396 | 0.000 | -14.001 | Likely Pathogenic | 0.791 | Likely Pathogenic | Ambiguous | 0.99 | Ambiguous | 0.1 | 2.15 | Destabilizing | 1.57 | Ambiguous | 0.93 | Ambiguous | 0.335 | Likely Benign | -7.22 | Deleterious | 1.000 | Probably Damaging | 0.997 | Probably Damaging | 1.55 | Pathogenic | 0.01 | Affected | 0.1395 | 0.3009 | 0 | -2 | -2.9 | 59.07 | |||||||||||||||||||||||||||||
| c.1151G>A | G384D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G384D is not reported in ClinVar (ClinVar status: none) but is present in gnomAD (ID 6‑33438056‑G‑A). Prediction tools that classify the variant as benign include REVEL, PROVEAN, and AlphaMissense‑Optimized. Those that predict pathogenicity are FoldX, Rosetta, Foldetta, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, and AlphaMissense‑Default; premPS is uncertain. Separately, the high‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as pathogenic. Overall, the majority of tools—including the high‑accuracy methods—indicate a pathogenic effect. This prediction does not contradict any ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.728858 | Disordered | 0.427831 | Uncertain | 0.323 | 0.934 | 0.750 | 6-33438056-G-A | -9.142 | Likely Pathogenic | 0.610 | Likely Pathogenic | Likely Benign | 2.06 | Destabilizing | 0.5 | 2.15 | Destabilizing | 2.11 | Destabilizing | 0.53 | Ambiguous | 0.439 | Likely Benign | -0.93 | Neutral | 0.994 | Probably Damaging | 0.986 | Probably Damaging | 1.32 | Pathogenic | 0.04 | Affected | 4.32 | 2 | 0.2071 | 0.2235 | -1 | 1 | -3.1 | 58.04 | ||||||||||||||||||||||||||
| c.1216T>C | Y406H 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant Y406H is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL and FATHMM, whereas the majority of other in silico predictors—FoldX, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default—consistently classify the substitution as pathogenic. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized is inconclusive, SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) reports a likely pathogenic outcome, and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, predicts a pathogenic effect. Taken together, the preponderance of evidence indicates that Y406H is most likely pathogenic, and this conclusion does not conflict with the absence of a ClinVar entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.179055 | Structured | 0.393707 | Uncertain | 0.946 | 0.291 | 0.000 | -9.015 | Likely Pathogenic | 0.858 | Likely Pathogenic | Ambiguous | 2.13 | Destabilizing | 0.1 | 2.15 | Destabilizing | 2.14 | Destabilizing | 1.03 | Destabilizing | 0.239 | Likely Benign | -4.21 | Deleterious | 0.997 | Probably Damaging | 0.966 | Probably Damaging | 3.80 | Benign | 0.03 | Affected | 0.2666 | 0.0811 | 0 | 2 | -1.9 | -26.03 | |||||||||||||||||||||||||||||
| c.1789T>A | F597I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F597I is not reported in ClinVar and is absent from gnomAD. Functional prediction tools uniformly indicate a deleterious effect. Benign predictions: none. Pathogenic predictions: SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments corroborate this: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. All available evidence points to a pathogenic effect. Therefore, the variant is most likely pathogenic, and this conclusion is consistent with the absence of a ClinVar entry. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.010926 | Structured | 0.142961 | Uncertain | 0.944 | 0.151 | 0.000 | -13.674 | Likely Pathogenic | 0.986 | Likely Pathogenic | Likely Pathogenic | 3.62 | Destabilizing | 0.9 | 2.15 | Destabilizing | 2.89 | Destabilizing | 1.45 | Destabilizing | 0.951 | Likely Pathogenic | -5.97 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | -2.18 | Pathogenic | 0.01 | Affected | 0.2008 | 0.1815 | 1 | 0 | 1.7 | -34.02 | |||||||||||||||||||||||||||||
| c.810G>C | E270D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E270D missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only REVEL, which scores the variant as benign. The majority of tools predict a pathogenic impact: PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN), and Rosetta. Tools with inconclusive results—FoldX, Foldetta, and premPS—are listed as uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as likely pathogenic, and Foldetta as uncertain. Overall, the preponderance of evidence indicates that the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.144935 | Structured | 0.382573 | Uncertain | 0.938 | 0.231 | 0.125 | -11.954 | Likely Pathogenic | 0.976 | Likely Pathogenic | Likely Pathogenic | 1.52 | Ambiguous | 0.1 | 2.15 | Destabilizing | 1.84 | Ambiguous | 0.91 | Ambiguous | 0.379 | Likely Benign | -2.76 | Deleterious | 0.997 | Probably Damaging | 0.992 | Probably Damaging | 1.60 | Pathogenic | 0.02 | Affected | 0.1908 | 0.2540 | 3 | 2 | 0.0 | -14.03 | |||||||||||||||||||||||||||||
| c.810G>T | E270D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E270D missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only REVEL, which scores the variant as benign. The majority of tools predict a pathogenic impact: PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, Rosetta, and the SGM‑Consensus (which is “Likely Pathogenic”). Predictions that are inconclusive or uncertain are FoldX, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus also indicating a likely pathogenic outcome, while Foldetta’s stability analysis is uncertain. Overall, the preponderance of evidence points to a pathogenic effect for E270D, and this conclusion is consistent with the lack of ClinVar annotation (no contradiction). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.144935 | Structured | 0.382573 | Uncertain | 0.938 | 0.231 | 0.125 | -11.954 | Likely Pathogenic | 0.976 | Likely Pathogenic | Likely Pathogenic | 1.52 | Ambiguous | 0.1 | 2.15 | Destabilizing | 1.84 | Ambiguous | 0.91 | Ambiguous | 0.383 | Likely Benign | -2.76 | Deleterious | 0.997 | Probably Damaging | 0.992 | Probably Damaging | 1.60 | Pathogenic | 0.02 | Affected | 0.1908 | 0.2540 | 3 | 2 | 0.0 | -14.03 | |||||||||||||||||||||||||||||
| c.1151G>C | G384A 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant G384A is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that classify the variant as benign include REVEL, premPS, PROVEAN, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Benign). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, FATHMM, and Rosetta. Predictions from FoldX and Foldetta are uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus as Likely Benign, and Foldetta as uncertain. Overall, the majority of evidence points to a benign impact, and this is consistent with the lack of ClinVar annotation. Therefore, the variant is most likely benign and does not contradict ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.728858 | Disordered | 0.427831 | Uncertain | 0.323 | 0.934 | 0.750 | -6.927 | Likely Benign | 0.112 | Likely Benign | Likely Benign | 1.81 | Ambiguous | 0.2 | 2.16 | Destabilizing | 1.99 | Ambiguous | 0.04 | Likely Benign | 0.307 | Likely Benign | -0.40 | Neutral | 0.953 | Possibly Damaging | 0.952 | Probably Damaging | 1.33 | Pathogenic | 0.05 | Affected | 0.3986 | 0.4576 | 1 | 0 | 2.2 | 14.03 | |||||||||||||||||||||||||||||
| c.2174T>G | L725R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L725R is not reported in ClinVar (ClinVar ID = None) and is absent from gnomAD (gnomAD ID = None). Prediction tools that agree on a benign effect include only REVEL, which scores the variant as benign. All other evaluated algorithms—polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, PROVEAN, AlphaMissense‑Default, AlphaMissense‑Optimized, premPS, Rosetta, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN)—classify the variant as pathogenic. FoldX and Foldetta report uncertain results and are therefore not considered evidence for either side. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts pathogenic, SGM‑Consensus indicates likely pathogenic, while Foldetta remains uncertain. Based on the overwhelming majority of pathogenic predictions and the lack of benign consensus, the variant is most likely pathogenic, which is consistent with the absence of a ClinVar entry and gnomAD observation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | 0.557691 | Disordered | 0.455613 | Uncertain | 0.911 | 0.491 | 0.625 | -15.383 | Likely Pathogenic | 0.961 | Likely Pathogenic | Likely Pathogenic | 0.69 | Ambiguous | 0.3 | 2.16 | Destabilizing | 1.43 | Ambiguous | 1.49 | Destabilizing | 0.345 | Likely Benign | -5.46 | Deleterious | 0.999 | Probably Damaging | 0.998 | Probably Damaging | 1.28 | Pathogenic | 0.00 | Affected | 0.1374 | 0.0846 | -3 | -2 | -8.3 | 43.03 | ||||||||||||||||||||||||||||||
| c.2069C>T | S690F 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S690F is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL and FATHMM, whereas the majority of algorithms predict a pathogenic outcome: FoldX, Rosetta, Foldetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). The high‑accuracy assessments are consistent: AlphaMissense‑Optimized predicts pathogenic, the SGM‑Consensus indicates likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenic. No predictions are inconclusive or missing. Based on the collective evidence, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.055536 | Structured | 0.247926 | Uncertain | 0.944 | 0.253 | 0.000 | -14.325 | Likely Pathogenic | 0.992 | Likely Pathogenic | Likely Pathogenic | 9.85 | Destabilizing | 2.4 | 2.17 | Destabilizing | 6.01 | Destabilizing | 0.51 | Ambiguous | 0.384 | Likely Benign | -5.76 | Deleterious | 0.999 | Probably Damaging | 0.935 | Probably Damaging | 3.39 | Benign | 0.00 | Affected | 0.0498 | 0.4800 | -3 | -2 | 3.6 | 60.10 | |||||||||||||||||||||||||||||
| c.2120C>T | A707V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 A707V variant is not reported in ClinVar (no ClinVar ID) but is present in gnomAD (variant ID 6‑33441585‑C‑T). Functional prediction tools largely agree on a benign effect: REVEL, FoldX, premPS, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict benign, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also yields a benign classification. Only three tools—Rosetta, polyPhen‑2 HumDiv, and polyPhen‑2 HumVar—predict pathogenicity, while Foldetta reports an uncertain stability change. High‑accuracy assessments reinforce the benign prediction: AlphaMissense‑Optimized is benign, the SGM‑Consensus is benign, and Foldetta remains uncertain. Overall, the majority of evidence supports a benign impact for A707V, and this conclusion does not contradict any ClinVar status because no ClinVar assertion exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.203355 | Structured | 0.371229 | Uncertain | 0.927 | 0.365 | 0.000 | 6-33441585-C-T | 1 | 6.20e-7 | -6.479 | Likely Benign | 0.277 | Likely Benign | Likely Benign | 0.05 | Likely Benign | 0.0 | 2.17 | Destabilizing | 1.11 | Ambiguous | -0.30 | Likely Benign | 0.212 | Likely Benign | -1.62 | Neutral | 0.991 | Probably Damaging | 0.912 | Probably Damaging | 3.45 | Benign | 1.00 | Tolerated | 3.50 | 9 | 0.0794 | 0.4048 | 0 | 0 | 2.4 | 28.05 | ||||||||||||||||||||||||
| c.697T>C | C233R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C233R is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that agree on a benign effect include polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. In contrast, the majority of tools predict a pathogenic impact: SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy methods reinforce the pathogenic prediction: AlphaMissense‑Optimized is pathogenic, SGM‑Consensus is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. No predictions are missing or inconclusive. Based on the overall consensus of the majority of tools and the high‑accuracy methods, the variant is most likely pathogenic, which is consistent with the lack of ClinVar annotation and gnomAD presence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.239899 | Structured | 0.306787 | Uncertain | 0.868 | 0.322 | 0.000 | -16.789 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 3.79 | Destabilizing | 3.4 | 2.17 | Destabilizing | 2.98 | Destabilizing | 1.75 | Destabilizing | 0.830 | Likely Pathogenic | -10.68 | Deleterious | 0.002 | Benign | 0.002 | Benign | 5.71 | Benign | 0.01 | Affected | 0.1733 | 0.2212 | -4 | -3 | -7.0 | 53.05 | |||||||||||||||||||||||||||||
| c.980T>A | L327Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L327Q is not reported in ClinVar and is absent from gnomAD. Prediction tools that assess pathogenicity all converge on a deleterious effect: SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all indicate a pathogenic or likely pathogenic outcome. No tool in the dataset predicts a benign effect. High‑accuracy methods reinforce this consensus: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, is pathogenic. With all available evidence pointing to a harmful impact and no ClinVar entry to contradict this, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.335645 | Structured | 0.409189 | Uncertain | 0.939 | 0.490 | 0.000 | -14.243 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 3.03 | Destabilizing | 0.1 | 2.17 | Destabilizing | 2.60 | Destabilizing | 2.11 | Destabilizing | 0.605 | Likely Pathogenic | -5.26 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.52 | Pathogenic | 0.00 | Affected | 0.1131 | 0.1042 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.989A>G | D330G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant D330G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools that agree on a benign effect include REVEL, whereas the majority of algorithms predict a pathogenic impact: FoldX, Rosetta, Foldetta, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, and AlphaMissense‑Default. Two predictors (AlphaMissense‑Optimized and premPS) give uncertain results. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized is inconclusive, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is “Likely Pathogenic,” and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, classifies the variant as pathogenic. Overall, the preponderance of evidence indicates that D330G is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.380708 | Structured | 0.360008 | Uncertain | 0.805 | 0.488 | 0.250 | -13.064 | Likely Pathogenic | 0.950 | Likely Pathogenic | Ambiguous | 2.69 | Destabilizing | 0.2 | 2.17 | Destabilizing | 2.43 | Destabilizing | 0.63 | Ambiguous | 0.373 | Likely Benign | -5.03 | Deleterious | 0.980 | Probably Damaging | 0.782 | Possibly Damaging | 0.94 | Pathogenic | 0.01 | Affected | 0.3935 | 0.5015 | 1 | -1 | 3.1 | -58.04 | |||||||||||||||||||||||||||||
| c.1382C>A | A461D 2D  AIThe SynGAP1 missense variant A461D is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL and FATHMM. The majority of tools predict a pathogenic impact: premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). FoldX and Foldetta are inconclusive, providing no definitive evidence. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as likely pathogenic, and Foldetta remains uncertain. Overall, the preponderance of evidence indicates that A461D is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.179055 | Structured | 0.292531 | Uncertain | 0.936 | 0.151 | 0.125 | -14.918 | Likely Pathogenic | 0.990 | Likely Pathogenic | Likely Pathogenic | 0.89 | Ambiguous | 1.1 | 2.18 | Destabilizing | 1.54 | Ambiguous | 1.09 | Destabilizing | 0.477 | Likely Benign | -5.47 | Deleterious | 0.997 | Probably Damaging | 0.792 | Possibly Damaging | 3.32 | Benign | 0.01 | Affected | 0.1533 | 0.2016 | 0 | -2 | -5.3 | 44.01 | |||||||||||||||||||||||||||||
| c.1537T>A | F513I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F513I is not reported in ClinVar and has no entries in gnomAD. Functional prediction tools largely converge on a deleterious effect: SIFT is the sole benign caller, whereas REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict pathogenicity. Grouping by consensus, the single benign prediction (SIFT) is outweighed by the 13 pathogenic calls. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized reports a pathogenic effect; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is labeled Likely Pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, also predicts a pathogenic outcome. Consequently, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.102787 | Structured | 0.250651 | Uncertain | 0.949 | 0.269 | 0.000 | -12.003 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 3.42 | Destabilizing | 0.4 | 2.18 | Destabilizing | 2.80 | Destabilizing | 1.12 | Destabilizing | 0.766 | Likely Pathogenic | -5.70 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | -1.24 | Pathogenic | 0.22 | Tolerated | 0.1473 | 0.1766 | 1 | 0 | 1.7 | -34.02 | |||||||||||||||||||||||||||||
| c.1630C>G | R544G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R544G is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Prediction tools cluster into two groups: the single benign prediction comes from SIFT, while the remaining eleven tools (REVEL, FoldX, Rosetta, Foldetta, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized) all predict pathogenicity. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized scores the variant as pathogenic; the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—labels it Likely Pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, also predicts a pathogenic effect. Taken together, the consensus of the majority of tools and the high‑accuracy methods indicates that R544G is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.038858 | Structured | 0.016004 | Uncertain | 0.967 | 0.333 | 0.000 | -12.971 | Likely Pathogenic | 0.986 | Likely Pathogenic | Likely Pathogenic | 2.58 | Destabilizing | 0.2 | 2.18 | Destabilizing | 2.38 | Destabilizing | 0.74 | Ambiguous | 0.714 | Likely Pathogenic | -5.33 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.48 | Pathogenic | 0.09 | Tolerated | 0.2711 | 0.2123 | -3 | -2 | 4.1 | -99.14 | |||||||||||||||||||||||||||||
| c.1957C>G | L653V 2D  AIThe SynGAP1 missense variant L653V is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that classify the variant as benign include REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are FoldX, Rosetta, and premPS, while ESM1b is inconclusive. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as Likely Benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as pathogenic. Overall, the majority of evidence points to a benign impact, and this does not contradict the ClinVar “Uncertain” classification. Thus, based on current predictions, the variant is most likely benign. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.049374 | Structured | 0.335213 | Uncertain | 0.963 | 0.332 | 0.000 | Uncertain | 1 | -7.050 | In-Between | 0.301 | Likely Benign | Likely Benign | 3.28 | Destabilizing | 0.3 | 2.18 | Destabilizing | 2.73 | Destabilizing | 1.32 | Destabilizing | 0.146 | Likely Benign | -2.25 | Neutral | 0.227 | Benign | 0.039 | Benign | 3.28 | Benign | 0.08 | Tolerated | 0.1436 | 0.3557 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||
| c.602A>T | D201V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D201V missense variant has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, premPS, and FATHMM, while those that predict a pathogenic impact are SGM‑Consensus (Likely Pathogenic), Rosetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as Uncertain, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as Uncertain. With the majority of tools indicating pathogenicity and no ClinVar record to contradict this, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.366687 | Structured | 0.428570 | Uncertain | 0.698 | 0.447 | 0.125 | -10.283 | Likely Pathogenic | 0.906 | Likely Pathogenic | Ambiguous | 0.87 | Ambiguous | 0.1 | 2.18 | Destabilizing | 1.53 | Ambiguous | 0.31 | Likely Benign | 0.305 | Likely Benign | -5.01 | Deleterious | 0.999 | Probably Damaging | 0.946 | Probably Damaging | 4.04 | Benign | 0.02 | Affected | 0.0572 | 0.5207 | -2 | -3 | 7.7 | -15.96 | |||||||||||||||||||||||||||||
| c.788T>A | V263E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 V263E missense variant is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools largely agree on a deleterious effect: FATHMM predicts the variant as benign, while the remaining twelve tools—SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default—classify it as pathogenic. AlphaMissense‑Optimized is uncertain. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized remains uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates “Likely Pathogenic,” and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts a destabilizing, pathogenic effect. Overall, the preponderance of evidence points to a pathogenic impact for V263E, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.268042 | Structured | 0.356141 | Uncertain | 0.918 | 0.257 | 0.000 | -13.498 | Likely Pathogenic | 0.809 | Likely Pathogenic | Ambiguous | 2.04 | Destabilizing | 0.3 | 2.18 | Destabilizing | 2.11 | Destabilizing | 1.99 | Destabilizing | 0.862 | Likely Pathogenic | -3.84 | Deleterious | 0.999 | Probably Damaging | 0.991 | Probably Damaging | 5.96 | Benign | 0.01 | Affected | 0.0886 | 0.1436 | -2 | -2 | -7.7 | 29.98 | |||||||||||||||||||||||||||||
| c.1073T>G | F358C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F358C is not reported in ClinVar and is absent from gnomAD. Consensus from standard in‑silico predictors shows a split: benign calls come from REVEL, SIFT, and FATHMM, whereas pathogenic calls arise from Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar) and AlphaMissense‑Default. High‑accuracy assessments are less definitive: AlphaMissense‑Optimized is inconclusive, Foldetta is inconclusive, and the SGM Consensus—derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—leans toward pathogenic. Because the majority of available predictions favor a damaging effect, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation (none exists). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.222385 | Structured | 0.407113 | Uncertain | 0.912 | 0.441 | 0.250 | -7.966 | In-Between | 0.927 | Likely Pathogenic | Ambiguous | 1.68 | Ambiguous | 0.1 | 2.19 | Destabilizing | 1.94 | Ambiguous | 1.18 | Destabilizing | 0.460 | Likely Benign | -6.36 | Deleterious | 0.999 | Probably Damaging | 0.993 | Probably Damaging | 4.02 | Benign | 0.06 | Tolerated | 0.2364 | 0.1800 | -4 | -2 | -0.3 | -44.04 | ||||||||||||||||||||||||||||||
| c.1497A>C | R499S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R499S is catalogued in gnomAD (ID 6‑33438529‑A‑C) but has no ClinVar entry. Functional prediction tools largely converge on a deleterious effect: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) all indicate pathogenicity. No tool reports a benign outcome. High‑accuracy assessments are mixed: AlphaMissense‑Optimized and Foldetta are uncertain, whereas the SGM‑Consensus remains likely pathogenic. Protein‑stability predictions are inconclusive (FoldX uncertain, Rosetta pathogenic, Foldetta uncertain). Taken together, the overwhelming majority of evidence supports a pathogenic classification for R499S. This conclusion is consistent with the absence of a ClinVar status, so there is no contradiction with existing clinical annotations. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.071867 | Structured | 0.386723 | Uncertain | 0.899 | 0.146 | 0.000 | 6-33438529-A-C | 1 | 6.20e-7 | -9.559 | Likely Pathogenic | 0.935 | Likely Pathogenic | Ambiguous | 1.03 | Ambiguous | 0.0 | 2.19 | Destabilizing | 1.61 | Ambiguous | 1.40 | Destabilizing | 0.632 | Likely Pathogenic | -2.69 | Deleterious | 0.958 | Probably Damaging | 0.702 | Possibly Damaging | -1.43 | Pathogenic | 0.01 | Affected | 3.37 | 35 | 0.2443 | 0.1649 | -1 | 0 | 3.7 | -69.11 | ||||||||||||||||||||||||
| c.1497A>T | R499S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R499S is not reported in ClinVar and has no entry in gnomAD. Consensus from multiple in silico predictors indicates a pathogenic effect: SGM‑Consensus, REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and Rosetta all predict pathogenicity, while FoldX, AlphaMissense‑Optimized, and Foldetta are uncertain. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized is inconclusive, SGM‑Consensus (derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) reports likely pathogenic, and Foldetta likewise yields an uncertain result. Taken together, the overwhelming majority of reliable tools predict a pathogenic effect, and there is no ClinVar annotation to contradict this assessment. Therefore, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.071867 | Structured | 0.386723 | Uncertain | 0.899 | 0.146 | 0.000 | -9.559 | Likely Pathogenic | 0.935 | Likely Pathogenic | Ambiguous | 1.03 | Ambiguous | 0.0 | 2.19 | Destabilizing | 1.61 | Ambiguous | 1.40 | Destabilizing | 0.632 | Likely Pathogenic | -2.69 | Deleterious | 0.958 | Probably Damaging | 0.702 | Possibly Damaging | -1.43 | Pathogenic | 0.01 | Affected | 3.37 | 35 | 0.2443 | 0.1649 | -1 | 0 | 3.7 | -69.11 | |||||||||||||||||||||||||||
| c.1652T>A | L551Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L551Q is not reported in ClinVar and is present in gnomAD (allele ID 6‑33438895‑T‑A). In silico prediction tools uniformly indicate a deleterious effect: benign‑predicting tools: none; pathogenic‑predicting tools: REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. No predictions are inconclusive or missing. **Thus, the variant is most likely pathogenic based on the available predictions, and this conclusion does not contradict the ClinVar status (no entry).** Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.009977 | Structured | 0.006653 | Uncertain | 0.960 | 0.254 | 0.000 | 6-33438895-T-A | 1 | 6.20e-7 | -13.632 | Likely Pathogenic | 0.990 | Likely Pathogenic | Likely Pathogenic | 2.48 | Destabilizing | 0.1 | 2.19 | Destabilizing | 2.34 | Destabilizing | 2.37 | Destabilizing | 0.936 | Likely Pathogenic | -3.68 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.60 | Pathogenic | 0.01 | Affected | 3.37 | 35 | 0.0983 | 0.0688 | -2 | -2 | -7.3 | 14.97 | ||||||||||||||||||||||||
| c.971G>C | R324P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R324P is not reported in ClinVar and is absent from gnomAD. Prediction tools that indicate a benign effect include REVEL, PROVEAN, and SIFT, whereas the remaining tools (FoldX, Rosetta, Foldetta, premPS, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized) all predict a pathogenic outcome. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is “Likely Pathogenic” because three of the four contributing tools predict pathogenicity; and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, also predicts pathogenic. Based on the preponderance of pathogenic predictions—including the high‑accuracy tools—the variant is most likely pathogenic, and this conclusion is not contradicted by ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.257454 | Structured | 0.426893 | Uncertain | 0.954 | 0.397 | 0.000 | -14.533 | Likely Pathogenic | 0.993 | Likely Pathogenic | Likely Pathogenic | 4.61 | Destabilizing | 0.5 | 2.19 | Destabilizing | 3.40 | Destabilizing | 1.03 | Destabilizing | 0.486 | Likely Benign | -2.15 | Neutral | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.83 | Pathogenic | 0.29 | Tolerated | 0.2291 | 0.5556 | 0 | -2 | 2.9 | -59.07 | |||||||||||||||||||||||||||||
| c.1043T>A | V348E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V348E is not reported in ClinVar and is absent from gnomAD. Prediction tools cluster into two groups: the single benign prediction comes from REVEL, while all other evaluated algorithms (FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized) indicate pathogenicity. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is “Likely Pathogenic”; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, also predicts pathogenic. No prediction or stability result is missing or inconclusive. Based on the overwhelming agreement among the majority of tools, the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no entry for V348E. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.170161 | Structured | 0.346556 | Uncertain | 0.951 | 0.414 | 0.000 | -11.135 | Likely Pathogenic | 0.993 | Likely Pathogenic | Likely Pathogenic | 2.41 | Destabilizing | 0.1 | 2.20 | Destabilizing | 2.31 | Destabilizing | 2.14 | Destabilizing | 0.470 | Likely Benign | -5.26 | Deleterious | 0.989 | Probably Damaging | 0.637 | Possibly Damaging | 1.56 | Pathogenic | 0.01 | Affected | 0.1089 | 0.1922 | -2 | -2 | -7.7 | 29.98 | |||||||||||||||||||||||||||||
| c.1282T>C | Y428H 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant Y428H is not reported in ClinVar but is present in gnomAD (ID 6‑33438187‑T‑C). Prediction tools that indicate a benign effect include REVEL and FATHMM, whereas the majority of other in silico predictors (FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) and the SGM Consensus score (Likely Pathogenic) all suggest a deleterious impact. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. Consequently, the variant is most likely pathogenic, and this prediction does not contradict any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.118441 | Structured | 0.389652 | Uncertain | 0.965 | 0.292 | 0.000 | 6-33438187-T-C | 1 | 6.19e-7 | -8.566 | Likely Pathogenic | 0.961 | Likely Pathogenic | Likely Pathogenic | 3.18 | Destabilizing | 0.1 | 2.20 | Destabilizing | 2.69 | Destabilizing | 1.80 | Destabilizing | 0.458 | Likely Benign | -4.77 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.40 | Benign | 0.04 | Affected | 3.38 | 25 | 0.2685 | 0.0711 | 2 | 0 | -1.9 | -26.03 | ||||||||||||||||||||||||
| c.1402A>G | M468V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M468V is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from PROVEAN, SIFT, and AlphaMissense‑Optimized, while pathogenic predictions are made by REVEL, FoldX, Rosetta, Foldetta, polyPhen‑2 (HumDiv and HumVar), ESM1b, and FATHMM. The remaining tools, premPS and AlphaMissense‑Default, return uncertain results. High‑accuracy assessments further clarify the variant’s impact: AlphaMissense‑Optimized predicts a benign effect; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, indicates pathogenicity; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, also classifies the variant as pathogenic. Overall, the preponderance of evidence points to a pathogenic effect, which does not contradict the ClinVar designation of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.284882 | Structured | 0.339253 | Uncertain | 0.932 | 0.257 | 0.000 | Uncertain | 1 | -9.461 | Likely Pathogenic | 0.361 | Ambiguous | Likely Benign | 2.69 | Destabilizing | 0.1 | 2.20 | Destabilizing | 2.45 | Destabilizing | 0.89 | Ambiguous | 0.570 | Likely Pathogenic | -1.66 | Neutral | 0.998 | Probably Damaging | 0.993 | Probably Damaging | -1.21 | Pathogenic | 0.08 | Tolerated | 3.37 | 31 | 0.3309 | 0.3845 | 1 | 2 | 2.3 | -32.06 | ||||||||||||||||||||||||||
| c.1784T>G | L595R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L595R is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include only FATHMM, while the majority of tools (SGM‑Consensus, REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic impact; FoldX and Foldetta are uncertain. High‑accuracy methods give a pathogenic signal: AlphaMissense‑Optimized is pathogenic, SGM‑Consensus is likely pathogenic, and Foldetta remains uncertain. Overall, the evidence strongly favors a pathogenic effect, and this conclusion does not contradict any ClinVar annotation, as no ClinVar record exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.015344 | Structured | 0.128444 | Uncertain | 0.920 | 0.150 | 0.000 | -14.601 | Likely Pathogenic | 0.989 | Likely Pathogenic | Likely Pathogenic | 1.22 | Ambiguous | 0.0 | 2.20 | Destabilizing | 1.71 | Ambiguous | 1.52 | Destabilizing | 0.707 | Likely Pathogenic | -5.97 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 2.76 | Benign | 0.00 | Affected | 0.1344 | 0.1005 | -3 | -2 | -8.3 | 43.03 | |||||||||||||||||||||||||||||
| c.2036T>C | F679S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F679S is not reported in ClinVar and is absent from gnomAD. Prediction tools largely agree on a deleterious effect: benign predictions are limited to FATHMM, while the remaining 12 tools (SGM‑Consensus, REVEL, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) all classify the variant as pathogenic. FoldX reports an uncertain outcome and is therefore not counted as evidence. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenicity. Based on the preponderance of pathogenic predictions and the absence of any benign consensus, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status (none available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.222385 | Structured | 0.129316 | Uncertain | 0.700 | 0.320 | 0.000 | -12.159 | Likely Pathogenic | 0.963 | Likely Pathogenic | Likely Pathogenic | 1.86 | Ambiguous | 0.4 | 2.20 | Destabilizing | 2.03 | Destabilizing | 1.28 | Destabilizing | 0.575 | Likely Pathogenic | -7.86 | Deleterious | 0.998 | Probably Damaging | 0.986 | Probably Damaging | 3.50 | Benign | 0.04 | Affected | 0.4276 | 0.0200 | -3 | -2 | -3.6 | -60.10 | |||||||||||||||||||||||||||||
| c.653T>G | F218C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F218C is not reported in ClinVar (ClinVar ID None) but is present in gnomAD (ID 6‑33435295‑T‑G). Prediction tools that agree on a benign effect include only FATHMM, whereas the majority of tools (REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and AlphaMissense‑Default) predict a pathogenic impact. Results that are uncertain or unavailable are FoldX, ESM1b, AlphaMissense‑Optimized, and Foldetta. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) yields a pathogenic prediction (2 pathogenic vs. 1 benign votes); and Foldetta remains uncertain. Overall, the preponderance of evidence points to a pathogenic effect for F218C, and this conclusion does not contradict any ClinVar annotation, as none exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | PH | 0.281712 | Structured | 0.408725 | Uncertain | 0.848 | 0.272 | 0.000 | 6-33435295-T-G | 1 | 6.20e-7 | -7.234 | In-Between | 0.948 | Likely Pathogenic | Ambiguous | 1.49 | Ambiguous | 0.1 | 2.20 | Destabilizing | 1.85 | Ambiguous | 1.02 | Destabilizing | 0.744 | Likely Pathogenic | -4.92 | Deleterious | 0.994 | Probably Damaging | 0.667 | Possibly Damaging | 5.78 | Benign | 0.03 | Affected | 3.41 | 13 | 0.2330 | 0.1321 | -2 | -4 | -0.3 | -44.04 | |||||||||||||||||||||||||
| c.655T>C | C219R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C219R is not reported in ClinVar (ClinVar ID None) and has no entries in gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only FATHMM, while the remaining 13 tools—SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—consistently predict a pathogenic or likely pathogenic impact. High‑accuracy methods reinforce this view: AlphaMissense‑Optimized reports Pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates Likely Pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts Pathogenic. No prediction or folding stability result is missing or inconclusive. Based on the collective evidence, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.254060 | Structured | 0.426845 | Uncertain | 0.903 | 0.279 | 0.000 | -15.727 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 3.00 | Destabilizing | 1.5 | 2.20 | Destabilizing | 2.60 | Destabilizing | 1.33 | Destabilizing | 0.904 | Likely Pathogenic | -9.90 | Deleterious | 0.969 | Probably Damaging | 0.680 | Possibly Damaging | 5.80 | Benign | 0.00 | Affected | 0.1381 | 0.1837 | -4 | -3 | -7.0 | 53.05 | |||||||||||||||||||||||||||||
| c.801G>C | W267C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant W267C is not reported in ClinVar and is absent from gnomAD. All available in‑silico predictors classify it as pathogenic: SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool predicts a benign effect. High‑accuracy assessments reinforce this: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates Likely Pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. Thus, the variant is most likely pathogenic based on the consensus of all predictions, and this conclusion does not contradict the current ClinVar status, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.216401 | Structured | 0.298060 | Uncertain | 0.943 | 0.274 | 0.000 | -12.351 | Likely Pathogenic | 0.996 | Likely Pathogenic | Likely Pathogenic | 2.81 | Destabilizing | 0.2 | 2.20 | Destabilizing | 2.51 | Destabilizing | 1.00 | Destabilizing | 0.705 | Likely Pathogenic | -11.95 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.90 | Pathogenic | 0.01 | Affected | 0.3573 | 0.1700 | -8 | -2 | 3.4 | -83.07 | |||||||||||||||||||||||||||||
| c.801G>T | W267C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant W267C is not reported in ClinVar and is absent from gnomAD. All available in‑silico predictors classify it as pathogenic: SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool predicts a benign effect. High‑accuracy assessments reinforce this: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates Likely Pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. Thus, the variant is most likely pathogenic based on the consensus of all predictions, and this conclusion does not contradict the current ClinVar status, which has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.216401 | Structured | 0.298060 | Uncertain | 0.943 | 0.274 | 0.000 | -12.351 | Likely Pathogenic | 0.996 | Likely Pathogenic | Likely Pathogenic | 2.81 | Destabilizing | 0.2 | 2.20 | Destabilizing | 2.51 | Destabilizing | 1.00 | Destabilizing | 0.705 | Likely Pathogenic | -11.95 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 1.90 | Pathogenic | 0.01 | Affected | 0.3573 | 0.1700 | -8 | -2 | 3.4 | -83.07 | |||||||||||||||||||||||||||||
| c.1078G>A | E360K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E360K is reported in gnomAD (variant ID 6-33437983‑G‑A) but has no ClinVar entry. Prediction tools that agree on a benign effect are limited to FoldX, which scores the variant as benign. In contrast, the majority of algorithms predict a pathogenic impact: REVEL, Rosetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Pathogenic). Tools with inconclusive results (Foldetta and premPS) are noted as unavailable. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts pathogenic, SGM‑Consensus indicates likely pathogenic, while Foldetta remains uncertain. Overall, the consensus of high‑confidence predictors points to a pathogenic effect for E360K. This conclusion is not contradicted by ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.250310 | Structured | 0.421183 | Uncertain | 0.955 | 0.498 | 0.250 | 6-33437983-G-A | -16.006 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 0.27 | Likely Benign | 0.0 | 2.21 | Destabilizing | 1.24 | Ambiguous | 0.55 | Ambiguous | 0.526 | Likely Pathogenic | -3.68 | Deleterious | 0.997 | Probably Damaging | 0.980 | Probably Damaging | 1.68 | Pathogenic | 0.04 | Affected | 3.37 | 25 | 0.3106 | 0.8594 | 1 | 0 | -0.4 | -0.94 | ||||||||||||||||||||||||||
| c.1893G>C | Q631H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant Q631H is reported in gnomAD (6‑33440945‑G‑C) but has no ClinVar entry. Prediction tools that agree on a benign effect include REVEL and FATHMM, whereas the majority of other in silico predictors (Rosetta, PROVEAN, polyPhen‑2 HumDiv/HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) and the SGM‑Consensus score (Likely Pathogenic) all indicate a pathogenic impact. Uncertain results come from FoldX, Foldetta, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. Based on the preponderance of pathogenic predictions, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar classification exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.041405 | Structured | 0.038963 | Uncertain | 0.948 | 0.230 | 0.000 | 6-33440945-G-C | 2 | 1.24e-6 | -13.282 | Likely Pathogenic | 0.978 | Likely Pathogenic | Likely Pathogenic | 0.84 | Ambiguous | 0.2 | 2.21 | Destabilizing | 1.53 | Ambiguous | 0.84 | Ambiguous | 0.475 | Likely Benign | -4.98 | Deleterious | 0.995 | Probably Damaging | 0.986 | Probably Damaging | 2.75 | Benign | 0.00 | Affected | 3.37 | 34 | 0.0834 | 0.1757 | 0 | 3 | 0.3 | 9.01 | ||||||||||||||||||||||||
| c.1893G>T | Q631H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant Q631H is not reported in ClinVar and has no entries in gnomAD. Functional prediction tools show a split: benign predictions come from REVEL and FATHMM, whereas the majority of algorithms—including AlphaMissense‑Default, AlphaMissense‑Optimized, ESM1b, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, Rosetta, and the SGM Consensus—classify the change as pathogenic. Predictions from FoldX, Foldetta, and premPS are inconclusive. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates likely pathogenic, while Foldetta’s stability analysis is uncertain. Overall, the preponderance of evidence points to a pathogenic impact for Q631H, which is consistent with the absence of a benign ClinVar annotation and the lack of population data in gnomAD. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.041405 | Structured | 0.038963 | Uncertain | 0.948 | 0.230 | 0.000 | -13.282 | Likely Pathogenic | 0.978 | Likely Pathogenic | Likely Pathogenic | 0.84 | Ambiguous | 0.2 | 2.21 | Destabilizing | 1.53 | Ambiguous | 0.84 | Ambiguous | 0.475 | Likely Benign | -4.98 | Deleterious | 0.995 | Probably Damaging | 0.986 | Probably Damaging | 2.75 | Benign | 0.00 | Affected | 3.37 | 34 | 0.0834 | 0.1757 | 0 | 3 | 0.3 | 9.01 | |||||||||||||||||||||||||||
| c.2107C>A | L703I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L703I is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, FATHMM, and AlphaMissense‑Optimized, while those that agree on a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, and Rosetta. Predictions that are uncertain or inconclusive (FoldX, Foldetta, premPS, AlphaMissense‑Default) are treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as benign, and Foldetta as uncertain. Overall, the majority of high‑confidence tools predict a benign impact, and this conclusion does not contradict the ClinVar status, which has no pathogenic classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.144935 | Structured | 0.388282 | Uncertain | 0.929 | 0.353 | 0.000 | -9.332 | Likely Pathogenic | 0.345 | Ambiguous | Likely Benign | 1.44 | Ambiguous | 0.1 | 2.21 | Destabilizing | 1.83 | Ambiguous | 0.61 | Ambiguous | 0.108 | Likely Benign | -1.50 | Neutral | 0.982 | Probably Damaging | 0.758 | Possibly Damaging | 3.38 | Benign | 0.00 | Affected | 0.0909 | 0.3079 | 2 | 2 | 0.7 | 0.00 | ||||||||||||||||||||||||||||||
| c.1120T>C | S374P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S374P is reported in gnomAD (6‑33438025‑T‑C) but has no ClinVar entry. Functional prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are SIFT and Rosetta; FoldX and Foldetta are inconclusive. The high‑accuracy consensus (SGM‑Consensus) is “Likely Benign,” derived from the unanimous benign calls of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN. AlphaMissense‑Optimized also predicts benign, while Foldetta remains uncertain. Overall, the majority of evidence points to a benign impact. There is no ClinVar status to contradict this assessment. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.642678 | Disordered | 0.428948 | Uncertain | 0.333 | 0.812 | 0.625 | 6-33438025-T-C | 1 | 7.85e-7 | -4.849 | Likely Benign | 0.125 | Likely Benign | Likely Benign | 0.66 | Ambiguous | 0.6 | 2.22 | Destabilizing | 1.44 | Ambiguous | 0.34 | Likely Benign | 0.388 | Likely Benign | -0.89 | Neutral | 0.396 | Benign | 0.099 | Benign | 5.30 | Benign | 0.02 | Affected | 4.32 | 13 | 0.3012 | 0.6813 | -1 | 1 | -0.8 | 10.04 | ||||||||||||||||||||||||
| c.1406C>G | A469G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A469G is not reported in ClinVar (ClinVar status: not listed) and is absent from gnomAD. Prediction tools that agree on a benign effect include PROVEAN, SIFT, ESM1b, and AlphaMissense‑Optimized. Those that predict a pathogenic effect comprise REVEL, Rosetta, Foldetta, premPS, polyPhen‑2 (HumDiv and HumVar), FATHMM, and AlphaMissense‑Default; FoldX is uncertain. High‑accuracy methods give mixed results: AlphaMissense‑Optimized indicates benign, Foldetta indicates pathogenic, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive due to a 2‑to‑2 split. Overall, the majority of tools (12 vs. 4) predict pathogenicity, and there is no ClinVar entry to contradict this assessment. Thus, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.278302 | Structured | 0.343926 | Uncertain | 0.910 | 0.276 | 0.000 | -4.438 | Likely Benign | 0.733 | Likely Pathogenic | Likely Benign | 1.99 | Ambiguous | 0.1 | 2.22 | Destabilizing | 2.11 | Destabilizing | 1.01 | Destabilizing | 0.569 | Likely Pathogenic | -2.42 | Neutral | 0.997 | Probably Damaging | 0.990 | Probably Damaging | -1.32 | Pathogenic | 0.37 | Tolerated | 0.1797 | 0.3078 | 1 | 0 | -2.2 | -14.03 | ||||||||||||||||||||||||||||||
| c.1664T>G | V555G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 V555G missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on pathogenicity include REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default; no tool predicts it benign. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as pathogenic. No predictions are inconclusive or missing. Based on the collective evidence, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status because the variant is not yet catalogued there. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.013265 | Structured | 0.008218 | Uncertain | 0.943 | 0.225 | 0.000 | -13.327 | Likely Pathogenic | 0.899 | Likely Pathogenic | Ambiguous | 2.07 | Destabilizing | 0.1 | 2.22 | Destabilizing | 2.15 | Destabilizing | 1.07 | Destabilizing | 0.798 | Likely Pathogenic | -6.35 | Deleterious | 0.984 | Probably Damaging | 1.000 | Probably Damaging | -1.39 | Pathogenic | 0.01 | Affected | 0.1994 | 0.1677 | -1 | -3 | -4.6 | -42.08 | ||||||||||||||||||||||||||||||
| c.1868T>A | L623H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L623H is not reported in ClinVar and is absent from gnomAD. Prediction tools that assess pathogenicity all converge on a deleterious effect: REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict pathogenicity, while no tool predicts a benign outcome. High‑accuracy methods reinforce this consensus: AlphaMissense‑Optimized indicates pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) reports “Likely Pathogenic,” and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenic. No contradictory evidence is present. Based on the unanimous computational predictions, the variant is most likely pathogenic, and this assessment does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.175930 | Structured | 0.060667 | Uncertain | 0.962 | 0.211 | 0.000 | -14.631 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 2.76 | Destabilizing | 0.3 | 2.22 | Destabilizing | 2.49 | Destabilizing | 2.40 | Destabilizing | 0.793 | Likely Pathogenic | -6.97 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 1.55 | Pathogenic | 0.00 | Affected | 0.1099 | 0.0541 | -2 | -3 | -7.0 | 23.98 | |||||||||||||||||||||||||||||
| c.2044T>A | Y682N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 Y682N variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only FATHMM, while the majority of tools (SGM‑Consensus, REVEL, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) predict a pathogenic impact; FoldX and AlphaMissense‑Optimized are uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as likely pathogenic (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN), and Foldetta as pathogenic. Overall, the evidence strongly favors a pathogenic effect, and this conclusion does not contradict any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.206376 | Structured | 0.141467 | Uncertain | 0.758 | 0.328 | 0.000 | -11.734 | Likely Pathogenic | 0.859 | Likely Pathogenic | Ambiguous | 1.86 | Ambiguous | 0.1 | 2.22 | Destabilizing | 2.04 | Destabilizing | 1.54 | Destabilizing | 0.564 | Likely Pathogenic | -8.61 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.34 | Benign | 0.02 | Affected | 0.2354 | 0.0928 | -2 | -2 | -2.2 | -49.07 | |||||||||||||||||||||||||||||
| c.2131C>G | L711V 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L711V is listed in ClinVar with an uncertain significance and is present in gnomAD (ID 6‑33441596‑C‑G). Prediction tools that indicate a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. The majority of other in silico predictors—FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default—classify the change as pathogenic, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) reports it as likely pathogenic. High‑accuracy assessments further show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as pathogenic. Overall, the preponderance of evidence points to a pathogenic effect, which does not conflict with the ClinVar designation of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.308712 | Structured | 0.377436 | Uncertain | 0.950 | 0.364 | 0.000 | Uncertain | 1 | 6-33441596-C-G | 1 | 6.20e-7 | -10.045 | Likely Pathogenic | 0.709 | Likely Pathogenic | Likely Benign | 3.48 | Destabilizing | 0.1 | 2.22 | Destabilizing | 2.85 | Destabilizing | 1.40 | Destabilizing | 0.170 | Likely Benign | -2.59 | Deleterious | 0.992 | Probably Damaging | 0.970 | Probably Damaging | 3.34 | Benign | 0.00 | Affected | 3.50 | 9 | 0.1318 | 0.3010 | 1 | 2 | 0.4 | -14.03 | ||||||||||||||||||||||
| c.728T>G | I243S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I243S is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that assess pathogenicity largely agree: SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict a pathogenic effect. Only FATHMM predicts a benign outcome. High‑accuracy methods reinforce the pathogenic signal: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta) is pathogenic. No predictions are missing or inconclusive. Based on the consensus of these tools, the variant is most likely pathogenic, and this assessment does not contradict the absence of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.363090 | Structured | 0.344471 | Uncertain | 0.842 | 0.347 | 0.000 | -14.097 | Likely Pathogenic | 0.975 | Likely Pathogenic | Likely Pathogenic | 2.52 | Destabilizing | 0.2 | 2.22 | Destabilizing | 2.37 | Destabilizing | 1.71 | Destabilizing | 0.802 | Likely Pathogenic | -3.55 | Deleterious | 0.995 | Probably Damaging | 0.795 | Possibly Damaging | 5.52 | Benign | 0.00 | Affected | 0.2658 | 0.0600 | -1 | -2 | -5.3 | -26.08 | |||||||||||||||||||||||||||||
| c.820C>G | L274V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L274V is not reported in ClinVar and is absent from gnomAD. Prediction tools that indicate a benign effect include REVEL, ESM1b, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect comprise FoldX, Rosetta, Foldetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, and AlphaMissense‑Default. The SGM‑Consensus, which aggregates the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is classified as Likely Pathogenic. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while both the SGM‑Consensus and Foldetta (combining FoldX‑MD and Rosetta outputs) predict pathogenicity. No predictions are missing or inconclusive. Overall, the preponderance of evidence from multiple independent tools points to a pathogenic impact for L274V. This conclusion is consistent with the lack of a ClinVar entry, so there is no contradiction with existing clinical annotations. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.066181 | Structured | 0.377483 | Uncertain | 0.866 | 0.195 | 0.250 | -5.634 | Likely Benign | 0.593 | Likely Pathogenic | Likely Benign | 2.13 | Destabilizing | 0.6 | 2.22 | Destabilizing | 2.18 | Destabilizing | 0.99 | Ambiguous | 0.378 | Likely Benign | -2.56 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | 0.10 | Pathogenic | 0.02 | Affected | 0.1448 | 0.1860 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.967C>G | L323V 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L323V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools cluster into two groups: benign predictions (REVEL, premPS, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) and pathogenic predictions (FoldX, Rosetta, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, FATHMM). The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is labeled Likely Benign. High‑accuracy assessments further refine the picture: AlphaMissense‑Optimized predicts benign; the SGM‑Consensus (majority of the four high‑accuracy tools) also indicates Likely Benign; Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, predicts pathogenic. Overall, the majority of evidence points to a benign effect, and this conclusion does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | C2 | 0.268042 | Structured | 0.428564 | Uncertain | 0.956 | 0.369 | 0.000 | -3.483 | Likely Benign | 0.107 | Likely Benign | Likely Benign | 2.18 | Destabilizing | 0.3 | 2.22 | Destabilizing | 2.20 | Destabilizing | 0.17 | Likely Benign | 0.227 | Likely Benign | 0.19 | Neutral | 0.898 | Possibly Damaging | 0.472 | Possibly Damaging | 0.80 | Pathogenic | 0.80 | Tolerated | 0.1277 | 0.2656 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.1403T>G | M468R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M468R is not reported in ClinVar and is absent from gnomAD. All evaluated in‑silico predictors classify the substitution as pathogenic: REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool predicts a benign effect. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized indicates pathogenicity; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) reports “Likely Pathogenic”; and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, also predicts pathogenicity. Consequently, the variant is most likely pathogenic based on the consensus of predictive tools, and this assessment does not contradict any ClinVar status (none available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.284882 | Structured | 0.339253 | Uncertain | 0.932 | 0.257 | 0.000 | -16.180 | Likely Pathogenic | 0.984 | Likely Pathogenic | Likely Pathogenic | 2.66 | Destabilizing | 0.2 | 2.23 | Destabilizing | 2.45 | Destabilizing | 2.43 | Destabilizing | 0.837 | Likely Pathogenic | -4.64 | Deleterious | 0.939 | Possibly Damaging | 0.943 | Probably Damaging | -1.34 | Pathogenic | 0.00 | Affected | 0.1717 | 0.0637 | 0 | -1 | -6.4 | 24.99 | |||||||||||||||||||||||||||||
| c.1418T>A | V473E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V473E is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools largely agree on a deleterious effect: all evaluated algorithms except FATHMM predict pathogenicity, while FATHMM alone predicts benign. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized classifies the variant as pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates a likely pathogenic outcome; and Foldetta, which integrates FoldX‑MD and Rosetta stability predictions, also reports a pathogenic effect. No predictions or stability results are missing or inconclusive. Based on the consensus of the majority of tools and the high‑accuracy methods, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.191378 | Structured | 0.362529 | Uncertain | 0.884 | 0.239 | 0.000 | -15.296 | Likely Pathogenic | 0.992 | Likely Pathogenic | Likely Pathogenic | 3.20 | Destabilizing | 0.2 | 2.24 | Destabilizing | 2.72 | Destabilizing | 2.22 | Destabilizing | 0.664 | Likely Pathogenic | -5.92 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.13 | Benign | 0.00 | Affected | 0.0903 | 0.1396 | -2 | -2 | -7.7 | 29.98 | |||||||||||||||||||||||||||||
| c.658T>C | F220L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F220L is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that classify the variant as benign include polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. All other evaluated predictors—SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—indicate a pathogenic effect. The SGM‑Consensus result is a majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN (3 pathogenic, 1 benign), thus supporting a pathogenic classification. High‑accuracy assessments further corroborate this: AlphaMissense‑Optimized predicts pathogenic, the SGM‑Consensus (majority vote) predicts pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. Consequently, the variant is most likely pathogenic based on the collective predictions, and this assessment does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.219301 | Structured | 0.429422 | Uncertain | 0.898 | 0.295 | 0.000 | -9.601 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 2.08 | Destabilizing | 0.1 | 2.24 | Destabilizing | 2.16 | Destabilizing | 1.33 | Destabilizing | 0.850 | Likely Pathogenic | -4.95 | Deleterious | 0.003 | Benign | 0.005 | Benign | 4.24 | Benign | 0.02 | Affected | 0.2589 | 0.4108 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.660T>A | F220L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F220L is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that classify the variant as benign include polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. All other evaluated predictors—SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—indicate a pathogenic effect. The SGM‑Consensus result is a majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN (3 pathogenic, 1 benign), thus supporting a pathogenic classification. High‑accuracy assessments further corroborate this: AlphaMissense‑Optimized predicts pathogenic, the SGM‑Consensus (majority vote) predicts pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. Consequently, the variant is most likely pathogenic based on the collective predictions, and this assessment does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.219301 | Structured | 0.429422 | Uncertain | 0.898 | 0.295 | 0.000 | -9.601 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 2.08 | Destabilizing | 0.1 | 2.24 | Destabilizing | 2.16 | Destabilizing | 1.33 | Destabilizing | 0.752 | Likely Pathogenic | -4.95 | Deleterious | 0.003 | Benign | 0.005 | Benign | 4.24 | Benign | 0.02 | Affected | 0.2589 | 0.4108 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.660T>G | F220L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F220L is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that classify the variant as benign include polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. All other evaluated predictors—SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—indicate a pathogenic effect. The SGM‑Consensus result is a majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN (3 pathogenic, 1 benign), thus supporting a pathogenic classification. High‑accuracy assessments further corroborate this: AlphaMissense‑Optimized predicts pathogenic, the SGM‑Consensus (majority vote) predicts pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenic. Consequently, the variant is most likely pathogenic based on the collective predictions, and this assessment does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.219301 | Structured | 0.429422 | Uncertain | 0.898 | 0.295 | 0.000 | -9.601 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 2.08 | Destabilizing | 0.1 | 2.24 | Destabilizing | 2.16 | Destabilizing | 1.33 | Destabilizing | 0.752 | Likely Pathogenic | -4.95 | Deleterious | 0.003 | Benign | 0.005 | Benign | 4.24 | Benign | 0.02 | Affected | 0.2589 | 0.4108 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1294T>G | C432G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant C432G has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include only FATHMM. Those that predict a pathogenic effect comprise SGM‑Consensus, REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. Predictions that are inconclusive or uncertain are FoldX, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. Overall, the majority of evidence points to a pathogenic impact. This conclusion is consistent with the absence of a ClinVar classification; there is no contradiction with ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.111485 | Structured | 0.362533 | Uncertain | 0.960 | 0.285 | 0.000 | -10.247 | Likely Pathogenic | 0.831 | Likely Pathogenic | Ambiguous | 1.37 | Ambiguous | 0.0 | 2.25 | Destabilizing | 1.81 | Ambiguous | 1.60 | Destabilizing | 0.631 | Likely Pathogenic | -11.46 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 3.45 | Benign | 0.03 | Affected | 0.2796 | 0.2130 | -3 | -3 | -2.9 | -46.09 | |||||||||||||||||||||||||||||
| c.2065C>G | L689V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L689V is not reported in ClinVar and is absent from gnomAD. Prediction tools that indicate a benign effect include REVEL and FATHMM, whereas the majority of other in silico predictors (FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) and the SGM Consensus (Likely Pathogenic) predict a pathogenic impact. The high‑accuracy methods give the following results: AlphaMissense‑Optimized is uncertain; SGM Consensus is Likely Pathogenic; Foldetta predicts a pathogenic effect. Taken together, the preponderance of evidence points to a pathogenic effect for L689V. This conclusion is not contradicted by ClinVar status, which currently has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.042364 | Structured | 0.227227 | Uncertain | 0.963 | 0.248 | 0.000 | -11.387 | Likely Pathogenic | 0.862 | Likely Pathogenic | Ambiguous | 2.98 | Destabilizing | 0.1 | 2.25 | Destabilizing | 2.62 | Destabilizing | 1.32 | Destabilizing | 0.234 | Likely Benign | -2.97 | Deleterious | 0.926 | Possibly Damaging | 0.481 | Possibly Damaging | 3.27 | Benign | 0.00 | Affected | 0.1393 | 0.3189 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
| c.1873C>T | L625F 2D  AIThe SynGAP1 missense variant L625F is not reported in ClinVar and has no entry in gnomAD. Prediction tools that agree on a benign effect are REVEL and FATHMM. Tools that predict a pathogenic effect include SGM‑Consensus (Likely Pathogenic), Rosetta, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Predictions that are uncertain or inconclusive are FoldX, premPS, and Foldetta. High‑accuracy methods give the following results: AlphaMissense‑Optimized predicts Pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also predicts Likely Pathogenic; Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is Uncertain. Overall, the majority of evidence points to a pathogenic effect. This conclusion is not contradicted by ClinVar status, as no ClinVar classification exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.229226 | Structured | 0.045896 | Uncertain | 0.966 | 0.215 | 0.000 | -12.989 | Likely Pathogenic | 0.982 | Likely Pathogenic | Likely Pathogenic | 1.70 | Ambiguous | 1.3 | 2.26 | Destabilizing | 1.98 | Ambiguous | 0.74 | Ambiguous | 0.479 | Likely Benign | -3.82 | Deleterious | 1.000 | Probably Damaging | 0.997 | Probably Damaging | 3.01 | Benign | 0.01 | Affected | 0.0586 | 0.2998 | 2 | 0 | -1.0 | 34.02 | |||||||||||||||||||||||||||||
| c.1885G>C | V629L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V629L has no ClinVar entry and is not reported in gnomAD. Prediction tools that indicate a benign effect include REVEL, FoldX, and FATHMM, whereas the majority of algorithms predict a pathogenic outcome: Rosetta, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Two tools report uncertainty: Foldetta and premPS. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as likely pathogenic. High‑accuracy assessments further support this: AlphaMissense‑Optimized predicts pathogenicity, the SGM‑Consensus also indicates likely pathogenic, while Foldetta remains uncertain. Based on the collective evidence, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.040537 | Structured | 0.034796 | Uncertain | 0.970 | 0.236 | 0.000 | -12.785 | Likely Pathogenic | 0.972 | Likely Pathogenic | Likely Pathogenic | 0.06 | Likely Benign | 0.2 | 2.26 | Destabilizing | 1.16 | Ambiguous | 0.54 | Ambiguous | 0.391 | Likely Benign | -2.79 | Deleterious | 0.975 | Probably Damaging | 0.958 | Probably Damaging | 3.22 | Benign | 0.02 | Affected | 0.0805 | 0.3336 | 2 | 1 | -0.4 | 14.03 | |||||||||||||||||||||||||||||
| c.808G>A | E270K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E270K is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include only FoldX, whereas the majority of tools predict a pathogenic impact: SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Two tools give uncertain results: Foldetta (a combined FoldX‑MD/Rosetta stability assessment) and premPS. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, SGM‑Consensus is likely pathogenic, and Foldetta remains uncertain. Based on the overall consensus of the available predictions, the variant is most likely pathogenic, which is consistent with the lack of ClinVar reporting and gnomAD absence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.144935 | Structured | 0.382573 | Uncertain | 0.938 | 0.231 | 0.125 | -14.466 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | -0.06 | Likely Benign | 0.2 | 2.26 | Destabilizing | 1.10 | Ambiguous | 0.71 | Ambiguous | 0.530 | Likely Pathogenic | -3.68 | Deleterious | 0.999 | Probably Damaging | 0.995 | Probably Damaging | 1.68 | Pathogenic | 0.01 | Affected | 0.2296 | 0.4251 | 0 | 1 | -0.4 | -0.94 | |||||||||||||||||||||||||||||
| c.1222A>C | T408P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant T408P is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict a pathogenic effect comprise Rosetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and ESM1b. FoldX, premPS, and Foldetta are inconclusive. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is a tie and thus unavailable; Foldetta remains uncertain. Overall, the majority of available predictions (six pathogenic vs. four benign) indicate that the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.161087 | Structured | 0.370935 | Uncertain | 0.907 | 0.239 | 0.000 | -10.384 | Likely Pathogenic | 0.230 | Likely Benign | Likely Benign | 1.08 | Ambiguous | 0.3 | 2.27 | Destabilizing | 1.68 | Ambiguous | 0.73 | Ambiguous | 0.323 | Likely Benign | -4.19 | Deleterious | 0.998 | Probably Damaging | 0.963 | Probably Damaging | 4.07 | Benign | 0.05 | Affected | 0.1985 | 0.5779 | 0 | -1 | -0.9 | -3.99 | ||||||||||||||||||||||||||||||
| c.1889T>C | I630T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I630T has no ClinVar entry and is not reported in gnomAD. Prediction tools that agree on a benign effect include PROVEAN, SIFT, and AlphaMissense‑Optimized. Those that predict a pathogenic impact are REVEL, FoldX, Rosetta, Foldetta, premPS, polyPhen‑2 (HumDiv and HumVar), FATHMM, and AlphaMissense‑Default; ESM1b remains uncertain. High‑accuracy assessments further support a deleterious outcome: AlphaMissense‑Optimized indicates benign, whereas the SGM Consensus (derived from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) yields a pathogenic majority, and Foldetta also predicts pathogenic. No prediction is missing or inconclusive. Overall, the preponderance of evidence points to a pathogenic effect for I630T, and this conclusion does not conflict with the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.040537 | Structured | 0.036106 | Uncertain | 0.966 | 0.236 | 0.000 | -7.780 | In-Between | 0.754 | Likely Pathogenic | Likely Benign | 2.77 | Destabilizing | 0.1 | 2.27 | Destabilizing | 2.52 | Destabilizing | 1.94 | Destabilizing | 0.734 | Likely Pathogenic | -2.15 | Neutral | 0.997 | Probably Damaging | 0.961 | Probably Damaging | -1.46 | Pathogenic | 0.35 | Tolerated | 0.0985 | 0.0640 | 0 | -1 | -5.2 | -12.05 | ||||||||||||||||||||||||||||||
| c.1901C>G | A634G 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant A634G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a split: benign calls come from REVEL, SIFT, FATHMM, and AlphaMissense‑Optimized, while pathogenic calls are made by Rosetta, premPS, PROVEAN, both polyPhen‑2 versions, ESM1b, and AlphaMissense‑Default. FoldX and Foldetta give uncertain results. High‑accuracy assessments indicate AlphaMissense‑Optimized predicts benign, whereas the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts pathogenic. Foldetta remains uncertain. Overall, the majority of tools lean toward pathogenicity, and the high‑accuracy consensus also supports a pathogenic interpretation. Therefore, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar annotation because none exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.085092 | Structured | 0.052058 | Uncertain | 0.932 | 0.242 | 0.000 | -10.685 | Likely Pathogenic | 0.613 | Likely Pathogenic | Likely Benign | 1.63 | Ambiguous | 0.1 | 2.27 | Destabilizing | 1.95 | Ambiguous | 1.09 | Destabilizing | 0.418 | Likely Benign | -3.98 | Deleterious | 0.997 | Probably Damaging | 0.990 | Probably Damaging | 2.69 | Benign | 0.15 | Tolerated | 0.2182 | 0.3187 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.880A>G | T294A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 T294A missense variant is not reported in ClinVar (ClinVar status: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a pathogenic effect include SGM‑Consensus (Likely Pathogenic), REVEL, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized; the only tool with an uncertain call is FoldX. High‑accuracy methods give consistent results: AlphaMissense‑Optimized predicts Pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts Likely Pathogenic, and Foldetta predicts Pathogenic. Based on the overwhelming agreement among these predictions, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.328603 | Structured | 0.316932 | Uncertain | 0.919 | 0.267 | 0.125 | -12.371 | Likely Pathogenic | 0.971 | Likely Pathogenic | Likely Pathogenic | 1.87 | Ambiguous | 0.1 | 2.27 | Destabilizing | 2.07 | Destabilizing | 1.05 | Destabilizing | 0.719 | Likely Pathogenic | -4.60 | Deleterious | 0.997 | Probably Damaging | 0.992 | Probably Damaging | -0.18 | Pathogenic | 0.03 | Affected | 0.3687 | 0.3494 | 1 | 0 | 2.5 | -30.03 | |||||||||||||||||||||||||||||
| c.1045C>T | P349S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 P349S missense variant is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, SIFT, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict a pathogenic impact are Rosetta, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. Predictions that are inconclusive or uncertain are FoldX, ESM1b, and premPS. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as pathogenic. Overall, the majority of tools, including the high‑accuracy methods, predict a pathogenic effect. Thus, the variant is most likely pathogenic, which does not contradict its current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.167087 | Structured | 0.348607 | Uncertain | 0.947 | 0.396 | 0.000 | Uncertain | 1 | -7.654 | In-Between | 0.217 | Likely Benign | Likely Benign | 1.92 | Ambiguous | 0.1 | 2.28 | Destabilizing | 2.10 | Destabilizing | 0.87 | Ambiguous | 0.277 | Likely Benign | -6.13 | Deleterious | 1.000 | Probably Damaging | 0.996 | Probably Damaging | 1.66 | Pathogenic | 0.06 | Tolerated | 3.37 | 25 | 0.3771 | 0.5756 | 1 | -1 | 0.8 | -10.04 | 194.9 | -18.1 | -0.1 | 0.0 | 0.2 | 0.1 | X | X | Potentially Pathogenic | The cyclic pyrrolidine side chain of Pro349, located at the end of an anti-parallel β sheet strand (res. Gly341-Pro349), allows the strand to end and make a tight turn before a short α helical section within a loop connecting to another β strand (res. Thr359-Pro364). In the variant simulations, the hydroxyl group of Ser349 forms a hydrogen bond with the backbone amide group of Ala351 in the short helical section. Conversely, the backbone amide group of Ser349 (absent in proline) does not form any intra-protein hydrogen bonds. However, the β strand end connects to the α helical section in a more stable and consistent manner compared to the WT. Although the residue swap does not cause major adverse effects on the protein structure in the simulations, it is possible that the tight turn at the β strand end could not be created during folding without the presence of proline. | ||||||||||||||||
| c.1349C>G | A450G 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant A450G is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. The majority of other in silico predictors (SGM‑Consensus, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) classify the variant as pathogenic; FoldX is inconclusive. High‑accuracy assessments further support a pathogenic interpretation: AlphaMissense‑Optimized predicts benign, but the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) and Foldetta (combining FoldX‑MD and Rosetta) both predict pathogenic. Overall, the preponderance of evidence indicates that A450G is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.321458 | Structured | 0.306281 | Uncertain | 0.963 | 0.234 | 0.000 | -11.090 | Likely Pathogenic | 0.695 | Likely Pathogenic | Likely Benign | 1.79 | Ambiguous | 0.0 | 2.29 | Destabilizing | 2.04 | Destabilizing | 1.23 | Destabilizing | 0.355 | Likely Benign | -3.82 | Deleterious | 0.998 | Probably Damaging | 0.980 | Probably Damaging | 3.40 | Benign | 0.04 | Affected | 0.1670 | 0.3078 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.1394T>A | L465H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L465H is not reported in ClinVar and is absent from gnomAD. Functional prediction tools uniformly indicate a deleterious effect. Benign predictions: none. Pathogenic predictions: SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, predicts pathogenic. Based on the consensus of all available predictions, the variant is most likely pathogenic, and this conclusion is not contradicted by ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.346032 | Structured | 0.319240 | Uncertain | 0.956 | 0.202 | 0.000 | -16.751 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 3.15 | Destabilizing | 0.4 | 2.29 | Destabilizing | 2.72 | Destabilizing | 2.54 | Destabilizing | 0.764 | Likely Pathogenic | -6.96 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 2.29 | Pathogenic | 0.00 | Affected | 0.1289 | 0.1089 | -2 | -3 | -7.0 | 23.98 | |||||||||||||||||||||||||||||
| c.1581C>A | D527E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D527E missense variant is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a strong bias toward pathogenicity: REVEL, Rosetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default all predict a deleterious effect, whereas only FoldX and premPS predict a benign outcome. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as Likely Pathogenic. High‑accuracy assessments are mixed: AlphaMissense‑Optimized is inconclusive, Foldetta (combining FoldX‑MD and Rosetta outputs) is uncertain, and the SGM Consensus remains Likely Pathogenic. Overall, the preponderance of evidence points to a pathogenic impact, and this assessment does not conflict with the absence of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.139895 | Structured | 0.021908 | Uncertain | 0.913 | 0.408 | 0.000 | -11.125 | Likely Pathogenic | 0.884 | Likely Pathogenic | Ambiguous | 0.36 | Likely Benign | 0.8 | 2.29 | Destabilizing | 1.33 | Ambiguous | 0.50 | Likely Benign | 0.740 | Likely Pathogenic | -3.74 | Deleterious | 0.929 | Possibly Damaging | 0.938 | Probably Damaging | -2.31 | Pathogenic | 0.02 | Affected | 0.1103 | 0.3428 | 3 | 2 | 0.0 | 14.03 | |||||||||||||||||||||||||||||
| c.1581C>G | D527E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D527E missense variant is not reported in ClinVar and is absent from gnomAD. Functional prediction tools show a strong bias toward pathogenicity: REVEL, Rosetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default all predict a deleterious effect, whereas only FoldX and premPS predict a benign outcome. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as Likely Pathogenic. High‑accuracy assessments are mixed: AlphaMissense‑Optimized is inconclusive, Foldetta (combining FoldX‑MD and Rosetta outputs) is uncertain, and the SGM Consensus remains Likely Pathogenic. Overall, the preponderance of evidence points to a pathogenic impact, and this assessment does not conflict with the absence of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.139895 | Structured | 0.021908 | Uncertain | 0.913 | 0.408 | 0.000 | -11.125 | Likely Pathogenic | 0.884 | Likely Pathogenic | Ambiguous | 0.36 | Likely Benign | 0.8 | 2.29 | Destabilizing | 1.33 | Ambiguous | 0.50 | Likely Benign | 0.740 | Likely Pathogenic | -3.74 | Deleterious | 0.929 | Possibly Damaging | 0.938 | Probably Damaging | -2.31 | Pathogenic | 0.02 | Affected | 0.1103 | 0.3428 | 3 | 2 | 0.0 | 14.03 | |||||||||||||||||||||||||||||
| c.2108T>A | L703H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L703H is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: benign predictions come from REVEL and FATHMM, whereas pathogenic predictions are made by FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as Likely Pathogenic. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized predicts Pathogenic; the SGM‑Consensus itself is Likely Pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, also predicts Pathogenic. Taken together, the overwhelming majority of evidence indicates that the variant is most likely pathogenic, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.144935 | Structured | 0.388282 | Uncertain | 0.929 | 0.353 | 0.000 | -12.886 | Likely Pathogenic | 0.957 | Likely Pathogenic | Likely Pathogenic | 2.52 | Destabilizing | 0.0 | 2.29 | Destabilizing | 2.41 | Destabilizing | 1.75 | Destabilizing | 0.420 | Likely Benign | -5.71 | Deleterious | 1.000 | Probably Damaging | 0.993 | Probably Damaging | 3.09 | Benign | 0.00 | Affected | 0.1038 | 0.0288 | -2 | -3 | -7.0 | 23.98 | |||||||||||||||||||||||||||||
| c.1934T>C | F645S 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant F645S is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: benign calls come from REVEL, polyPhen‑2 HumVar, and FATHMM, whereas pathogenic calls are made by FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, while both the SGM Consensus and Foldetta predict pathogenicity. No evidence from ClinVar contradicts these findings. Overall, the preponderance of predictions supports a pathogenic classification for F645S. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.046336 | Structured | 0.276445 | Uncertain | 0.921 | 0.325 | 0.000 | -9.748 | Likely Pathogenic | 0.947 | Likely Pathogenic | Ambiguous | 2.49 | Destabilizing | 0.2 | 2.30 | Destabilizing | 2.40 | Destabilizing | 1.57 | Destabilizing | 0.326 | Likely Benign | -5.34 | Deleterious | 0.755 | Possibly Damaging | 0.112 | Benign | 3.37 | Benign | 0.03 | Affected | 0.3899 | 0.0547 | -3 | -2 | -3.6 | -60.10 | |||||||||||||||||||||||||||||
| c.691T>G | F231V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F231V is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect are polyPhen‑2 HumVar and FATHMM. All other evaluated algorithms—REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—classify the change as pathogenic. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized predicts pathogenicity; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) reports a likely pathogenic status; and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, also indicates pathogenicity. Taken together, the evidence overwhelmingly points to a pathogenic effect for F231V, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.366687 | Structured | 0.306467 | Uncertain | 0.895 | 0.300 | 0.000 | -13.201 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 2.16 | Destabilizing | 0.3 | 2.30 | Destabilizing | 2.23 | Destabilizing | 1.13 | Destabilizing | 0.910 | Likely Pathogenic | -5.90 | Deleterious | 0.759 | Possibly Damaging | 0.328 | Benign | 5.72 | Benign | 0.00 | Affected | 0.2212 | 0.3030 | -1 | -1 | 1.4 | -48.04 | |||||||||||||||||||||||||||||
| c.1802C>G | A601G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 A601G missense variant is listed in gnomAD (ID 6‑33440854‑C‑G) but has no ClinVar entry. Prediction tools that agree on a benign effect are FATHMM and AlphaMissense‑Optimized; those that agree on a pathogenic effect are REVEL, FoldX, Rosetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and ESM1b. The high‑accuracy consensus methods give a pathogenic verdict: the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is also pathogenic. AlphaMissense‑Default and premPS are uncertain, and no evidence is available from other folding‑stability tools. Overall, the preponderance of evidence points to a pathogenic impact for A601G, and this assessment does not contradict the absence of a ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.008895 | Structured | 0.174517 | Uncertain | 0.955 | 0.156 | 0.000 | 6-33440854-C-G | 1 | 6.19e-7 | -11.772 | Likely Pathogenic | 0.543 | Ambiguous | Likely Benign | 2.03 | Destabilizing | 0.0 | 2.31 | Destabilizing | 2.17 | Destabilizing | 0.94 | Ambiguous | 0.511 | Likely Pathogenic | -3.98 | Deleterious | 1.000 | Probably Damaging | 0.997 | Probably Damaging | 2.55 | Benign | 0.01 | Affected | 3.37 | 35 | 0.2337 | 0.4370 | 0 | 1 | -2.2 | -14.03 | |||||||||||||||||||||||||
| c.990C>A | D330E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D330E missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are Rosetta and FATHMM. The remaining tools—FoldX, AlphaMissense‑Default, and Foldetta—return uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as benign, and Foldetta as uncertain. Overall, the majority of evidence points to a benign impact. This conclusion is not contradicted by any ClinVar annotation, as no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.380708 | Structured | 0.360008 | Uncertain | 0.805 | 0.488 | 0.250 | -3.427 | Likely Benign | 0.407 | Ambiguous | Likely Benign | 0.98 | Ambiguous | 0.2 | 2.31 | Destabilizing | 1.65 | Ambiguous | 0.39 | Likely Benign | 0.073 | Likely Benign | -1.38 | Neutral | 0.122 | Benign | 0.030 | Benign | 0.97 | Pathogenic | 0.67 | Tolerated | 0.1464 | 0.4416 | 3 | 2 | 0.0 | 14.03 | ||||||||||||||||||||||||||||||
| c.990C>G | D330E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 D330E missense variant is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are Rosetta and FATHMM. The remaining tools—FoldX, AlphaMissense‑Default, and Foldetta—return uncertain or inconclusive results. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as benign, and Foldetta as uncertain. Overall, the majority of evidence points to a benign impact. This conclusion is not contradicted by any ClinVar annotation, as no ClinVar entry exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.380708 | Structured | 0.360008 | Uncertain | 0.805 | 0.488 | 0.250 | -3.427 | Likely Benign | 0.407 | Ambiguous | Likely Benign | 0.98 | Ambiguous | 0.2 | 2.31 | Destabilizing | 1.65 | Ambiguous | 0.39 | Likely Benign | 0.075 | Likely Benign | -1.38 | Neutral | 0.122 | Benign | 0.030 | Benign | 0.97 | Pathogenic | 0.67 | Tolerated | 0.1464 | 0.4416 | 3 | 2 | 0.0 | 14.03 | ||||||||||||||||||||||||||||||
| c.1362C>G | I454M 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I454M is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include REVEL and FATHMM, whereas a majority of tools (SGM‑Consensus, Rosetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) predict a pathogenic impact. Tools with inconclusive results are FoldX, Foldetta, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta as uncertain. Overall, the preponderance of evidence points to a pathogenic effect for I454M. This conclusion is not contradicted by ClinVar, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.254060 | Structured | 0.312811 | Uncertain | 0.965 | 0.182 | 0.000 | -8.437 | Likely Pathogenic | 0.871 | Likely Pathogenic | Ambiguous | 1.06 | Ambiguous | 0.1 | 2.32 | Destabilizing | 1.69 | Ambiguous | 1.08 | Destabilizing | 0.292 | Likely Benign | -2.86 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.27 | Benign | 0.01 | Affected | 0.0566 | 0.2524 | 2 | 1 | -2.6 | 18.03 | |||||||||||||||||||||||||||||
| c.1778T>A | L593H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L593H is listed in ClinVar with an uncertain significance and is not present in gnomAD. In silico predictors that classify the variant as benign include only FATHMM. All other evaluated tools—REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—predict a pathogenic effect. High‑accuracy methods further support pathogenicity: AlphaMissense‑Optimized is pathogenic; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, is pathogenic. No prediction or stability result is missing or inconclusive. Overall, the variant is most likely pathogenic based on the consensus of predictions, and this assessment does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.009728 | Structured | 0.110534 | Uncertain | 0.941 | 0.151 | 0.000 | Uncertain | 1 | -16.504 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 2.52 | Destabilizing | 0.2 | 2.32 | Destabilizing | 2.42 | Destabilizing | 2.75 | Destabilizing | 0.812 | Likely Pathogenic | -6.77 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 2.77 | Benign | 0.00 | Affected | 3.37 | 35 | 0.1101 | 0.0541 | -2 | -3 | -7.0 | 23.98 | 222.0 | 20.7 | 0.0 | 0.0 | 0.2 | 0.0 | X | X | Potentially Pathogenic | The iso-propyl side chain of Leu593, located in an α helix (res. Glu582-Met603), packs favourably with multiple hydrophobic residues in the inter-helix space (e.g., Leu598, Ile589, Phe594, Phe561).In the variant simulations, His593 retains a similar packing arrangement via its aromatic imidazole ring. However, the polar nitrogen atoms introduce hydrogen bond donors and acceptors into the previously hydrophobic space. The epsilon protonated nitrogen of His593 forms a stable hydrogen bond with the phenol group of the Tyr505 side chain in an α helix (res. Gln503-Tyr518).While the residue swap could affect the tertiary assembly and the underlying protein folding process, it is difficult to determine if the mutation would be tolerated based solely on the variant simulations. | |||||||||||||||
| c.2018T>A | L673Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L673Q is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. Tools that agree on a pathogenic effect include Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and ESM1b. FoldX and Foldetta, as well as AlphaMissense‑Default, are inconclusive and are treated as unavailable evidence. High‑accuracy assessments show AlphaMissense‑Optimized predicting benign, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) predicts pathogenic. Foldetta remains uncertain. Overall, the majority of available predictions (seven pathogenic vs. three benign) indicate that the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar annotation because no ClinVar claim exists for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.060549 | Structured | 0.104692 | Uncertain | 0.545 | 0.369 | 0.000 | -11.963 | Likely Pathogenic | 0.445 | Ambiguous | Likely Benign | 1.56 | Ambiguous | 0.2 | 2.32 | Destabilizing | 1.94 | Ambiguous | 1.02 | Destabilizing | 0.190 | Likely Benign | -3.40 | Deleterious | 1.000 | Probably Damaging | 0.968 | Probably Damaging | 3.32 | Benign | 0.03 | Affected | 0.1045 | 0.1172 | -2 | -2 | -7.3 | 14.97 | ||||||||||||||||||||||||||||||
| c.635C>A | S212Y 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S212Y is not reported in ClinVar (ClinVar status: not listed) and is absent from gnomAD (no gnomAD ID). Prediction tools that agree on a benign effect include FoldX, FATHMM, and premPS. In contrast, a majority of tools predict a pathogenic impact: REVEL, SGM‑Consensus, AlphaMissense‑Default, AlphaMissense‑Optimized, ESM1b, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT. Foldetta, a protein‑folding stability method that integrates FoldX‑MD and Rosetta outputs, is inconclusive for this variant. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized classifies the variant as pathogenic; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) reports it as likely pathogenic; Foldetta remains uncertain. Overall, the preponderance of evidence from multiple in silico predictors indicates that S212Y is most likely pathogenic, and this conclusion does not contradict any existing ClinVar annotation, as none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.158265 | Structured | 0.381517 | Uncertain | 0.846 | 0.278 | 0.125 | -14.812 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.43 | Likely Benign | 0.5 | 2.32 | Destabilizing | 1.38 | Ambiguous | 0.22 | Likely Benign | 0.819 | Likely Pathogenic | -5.15 | Deleterious | 0.995 | Probably Damaging | 0.986 | Probably Damaging | 5.76 | Benign | 0.00 | Affected | 0.0555 | 0.6120 | -3 | -2 | -0.5 | 76.10 | |||||||||||||||||||||||||||||
| c.1028T>C | V343A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V343A is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are REVEL and AlphaMissense‑Optimized, whereas the remaining tools—SGM‑Consensus, FoldX (uncertain), Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default—consistently predict a pathogenic impact. High‑accuracy assessments further support this: AlphaMissense‑Optimized classifies the variant as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates a likely pathogenic effect, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts pathogenicity. Overall, the preponderance of evidence points to a pathogenic effect for V343A, and this conclusion does not contradict the ClinVar status, which currently contains no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.291804 | Structured | 0.383911 | Uncertain | 0.882 | 0.497 | 0.250 | -8.088 | Likely Pathogenic | 0.588 | Likely Pathogenic | Likely Benign | 1.66 | Ambiguous | 0.1 | 2.33 | Destabilizing | 2.00 | Destabilizing | 1.69 | Destabilizing | 0.218 | Likely Benign | -3.15 | Deleterious | 0.826 | Possibly Damaging | 0.551 | Possibly Damaging | 1.63 | Pathogenic | 0.01 | Affected | 0.3137 | 0.3301 | 0 | 0 | -2.4 | -28.05 | |||||||||||||||||||||||||||||
| c.1303T>G | L435V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L435V has no ClinVar assertion and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, FATHMM, and AlphaMissense‑Optimized, while pathogenic predictions arise from FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), and AlphaMissense‑Default. ESM1b is uncertain. High‑accuracy assessments further support a pathogenic bias: AlphaMissense‑Optimized predicts benign, but the SGM Consensus—derived from a majority vote of AlphaMissense‑Default (pathogenic), ESM1b (uncertain), FATHMM (benign), and PROVEAN (pathogenic)—favors pathogenicity. Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, also predicts pathogenic. With the preponderance of pathogenic calls from both general and high‑accuracy tools, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar claim exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | 0.229226 | Structured | 0.333584 | Uncertain | 0.954 | 0.292 | 0.000 | -7.134 | In-Between | 0.571 | Likely Pathogenic | Likely Benign | 2.97 | Destabilizing | 0.1 | 2.33 | Destabilizing | 2.65 | Destabilizing | 1.36 | Destabilizing | 0.217 | Likely Benign | -2.80 | Deleterious | 0.995 | Probably Damaging | 0.970 | Probably Damaging | 3.23 | Benign | 0.06 | Tolerated | 0.1216 | 0.2628 | 2 | 1 | 0.4 | -14.03 | ||||||||||||||||||||||||||||||
| c.1463C>A | T488K 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant T488K is not reported in ClinVar and is absent from gnomAD. Prediction tools largely agree on a deleterious effect: FATHMM is the sole benign predictor, while the remaining eleven tools (SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) all classify the variant as pathogenic. The high‑accuracy methods—AlphaMissense‑Optimized, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN), and Foldetta (combining FoldX‑MD and Rosetta outputs)—all predict pathogenicity. No prediction or folding‑stability result is missing or inconclusive. Based on the consensus of these tools, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.206376 | Structured | 0.332663 | Uncertain | 0.928 | 0.233 | 0.125 | -14.391 | Likely Pathogenic | 0.994 | Likely Pathogenic | Likely Pathogenic | 2.09 | Destabilizing | 0.2 | 2.33 | Destabilizing | 2.21 | Destabilizing | 0.90 | Ambiguous | 0.692 | Likely Pathogenic | -5.64 | Deleterious | 1.000 | Probably Damaging | 0.996 | Probably Damaging | 3.24 | Benign | 0.00 | Affected | 0.0871 | 0.2104 | 0 | -1 | -3.2 | 27.07 | |||||||||||||||||||||||||||||
| c.1526C>G | A509G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A509G is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Prediction tools that clearly indicate benign effect include only AlphaMissense‑Optimized. All other evaluated tools that provide a definitive call predict pathogenicity: SGM‑Consensus, REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and FATHMM. Tools with inconclusive results (AlphaMissense‑Default, FoldX, and Foldetta) are treated as unavailable and do not influence the overall assessment. High‑accuracy methods give the following: AlphaMissense‑Optimized predicts benign; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic; Foldetta, combining FoldX‑MD (uncertain) and Rosetta (pathogenic), is uncertain. Overall, the majority of definitive predictions support a pathogenic effect. Thus, the variant is most likely pathogenic, and this conclusion does not contradict the ClinVar status, which currently has no entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.025762 | Structured | 0.250110 | Uncertain | 0.923 | 0.256 | 0.000 | -11.873 | Likely Pathogenic | 0.541 | Ambiguous | Likely Benign | 1.36 | Ambiguous | 0.2 | 2.33 | Destabilizing | 1.85 | Ambiguous | 1.14 | Destabilizing | 0.804 | Likely Pathogenic | -3.57 | Deleterious | 0.911 | Possibly Damaging | 0.706 | Possibly Damaging | -1.39 | Pathogenic | 0.00 | Affected | 0.2193 | 0.4213 | 1 | 0 | -2.2 | -14.03 | |||||||||||||||||||||||||||||
| c.1709C>A | A570D 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A570D is not reported in ClinVar (ClinVar status: not listed) and is absent from gnomAD (gnomAD ID: none). Prediction tools that assess pathogenicity unanimously classify the variant as deleterious: SGM‑Consensus (Likely Pathogenic), REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool predicts a benign effect, so the benign group is empty. High‑accuracy methods reinforce this view: AlphaMissense‑Optimized predicts Pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates Likely Pathogenic; and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts Pathogenic. Based on the collective evidence, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because no ClinVar entry exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.046336 | Structured | 0.054494 | Uncertain | 0.932 | 0.263 | 0.000 | -14.117 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 2.47 | Destabilizing | 1.2 | 2.33 | Destabilizing | 2.40 | Destabilizing | 1.36 | Destabilizing | 0.805 | Likely Pathogenic | -5.31 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | -1.28 | Pathogenic | 0.03 | Affected | 0.2006 | 0.2206 | 0 | -2 | -5.3 | 44.01 | |||||||||||||||||||||||||||||
| c.1879G>A | A627T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant A627T is not reported in ClinVar (ClinVar ID: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a benign effect include only FATHMM, whereas the remaining tools—SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default—consistently predict a pathogenic impact. Two tools, premPS and AlphaMissense‑Optimized, return uncertain results. High‑accuracy assessments further support pathogenicity: the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) yields a pathogenic verdict, and Foldetta also predicts a destabilizing, pathogenic effect. AlphaMissense‑Optimized remains uncertain. Overall, the preponderance of evidence indicates that A627T is most likely pathogenic, and this conclusion does not contradict the absence of ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.100716 | Structured | 0.037862 | Uncertain | 0.970 | 0.210 | 0.000 | -8.613 | Likely Pathogenic | 0.937 | Likely Pathogenic | Ambiguous | 2.27 | Destabilizing | 0.3 | 2.33 | Destabilizing | 2.30 | Destabilizing | 0.80 | Ambiguous | 0.505 | Likely Pathogenic | -3.95 | Deleterious | 0.994 | Probably Damaging | 0.807 | Possibly Damaging | 2.56 | Benign | 0.01 | Affected | 0.1069 | 0.5403 | 1 | 0 | -2.5 | 30.03 | |||||||||||||||||||||||||||||
| c.772C>G | R258G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R258G is not reported in ClinVar and is absent from gnomAD. Prediction tools largely agree on a deleterious effect: FATHMM is the sole benign predictor, while the remaining twelve tools (SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) all classify the variant as pathogenic; premPS remains uncertain. High‑accuracy methods reinforce this view: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) yields a pathogenic verdict; and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, also predicts pathogenic. No prediction is missing or inconclusive. Consequently, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status because none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.295083 | Structured | 0.293667 | Uncertain | 0.894 | 0.260 | 0.250 | -14.239 | Likely Pathogenic | 0.980 | Likely Pathogenic | Likely Pathogenic | 2.42 | Destabilizing | 0.5 | 2.33 | Destabilizing | 2.38 | Destabilizing | 0.94 | Ambiguous | 0.788 | Likely Pathogenic | -5.83 | Deleterious | 0.997 | Probably Damaging | 0.987 | Probably Damaging | 5.78 | Benign | 0.04 | Affected | 0.3166 | 0.3447 | -3 | -2 | 4.1 | -99.14 | |||||||||||||||||||||||||||||
| c.1159G>T | G387C 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant G387C is not reported in ClinVar (ClinVar ID None) but is present in gnomAD (ID 6‑33438064‑G‑T). Prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, AlphaMissense‑Default, AlphaMissense‑Optimized, and polyPhen‑2 HumVar. Those that predict a pathogenic effect are FoldX, Rosetta, Foldetta, polyPhen‑2 HumDiv, SIFT, and FATHMM; ESM1b is uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as pathogenic. Overall, the majority of high‑confidence tools lean toward a benign interpretation, and this does not contradict the ClinVar status, which has no pathogenic classification for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.642678 | Disordered | 0.422910 | Uncertain | 0.293 | 0.861 | 0.750 | 6-33438064-G-T | 1 | 6.21e-7 | -7.609 | In-Between | 0.146 | Likely Benign | Likely Benign | 2.88 | Destabilizing | 0.6 | 2.34 | Destabilizing | 2.61 | Destabilizing | -0.03 | Likely Benign | 0.430 | Likely Benign | -0.58 | Neutral | 0.859 | Possibly Damaging | 0.346 | Benign | 1.32 | Pathogenic | 0.01 | Affected | 4.32 | 3 | 0.1589 | 0.4240 | -3 | -3 | 2.9 | 46.09 | |||||||||||||||||||||||||
| c.1439A>G | E480G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 E480G missense variant is not reported in ClinVar and has no gnomAD entry. Consensus from multiple in‑silico predictors indicates a pathogenic effect: SGM‑Consensus, REVEL, Rosetta, Foldetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default all classify it as pathogenic. Predictions that are uncertain—FoldX, premPS, and AlphaMissense‑Optimized—do not provide evidence for benignity. High‑accuracy assessments further support pathogenicity: the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic, Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic, and AlphaMissense‑Optimized remains uncertain. Consequently, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation because no ClinVar record exists. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.216401 | Structured | 0.426867 | Uncertain | 0.798 | 0.250 | 0.000 | -11.651 | Likely Pathogenic | 0.839 | Likely Pathogenic | Ambiguous | 1.83 | Ambiguous | 0.1 | 2.34 | Destabilizing | 2.09 | Destabilizing | 0.65 | Ambiguous | 0.778 | Likely Pathogenic | -5.44 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | -1.32 | Pathogenic | 0.03 | Affected | 0.2806 | 0.6172 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.1661T>C | V554A 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V554A is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect are REVEL and FATHMM. All other evaluated predictors—SGM‑Consensus, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default—consistently predict a pathogenic impact. High‑accuracy assessments further support this: AlphaMissense‑Optimized is uncertain, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts Pathogenic. No prediction or stability result is missing or inconclusive. Based on the overwhelming majority of pathogenic predictions, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status because none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.020522 | Structured | 0.007349 | Uncertain | 0.955 | 0.226 | 0.000 | -9.730 | Likely Pathogenic | 0.870 | Likely Pathogenic | Ambiguous | 2.07 | Destabilizing | 0.1 | 2.34 | Destabilizing | 2.21 | Destabilizing | 2.00 | Destabilizing | 0.419 | Likely Benign | -3.97 | Deleterious | 0.998 | Probably Damaging | 0.981 | Probably Damaging | 3.22 | Benign | 0.02 | Affected | 0.2131 | 0.1633 | 0 | 0 | -2.4 | -28.05 | |||||||||||||||||||||||||||||
| c.760A>G | K254E 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant K254E is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely converge on a deleterious effect: FATHMM is the sole benign predictor, while the remaining 13 tools (REVEL, FoldX, Rosetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and SGM‑Consensus) predict pathogenicity; FoldX and Foldetta are uncertain. High‑accuracy assessments reinforce this trend: AlphaMissense‑Optimized indicates pathogenic, SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) remains uncertain. Overall, the preponderance of evidence supports a pathogenic classification for K254E, and this conclusion does not conflict with ClinVar, which currently has no entry for the variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.513880 | Disordered | 0.207751 | Uncertain | 0.799 | 0.285 | 0.375 | -14.745 | Likely Pathogenic | 0.996 | Likely Pathogenic | Likely Pathogenic | 0.72 | Ambiguous | 0.1 | 2.34 | Destabilizing | 1.53 | Ambiguous | 1.17 | Destabilizing | 0.860 | Likely Pathogenic | -3.27 | Deleterious | 0.970 | Probably Damaging | 0.584 | Possibly Damaging | 5.87 | Benign | 0.05 | Affected | 0.3224 | 0.1352 | 0 | 1 | 0.4 | 0.94 | |||||||||||||||||||||||||||||
| c.1493T>G | M498R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M498R is listed in ClinVar as Pathogenic (ClinVar ID 3907767.0) and is not reported in gnomAD. Prediction tools that agree on a benign effect include only polyPhen‑2 HumVar; all other evaluated algorithms (REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic impact. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, is pathogenic. No predictions or stability results are missing or inconclusive. Based on the overwhelming consensus of pathogenic predictions, the variant is most likely pathogenic, and this conclusion aligns with its ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.092881 | Structured | 0.399612 | Uncertain | 0.932 | 0.158 | 0.000 | Likely Pathogenic | 1 | -8.812 | Likely Pathogenic | 0.988 | Likely Pathogenic | Likely Pathogenic | 3.85 | Destabilizing | 0.2 | 2.35 | Destabilizing | 3.10 | Destabilizing | 1.76 | Destabilizing | 0.869 | Likely Pathogenic | -4.53 | Deleterious | 0.464 | Possibly Damaging | 0.120 | Benign | -1.36 | Pathogenic | 0.00 | Affected | 0.1482 | 0.0757 | 0 | -1 | -6.4 | 24.99 | |||||||||||||||||||||||||||
| c.1652T>G | L551R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L551R is not reported in ClinVar (ClinVar status: None) and is absent from gnomAD (gnomAD ID: None). Prediction tools that agree on a pathogenic effect include REVEL, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized; the only tool with an uncertain outcome is FoldX. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenicity. Based on the concordant predictions from multiple independent algorithms and the absence of benign evidence, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.009977 | Structured | 0.006653 | Uncertain | 0.960 | 0.254 | 0.000 | -15.420 | Likely Pathogenic | 0.992 | Likely Pathogenic | Likely Pathogenic | 1.81 | Ambiguous | 0.2 | 2.35 | Destabilizing | 2.08 | Destabilizing | 2.09 | Destabilizing | 0.949 | Likely Pathogenic | -3.85 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | -1.60 | Pathogenic | 0.01 | Affected | 0.1278 | 0.0530 | -3 | -2 | -8.3 | 43.03 | |||||||||||||||||||||||||||||
| c.626T>A | V209E 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant V209E is not reported in ClinVar and is absent from gnomAD. Prediction tools that indicate a benign effect include REVEL and FATHMM, whereas the remaining 13 tools (FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus) all predict a pathogenic outcome. High‑accuracy methods further support a deleterious impact: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is pathogenic, and Foldetta (combining FoldX‑MD and Rosetta) is pathogenic. No prediction or stability result is missing or inconclusive. Based on the overwhelming majority of pathogenic predictions, V209E is most likely pathogenic, and this assessment does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.247041 | Structured | 0.397624 | Uncertain | 0.874 | 0.331 | 0.125 | -14.366 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 3.53 | Destabilizing | 1.3 | 2.35 | Destabilizing | 2.94 | Destabilizing | 1.52 | Destabilizing | 0.403 | Likely Benign | -4.54 | Deleterious | 0.995 | Probably Damaging | 0.829 | Possibly Damaging | 3.65 | Benign | 0.00 | Affected | 0.0826 | 0.1652 | -2 | -2 | -7.7 | 29.98 | |||||||||||||||||||||||||||||
| c.658T>A | F220I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F220I is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD (no gnomAD ID). Prediction tools that classify the variant as benign include polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM. All other evaluated algorithms—SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—predict a pathogenic effect. The SGM‑Consensus result is a majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, yielding 3 pathogenic versus 1 benign calls, thus supporting a pathogenic classification. High‑accuracy tools specifically report pathogenicity: AlphaMissense‑Optimized is pathogenic; the SGM‑Consensus (majority vote) is pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is pathogenic. Consequently, the variant is most likely pathogenic based on the collective predictions, and this assessment does not contradict any ClinVar status because none is available. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.219301 | Structured | 0.429422 | Uncertain | 0.898 | 0.295 | 0.000 | -12.041 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 3.11 | Destabilizing | 0.1 | 2.35 | Destabilizing | 2.73 | Destabilizing | 1.03 | Destabilizing | 0.921 | Likely Pathogenic | -4.98 | Deleterious | 0.300 | Benign | 0.098 | Benign | 4.06 | Benign | 0.01 | Affected | 0.2292 | 0.3127 | 1 | 0 | 1.7 | -34.02 | |||||||||||||||||||||||||||||
| c.877C>G | R293G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R293G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools uniformly indicate a deleterious effect: SGM‑Consensus, REVEL, FoldX, Rosetta, Foldetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the change as pathogenic, while premPS remains uncertain. High‑accuracy assessments corroborate this trend: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) reports a likely pathogenic outcome, and Foldetta (integrating FoldX‑MD and Rosetta stability outputs) also indicates pathogenicity. No tool suggests a benign effect. Consequently, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.335645 | Structured | 0.338192 | Uncertain | 0.924 | 0.269 | 0.125 | -15.861 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 3.70 | Destabilizing | 0.2 | 2.35 | Destabilizing | 3.03 | Destabilizing | 0.57 | Ambiguous | 0.604 | Likely Pathogenic | -6.43 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | 1.45 | Pathogenic | 0.02 | Affected | 0.3235 | 0.3738 | -3 | -2 | 4.1 | -99.14 | |||||||||||||||||||||||||||||
| c.991T>C | S331P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S331P is not reported in ClinVar (ClinVar ID None) and is absent from gnomAD (gnomAD ID None). Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, PROVEAN, SIFT, ESM1b, polyPhen‑2 HumVar, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are Rosetta, polyPhen‑2 HumDiv, and FATHMM. Two tools give inconclusive results: AlphaMissense‑Default and Foldetta. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also favors benign, while Foldetta remains uncertain. Overall, the majority of evidence points to a benign impact. This conclusion is consistent with the lack of ClinVar annotation and gnomAD presence, so there is no contradiction with existing clinical data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | C2 | 0.433034 | Structured | 0.346458 | Uncertain | 0.658 | 0.475 | 0.250 | -5.104 | Likely Benign | 0.409 | Ambiguous | Likely Benign | 0.33 | Likely Benign | 0.2 | 2.35 | Destabilizing | 1.34 | Ambiguous | 0.44 | Likely Benign | 0.186 | Likely Benign | -1.81 | Neutral | 0.784 | Possibly Damaging | 0.390 | Benign | 1.83 | Pathogenic | 0.26 | Tolerated | 0.2306 | 0.4086 | 1 | -1 | -0.8 | 10.04 | ||||||||||||||||||||||||||||||
| c.1735C>G | R579G 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant R579G is reported in gnomAD (ID 6‑33440787‑C‑G) and has no ClinVar entry. Prediction tools that assess pathogenicity uniformly favor a deleterious effect: SGM‑Consensus (Likely Pathogenic), REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default all predict pathogenic. No tool in the dataset reports a benign outcome; the only uncertain calls are from FoldX, AlphaMissense‑Optimized, and Foldetta. High‑accuracy assessments further support pathogenicity: the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is pathogenic, while AlphaMissense‑Optimized and Foldetta remain uncertain. Consequently, the collective evidence indicates that R579G is most likely pathogenic, and this conclusion is not contradicted by ClinVar status, which is currently absent. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.053060 | Structured | 0.022872 | Uncertain | 0.877 | 0.244 | 0.000 | 6-33440787-C-G | 1 | 6.20e-7 | -14.298 | Likely Pathogenic | 0.948 | Likely Pathogenic | Ambiguous | 1.43 | Ambiguous | 0.0 | 2.36 | Destabilizing | 1.90 | Ambiguous | 1.32 | Destabilizing | 0.680 | Likely Pathogenic | -5.96 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.40 | Pathogenic | 0.01 | Affected | 3.37 | 34 | 0.3078 | 0.2554 | -2 | -3 | 4.1 | -99.14 | ||||||||||||||||||||||||
| c.1774T>C | S592P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S592P is not reported in ClinVar (no ClinVar ID) and is absent from gnomAD. Prediction tools that agree on a benign effect include only SIFT, whereas the remaining tools—REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized—consistently predict a pathogenic impact. High‑accuracy assessments further support this: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is likely pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, is pathogenic. No predictions are inconclusive or missing. Based on the overwhelming agreement among pathogenic predictors and the absence of benign evidence, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status (none is available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.012270 | Structured | 0.100070 | Uncertain | 0.913 | 0.182 | 0.000 | -14.621 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 4.26 | Destabilizing | 0.1 | 2.36 | Destabilizing | 3.31 | Destabilizing | 1.24 | Destabilizing | 0.909 | Likely Pathogenic | -4.77 | Deleterious | 0.999 | Probably Damaging | 0.993 | Probably Damaging | -1.29 | Pathogenic | 0.11 | Tolerated | 0.2412 | 0.4259 | 1 | -1 | -0.8 | 10.04 | |||||||||||||||||||||||||||||
| c.1786C>G | R596G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant R596G is not reported in ClinVar and is absent from gnomAD. Prediction tools that assess pathogenicity largely agree: REVEL, FoldX, Rosetta, Foldetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict a deleterious effect, while premPS remains uncertain. No tools predict a benign outcome. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized is pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is likely pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is pathogenic. Based on the consensus of these predictions, the variant is most likely pathogenic, and this conclusion does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.017797 | Structured | 0.135423 | Uncertain | 0.918 | 0.134 | 0.000 | -13.525 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 4.02 | Destabilizing | 0.1 | 2.36 | Destabilizing | 3.19 | Destabilizing | 0.81 | Ambiguous | 0.629 | Likely Pathogenic | -6.96 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 2.41 | Pathogenic | 0.00 | Affected | 0.3214 | 0.2316 | -3 | -2 | 4.1 | -99.14 | |||||||||||||||||||||||||||||
| c.2119G>A | A707T 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant A707T is reported in gnomAD (ID 6‑33441584‑G‑A) but has no ClinVar entry. Functional prediction tools that agree on a benign effect include REVEL, premPS, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv and polyPhen‑2 HumVar. High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized is benign, and the SGM‑Consensus (derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is also benign. The Foldetta stability prediction is inconclusive and therefore not considered evidence. Overall, the consensus of available predictions indicates that the variant is most likely benign, and this assessment does not contradict any ClinVar classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | 0.203355 | Structured | 0.371229 | Uncertain | 0.927 | 0.365 | 0.000 | 6-33441584-G-A | 1 | 6.20e-7 | -0.836 | Likely Benign | 0.175 | Likely Benign | Likely Benign | 0.50 | Ambiguous | 0.0 | 2.36 | Destabilizing | 1.43 | Ambiguous | 0.10 | Likely Benign | 0.252 | Likely Benign | -0.57 | Neutral | 0.980 | Probably Damaging | 0.947 | Probably Damaging | 3.52 | Benign | 0.44 | Tolerated | 3.50 | 9 | 0.0880 | 0.4330 | 0 | 1 | -2.5 | 30.03 | ||||||||||||||||||||||||
| c.652T>C | F218L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F218L is not reported in ClinVar and is absent from gnomAD. Prediction tools that agree on a benign effect include polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and FATHMM. Those that predict a pathogenic effect comprise REVEL, Rosetta, premPS, PROVEAN, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). FoldX and Foldetta provide uncertain results and are therefore considered unavailable for interpretation. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, the SGM Consensus as likely pathogenic, and Foldetta as uncertain. Overall, the majority of evidence points to a pathogenic impact for F218L, and this conclusion is not contradicted by any ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.281712 | Structured | 0.408725 | Uncertain | 0.848 | 0.272 | 0.000 | -8.861 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.96 | Ambiguous | 0.4 | 2.36 | Destabilizing | 1.66 | Ambiguous | 1.08 | Destabilizing | 0.546 | Likely Pathogenic | -3.80 | Deleterious | 0.158 | Benign | 0.025 | Benign | 5.90 | Benign | 0.15 | Tolerated | 0.2331 | 0.3241 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.654C>A | F218L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F218L is not reported in ClinVar (ClinVar status: none) and is absent from gnomAD (gnomAD ID: none). Prediction tools that agree on a benign effect include REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and FATHMM. Tools that agree on a pathogenic effect comprise SGM‑Consensus (Likely Pathogenic), Rosetta, premPS, PROVEAN, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. FoldX and Foldetta are inconclusive and are treated as unavailable. High‑accuracy methods further support pathogenicity: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also predicts Likely Pathogenic; Foldetta remains uncertain. Overall, the majority of evidence points to a pathogenic impact. There is no ClinVar annotation to contradict this assessment, so the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.281712 | Structured | 0.408725 | Uncertain | 0.848 | 0.272 | 0.000 | -8.861 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.96 | Ambiguous | 0.4 | 2.36 | Destabilizing | 1.66 | Ambiguous | 1.08 | Destabilizing | 0.477 | Likely Benign | -3.80 | Deleterious | 0.158 | Benign | 0.025 | Benign | 5.90 | Benign | 0.15 | Tolerated | 0.2331 | 0.3241 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.654C>G | F218L 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant F218L is not reported in ClinVar (ClinVar status: none) and is absent from gnomAD (gnomAD ID: none). Prediction tools that agree on a benign effect include REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and FATHMM. Tools that agree on a pathogenic effect comprise SGM‑Consensus (Likely Pathogenic), Rosetta, premPS, PROVEAN, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. FoldX and Foldetta are inconclusive and are treated as unavailable. High‑accuracy methods further support pathogenicity: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also predicts Likely Pathogenic; Foldetta remains uncertain. Overall, the majority of evidence points to a pathogenic impact. There is no ClinVar annotation to contradict this assessment, so the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | PH | 0.281712 | Structured | 0.408725 | Uncertain | 0.848 | 0.272 | 0.000 | -8.861 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.96 | Ambiguous | 0.4 | 2.36 | Destabilizing | 1.66 | Ambiguous | 1.08 | Destabilizing | 0.477 | Likely Benign | -3.80 | Deleterious | 0.158 | Benign | 0.025 | Benign | 5.90 | Benign | 0.15 | Tolerated | 0.2331 | 0.3241 | 2 | 0 | 1.0 | -34.02 | |||||||||||||||||||||||||||||
| c.1292T>A | L431Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L431Q resides in the GAP domain. ClinVar has no entry for this variant, and it is not present in gnomAD. Prediction tools that agree on a benign effect are REVEL and FATHMM; all other evaluated algorithms—including FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized—consistently predict a pathogenic impact. High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is Likely Pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. No predictions or stability results are missing or inconclusive. Based on the overwhelming agreement among pathogenic predictions and the high‑accuracy tools, the variant is most likely pathogenic; this conclusion is not contradicted by ClinVar status, which is currently absent. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.094817 | Structured | 0.374755 | Uncertain | 0.959 | 0.300 | 0.000 | -12.399 | Likely Pathogenic | 0.986 | Likely Pathogenic | Likely Pathogenic | 2.69 | Destabilizing | 0.0 | 2.37 | Destabilizing | 2.53 | Destabilizing | 1.99 | Destabilizing | 0.493 | Likely Benign | -5.40 | Deleterious | 1.000 | Probably Damaging | 0.992 | Probably Damaging | 2.92 | Benign | 0.01 | Affected | 0.1036 | 0.1088 | -2 | -2 | -7.3 | 14.97 | |||||||||||||||||||||||||||||
| c.1793T>A | L598H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L598H is not reported in ClinVar and is absent from gnomAD. Prediction tools largely agree on a deleterious effect: FATHMM is the sole benign predictor, while all other evaluated algorithms (REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict pathogenicity. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized returns a pathogenic score; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is Likely Pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, also predicts pathogenicity. No predictions are missing or inconclusive. Based on the overwhelming consensus of pathogenic predictions, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar status (none reported). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.007259 | Structured | 0.147872 | Uncertain | 0.953 | 0.154 | 0.000 | -17.210 | Likely Pathogenic | 0.990 | Likely Pathogenic | Likely Pathogenic | 2.52 | Destabilizing | 0.2 | 2.37 | Destabilizing | 2.45 | Destabilizing | 2.24 | Destabilizing | 0.648 | Likely Pathogenic | -6.87 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.10 | Benign | 0.00 | Affected | 0.1060 | 0.0879 | -2 | -3 | -7.0 | 23.98 | |||||||||||||||||||||||||||||
| c.1865C>A | T622N 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant T622N has no ClinVar entry and is not reported in gnomAD. Functional prediction tools that agree on a benign effect are SIFT and AlphaMissense‑Optimized, whereas a majority of tools predict a pathogenic impact: REVEL, Rosetta, PROVEAN, polyPhen‑2 (HumDiv and HumVar), FATHMM, AlphaMissense‑Default, and the SGM‑Consensus. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as uncertain. No prediction or stability result is missing; all available data are considered. Overall, the preponderance of evidence points to a pathogenic effect for T622N, and this conclusion is not contradicted by ClinVar status, which currently lacks an entry for this variant. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.268042 | Structured | 0.071403 | Uncertain | 0.957 | 0.198 | 0.000 | -7.962 | In-Between | 0.693 | Likely Pathogenic | Likely Benign | 1.00 | Ambiguous | 0.1 | 2.37 | Destabilizing | 1.69 | Ambiguous | 0.94 | Ambiguous | 0.564 | Likely Pathogenic | -4.14 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | -1.24 | Pathogenic | 0.13 | Tolerated | 0.0943 | 0.2848 | 0 | 0 | -2.8 | 13.00 | |||||||||||||||||||||||||||||
| c.809A>G | E270G 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E270G is not reported in ClinVar and is absent from gnomAD. Functional prediction tools uniformly indicate a deleterious effect: SIFT, PolyPhen‑2 (HumDiv and HumVar), REVEL, ESM1b, FATHMM, PROVEAN, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the change as pathogenic. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is labeled Likely Pathogenic. Rosetta also predicts pathogenicity, while FoldX, Foldetta, and premPS are inconclusive. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, SGM‑Consensus as Likely Pathogenic, and Foldetta as uncertain. No tool predicts a benign effect. Consequently, the variant is most likely pathogenic, and this assessment does not contradict any ClinVar annotation (none available). Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | C2 | 0.144935 | Structured | 0.382573 | Uncertain | 0.938 | 0.231 | 0.125 | -13.759 | Likely Pathogenic | 0.968 | Likely Pathogenic | Likely Pathogenic | 1.23 | Ambiguous | 0.2 | 2.37 | Destabilizing | 1.80 | Ambiguous | 0.61 | Ambiguous | 0.576 | Likely Pathogenic | -6.43 | Deleterious | 1.000 | Probably Damaging | 0.996 | Probably Damaging | 1.63 | Pathogenic | 0.00 | Affected | 0.3254 | 0.3876 | 0 | -2 | 3.1 | -72.06 | |||||||||||||||||||||||||||||
| c.1363C>G | L455V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L455V is not reported in ClinVar and is absent from gnomAD. Functional prediction tools largely agree on a deleterious effect: benign predictions come from REVEL and FATHMM, whereas the remaining 11 tools (FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) predict pathogenicity. High‑accuracy assessments reinforce this trend: AlphaMissense‑Optimized is uncertain; the SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is pathogenic; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, also predicts pathogenicity. Taken together, the preponderance of evidence indicates that the variant is most likely pathogenic, and this conclusion is consistent with the absence of a ClinVar annotation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | 0.188120 | Structured | 0.310377 | Uncertain | 0.963 | 0.168 | 0.000 | -12.773 | Likely Pathogenic | 0.946 | Likely Pathogenic | Ambiguous | 2.89 | Destabilizing | 0.1 | 2.38 | Destabilizing | 2.64 | Destabilizing | 1.41 | Destabilizing | 0.276 | Likely Benign | -2.83 | Deleterious | 0.995 | Probably Damaging | 0.970 | Probably Damaging | 3.29 | Benign | 0.02 | Affected | 0.1149 | 0.3056 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||||
Found 8751 rows. Show 800 rows per page. Page 10/11 « Previous | Next »