

Table of SynGAP1 Isoform α2 (UniProt Q96PV0-1) Missense Variants.
| c.dna | Variant | SGM Consensus | Domain | ClinVar | gnomAD | ESM1b | AlphaMissense | REVEL | FoldX | Rosetta | Foldetta | PremPS | PROVEAN | PolyPhen-2 HumDiv | PolyPhen-2 HumVar | FATHMM | SIFT | PAM | Physical | SASA | Normalized B-factor backbone | Normalized B-factor sidechain | SynGAP Structural Annotation | DOI | |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Clinical Status | Review | Subm. | ID | Allele count | Allele freq. | LLR score | Prediction | Pathogenicity | Class | Optimized | Score | Prediction | Average ΔΔG | Prediction | StdDev | ΔΔG | Prediction | ΔΔG | Prediction | ΔΔG | Prediction | Score | Prediction | pph2_prob | Prediction | pph2_prob | Prediction | Nervous System Score | Prediction | Prediction | Status | Conservation | Sequences | PAM250 | PAM120 | Hydropathy Δ | MW Δ | Average | Δ | Δ | StdDev | Δ | StdDev | Secondary | Tertiary bonds | Inside out | GAP-Ras interface | At membrane | No effect | MD Alert | Verdict | Description | |||||
| c.1913A>G | K638R 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant K638R is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that uniformly indicate a benign effect include REVEL, FoldX, Rosetta, Foldetta, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Benign). In contrast, PROVEAN and polyPhen‑2 HumDiv predict a pathogenic impact, while premPS remains inconclusive. High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized is benign; the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is benign; and Foldetta, which integrates FoldX‑MD and Rosetta stability predictions, is benign. Overall, the majority of evidence points to a benign effect, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | Uncertain | 1 | -2.700 | Likely Benign | 0.110 | Likely Benign | Likely Benign | 0.216 | Likely Benign | 0.09 | Likely Benign | 0.1 | -0.04 | Likely Benign | 0.03 | Likely Benign | 0.53 | Ambiguous | -2.55 | Deleterious | 0.649 | Possibly Damaging | 0.240 | Benign | 3.41 | Benign | 0.13 | Tolerated | 3.37 | 31 | 2 | 3 | -0.6 | 28.01 | ||||||||||||||||||||
| c.1942T>C | F648L 2D  AISynGAP1 missense variant F648L is listed in ClinVar with an uncertain significance (ClinVar ID 3383902.0) and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, and FATHMM, whereas the remaining tools—FoldX, Rosetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, AlphaMissense‑Optimized, and ESM1b—consistently predict pathogenicity. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also indicates likely pathogenic. High‑accuracy assessments further support a deleterious effect: AlphaMissense‑Optimized scores pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts a destabilizing, pathogenic change. Taken together, the preponderance of evidence points to a pathogenic impact for F648L, which contradicts the current ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | Uncertain | 1 | -9.296 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.468 | Likely Benign | 2.71 | Destabilizing | 0.8 | 2.08 | Destabilizing | 2.40 | Destabilizing | 1.04 | Destabilizing | -5.98 | Deleterious | 0.999 | Probably Damaging | 0.976 | Probably Damaging | 3.45 | Benign | 0.08 | Tolerated | 2 | 0 | 1.0 | -34.02 | ||||||||||||||||||||||
| c.1947G>C | M649I 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant M649I is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that indicate a benign effect include REVEL, polyPhen‑2 HumVar, and FATHMM, whereas the majority of other in silico predictors (FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) report a pathogenic outcome; Rosetta is inconclusive. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is “Likely Pathogenic,” and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenicity. Overall, the preponderance of evidence points to a pathogenic effect for M649I, which is consistent with the ClinVar “Uncertain” classification rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | Uncertain | 1 | -9.361 | Likely Pathogenic | 0.995 | Likely Pathogenic | Likely Pathogenic | 0.449 | Likely Benign | 2.42 | Destabilizing | 0.2 | 1.96 | Ambiguous | 2.19 | Destabilizing | 1.01 | Destabilizing | -3.99 | Deleterious | 0.672 | Possibly Damaging | 0.093 | Benign | 3.40 | Benign | 0.02 | Affected | 3.38 | 27 | 2 | 1 | 2.6 | -18.03 | 243.7 | 21.5 | 0.0 | 0.1 | 0.0 | 0.1 | X | Potentially Benign | The thioether side chain of Met649, located on an α helix (res. Ser641-Glu666), bridges Phe652, Phe648, and Phe639 in an inter-helix hydrophobic cavity in the WT simulations. In the variant simulations, the sec-butyl side chain of Ile649 maintains hydrophobic interactions with nearby residues, with no significant effects on the protein structure.However, methionine is known as a bridging motif for aromatic residues, and these Met-aromatic interactions are lost in the variant. Indeed, in the second variant simulation,the bridging of Phe652, Phe648 and Phe639 is completely lost. In reality, the effect could be more severe on the structure during the protein folding. | |||||||||||
| c.194A>G | H65R 2D  AIThe SynGAP1 missense variant H65R is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33425802‑A‑G). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumVar, ESM1b, and FATHMM, while those that predict a pathogenic effect are polyPhen‑2 HumDiv, SIFT, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, whereas the SGM‑Consensus remains benign; Foldetta results are unavailable. Overall, the majority of evidence points to a benign impact, and this conclusion does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33425802-A-G | 1 | 6.20e-7 | -1.980 | Likely Benign | 0.967 | Likely Pathogenic | Likely Pathogenic | 0.073 | Likely Benign | -1.60 | Neutral | 0.462 | Possibly Damaging | 0.227 | Benign | 4.19 | Benign | 0.00 | Affected | 4.32 | 1 | 2 | 0 | -1.3 | 19.05 | |||||||||||||||||||||||||||
| c.1957C>G | L653V 2D  AIThe SynGAP1 missense variant L653V is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that classify the variant as benign include REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are FoldX, Rosetta, and premPS, while ESM1b is inconclusive. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as Likely Benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as pathogenic. Overall, the majority of evidence points to a benign impact, and this does not contradict the ClinVar “Uncertain” classification. Thus, based on current predictions, the variant is most likely benign. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | Uncertain | 1 | -7.050 | In-Between | 0.301 | Likely Benign | Likely Benign | 0.146 | Likely Benign | 3.28 | Destabilizing | 0.3 | 2.18 | Destabilizing | 2.73 | Destabilizing | 1.32 | Destabilizing | -2.25 | Neutral | 0.227 | Benign | 0.039 | Benign | 3.28 | Benign | 0.08 | Tolerated | 2 | 1 | 0.4 | -14.03 | ||||||||||||||||||||||
| c.1964T>A | L655Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L655Q is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, FoldX, Foldetta, premPS, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) all indicate benign or likely benign. Only the two polyPhen‑2 implementations (HumDiv and HumVar) predict pathogenicity, while Rosetta remains inconclusive. High‑accuracy assessments reinforce the benign consensus: AlphaMissense‑Optimized scores benign, the SGM‑Consensus is likely benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) predicts benign stability. Overall, the majority of evidence supports a benign impact for L655Q, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | Uncertain | 1 | -5.278 | Likely Benign | 0.144 | Likely Benign | Likely Benign | 0.139 | Likely Benign | -0.01 | Likely Benign | 0.0 | 0.69 | Ambiguous | 0.34 | Likely Benign | -0.15 | Likely Benign | 0.61 | Neutral | 0.955 | Possibly Damaging | 0.602 | Possibly Damaging | 3.59 | Benign | 0.65 | Tolerated | 3.39 | 24 | -2 | -2 | -7.3 | 14.97 | 229.9 | -8.6 | 0.0 | 0.0 | 0.4 | 0.0 | X | Potentially Benign | The iso-butyl side chain of Leu655, located on the surface of an α helix (res. Ser641-Glu666), is not involved in any interactions in the WT simulations. In the variant simulations, the carboxamide side chain of Gln655 dynamically interacts with neighboring residues (e.g., Glu651, Glu656, Arg544) on the protein surface, with no negative structural effects. | |||||||||||
| c.1966G>C | E656Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant E656Q is listed in ClinVar with an uncertain significance and is present in gnomAD (ID 6‑33441225‑G‑C). Functional prediction tools that agree on a benign effect include REVEL, FoldX, Foldetta, premPS, PROVEAN, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic outcome are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, and AlphaMissense‑Default; Rosetta reports an uncertain result. High‑accuracy assessments show AlphaMissense‑Optimized as benign, Foldetta as benign, while the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive due to a 2‑2 split. Overall, the majority of evidence points to a benign effect, and this conclusion does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | Uncertain | 1 | 6-33441225-G-C | 1 | 6.20e-7 | -9.145 | Likely Pathogenic | 0.766 | Likely Pathogenic | Likely Benign | 0.249 | Likely Benign | -0.14 | Likely Benign | 0.0 | -0.81 | Ambiguous | -0.48 | Likely Benign | 0.25 | Likely Benign | -2.29 | Neutral | 0.980 | Probably Damaging | 0.528 | Possibly Damaging | 3.46 | Benign | 0.02 | Affected | 3.39 | 24 | 2 | 2 | 0.0 | -0.98 | 224.3 | 1.7 | 0.0 | 0.1 | 0.1 | 0.0 | X | Potentially Benign | The carboxylate side chain of Glu656, located on an α helix (res. Ser641-Glu666), frequently forms a hydrogen bond with the nearby residue Ser659 on the same α helix. In the variant simulations, the carboxamide side chain of Gln656 alternatively forms a hydrogen bond with either Ser659 or Glu548 on an opposing helix (res. Ala533-Val560).Although the frequent interaction between Gln656 and Glu548 may strengthen or stabilize the tertiary structure assembly, the effect is likely to be marginal. | |||||||||
| c.196C>G | P66A 2D  AIThe SynGAP1 P66A missense variant (ClinVar ID 1303518.0) is listed as “Uncertain” and is not reported in gnomAD. Functional prediction tools that agree on benign impact include REVEL, PROVEAN, ESM1b, and FATHMM, while polyPhen‑2 (HumDiv and HumVar), SIFT, and AlphaMissense‑Default all predict pathogenicity. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” status. Separately, the high‑accuracy AlphaMissense‑Optimized result is “Uncertain,” the SGM‑Consensus remains “Likely Benign,” and Foldetta (a protein‑folding stability method combining FoldX‑MD and Rosetta outputs) has no available result for this variant. Overall, the predictions are mixed, but the majority of high‑confidence tools lean toward a benign effect. Thus, the variant is most likely benign based on current computational evidence, and this assessment does not contradict the ClinVar status of uncertainty. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -2.845 | Likely Benign | 0.891 | Likely Pathogenic | Ambiguous | 0.091 | Likely Benign | -1.56 | Neutral | 0.805 | Possibly Damaging | 0.539 | Possibly Damaging | 4.04 | Benign | 0.00 | Affected | 4.32 | 1 | 1 | -1 | 3.4 | -26.04 | ||||||||||||||||||||||||||||||
| c.1970G>T | W657L 2D 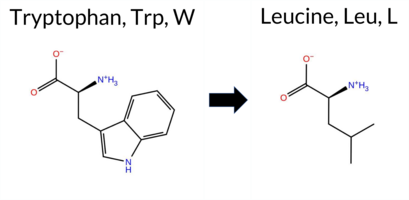 3DClick to see structure in 3D Viewer AISynGAP1 missense variant W657L is listed in ClinVar with an uncertain significance (ClinVar ID 2767440.0) and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, FoldX, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and FATHMM. Tools that agree on a pathogenic effect are PROVEAN, ESM1b, and AlphaMissense‑Default. Uncertain predictions come from Foldetta, premPS, and Rosetta. High‑accuracy assessments show AlphaMissense‑Optimized predicts pathogenicity, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also predicts pathogenic, and Foldetta predicts a benign folding‑stability change. Overall, the majority of evidence points toward a pathogenic impact, which is consistent with the ClinVar uncertain status but leans toward pathogenic rather than benign. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | Uncertain | 1 | -14.411 | Likely Pathogenic | 0.960 | Likely Pathogenic | Likely Pathogenic | 0.213 | Likely Benign | 0.14 | Likely Benign | 0.1 | 0.73 | Ambiguous | 0.44 | Likely Benign | 0.87 | Ambiguous | -10.86 | Deleterious | 0.277 | Benign | 0.078 | Benign | 3.52 | Benign | 0.14 | Tolerated | 3.39 | 24 | -2 | -2 | 4.7 | -73.05 | ||||||||||||||||||||
| c.1971G>C | W657C 2D 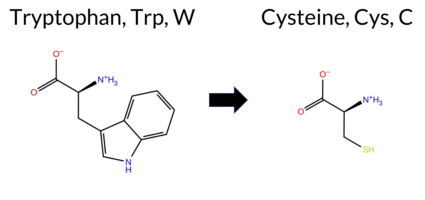 AISynGAP1 missense variant W657C is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools that classify the variant as benign include REVEL and FATHMM. Those that predict a deleterious effect are FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized; Rosetta reports an uncertain outcome. High‑accuracy assessments further support a damaging interpretation: AlphaMissense‑Optimized is pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. Overall, the preponderance of evidence indicates that W657C is most likely pathogenic, which does not contradict the current ClinVar status of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | Uncertain | 1 | -12.035 | Likely Pathogenic | 0.997 | Likely Pathogenic | Likely Pathogenic | 0.463 | Likely Benign | 2.74 | Destabilizing | 0.3 | 1.69 | Ambiguous | 2.22 | Destabilizing | 1.30 | Destabilizing | -11.06 | Deleterious | 1.000 | Probably Damaging | 0.982 | Probably Damaging | 3.43 | Benign | 0.03 | Affected | -8 | -2 | 3.4 | -83.07 | ||||||||||||||||||||||
| c.1973G>A | G658D 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant G658D is listed in ClinVar with an uncertain significance and is present in gnomAD (ID 6-33441232‑G‑A). Functional prediction tools that agree on a benign effect include REVEL, FoldX, premPS, polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, and AlphaMissense‑Optimized. Only PROVEAN predicts a pathogenic outcome, while Rosetta, Foldetta, ESM1b, and AlphaMissense‑Default are inconclusive. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive, and Foldetta, which integrates FoldX‑MD and Rosetta, is also inconclusive. Overall, the preponderance of evidence points to a benign effect, and this does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | Uncertain | 1 | 6-33441232-G-A | 3 | 1.86e-6 | -7.786 | In-Between | 0.442 | Ambiguous | Likely Benign | 0.144 | Likely Benign | -0.40 | Likely Benign | 0.1 | -0.59 | Ambiguous | -0.50 | Ambiguous | 0.46 | Likely Benign | -2.64 | Deleterious | 0.008 | Benign | 0.005 | Benign | 3.53 | Benign | 0.38 | Tolerated | 3.39 | 24 | 1 | -1 | -3.1 | 58.04 | 219.8 | -84.3 | 0.0 | 0.0 | 0.2 | 0.1 | X | Potentially Pathogenic | Gly658, located on the outer surface of an α helix (res. Ser641-Glu666), weakens the helix integrity at that spot, which is necessary for the kink in the middle of the long helix. In the variant simulations, the carboxylic acid side chain of Asp658 is on the surface of the α helix and is not involved in any interactions. However, aspartate is not as effective a breaker of the secondary structure element as glycine, which may lead to misfolding. | |||||||||
| c.1976C>T | S659F 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S659F is listed in ClinVar with an uncertain significance and is absent from gnomAD. Functional prediction tools that provide definitive calls cluster into two groups: benign predictions come from REVEL, Rosetta, premPS, polyPhen2_HumVar, FATHMM, and AlphaMissense‑Optimized; pathogenic predictions come from PROVEAN, polyPhen2_HumDiv, SIFT, ESM1b, AlphaMissense‑Default, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). High‑accuracy assessments show AlphaMissense‑Optimized predicts benign, SGM Consensus predicts pathogenic, and Foldetta (which integrates FoldX‑MD and Rosetta outputs) yields an uncertain result and is therefore unavailable. Overall, the majority of reliable tools favor a pathogenic effect. Thus, the variant is most likely pathogenic, a conclusion that does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | Uncertain | 1 | -10.925 | Likely Pathogenic | 0.662 | Likely Pathogenic | Likely Benign | 0.194 | Likely Benign | -0.81 | Ambiguous | 0.1 | -0.25 | Likely Benign | -0.53 | Ambiguous | 0.32 | Likely Benign | -4.59 | Deleterious | 0.806 | Possibly Damaging | 0.171 | Benign | 3.39 | Benign | 0.05 | Affected | 3.38 | 28 | -3 | -2 | 3.6 | 60.10 | 221.3 | -61.2 | 0.0 | 0.0 | 0.6 | 0.4 | X | Potentially Benign | In the WT simulations, the hydroxyl group of Ser659, located in a kink in the middle of the long α-helix (res. Ser641-Glu666), forms a hydrogen bond with the carboxylate group of Glu656. However, the phenol ring of the Phe659 side chain cannot form a similar hydrogen bond. Instead, it interacts with the hydrophobic isopropyl side chain of Val555 from the opposing α-helix (res. Ala533-Val560). This residue swap may therefore cause issues during protein folding. | |||||||||||
| c.1998G>C | E666D 2D  3DClick to see structure in 3D Viewer AISynGAP1 E666D is listed in ClinVar with an uncertain significance (ID 587483.0) and is not reported in gnomAD. Functional prediction tools show a mixed signal: benign calls come from REVEL, SIFT, FATHMM, AlphaMissense‑Optimized, and Rosetta; pathogenic calls come from premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Default. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is classified as likely pathogenic. High‑accuracy assessments give AlphaMissense‑Optimized a benign prediction, while the SGM Consensus remains pathogenic; Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, is inconclusive. Overall, the balance of evidence slightly favors a pathogenic interpretation, but the predictions are not unequivocal. Thus, the variant is most likely pathogenic according to the current computational data, and this does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | Uncertain | 1 | -8.820 | Likely Pathogenic | 0.704 | Likely Pathogenic | Likely Benign | 0.197 | Likely Benign | 0.88 | Ambiguous | 0.0 | 0.37 | Likely Benign | 0.63 | Ambiguous | 1.05 | Destabilizing | -2.69 | Deleterious | 0.992 | Probably Damaging | 0.603 | Possibly Damaging | 3.43 | Benign | 0.06 | Tolerated | 3.38 | 28 | 3 | 2 | 0.0 | -14.03 | 237.2 | 16.5 | 0.0 | 0.0 | -0.3 | 0.1 | X | Potentially Pathogenic | The carboxylate group of Glu666, located on the α-helix (res. Ser641-Glu666), is involved in a highly coordinated hydrogen-bonding network between residues from two α-helices (res. Ser641-Glu666 and res. Arg563-Glu578) and from the α-α loop connecting the two α-helices (res. Ser641-Glu666 and res. Leu685-Val699), such as Lys566, Thr672, and Asn669, in the WT simulations. In the variant simulations, the shorter side chain of Asp666 cannot maintain these interactions as efficiently as Glu666 in the WT, resulting in a less coordinated hydrogen-bond network. | |||||||||||
| c.2015C>A | T672K 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant T672K is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, FoldX, polyPhen‑2 HumVar, SIFT, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are SGM‑Consensus, PROVEAN, polyPhen‑2 HumDiv, ESM1b, and AlphaMissense‑Default. Uncertain predictions come from Foldetta, premPS, and Rosetta. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta as inconclusive. Overall, the majority of tools lean toward a benign interpretation, but the high‑accuracy consensus is split, leaving the variant’s impact uncertain. Thus, the variant is most likely benign based on the bulk of predictions, and this does not contradict its ClinVar status of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | Uncertain | 1 | -12.192 | Likely Pathogenic | 0.698 | Likely Pathogenic | Likely Benign | 0.065 | Likely Benign | 0.20 | Likely Benign | 0.5 | 1.21 | Ambiguous | 0.71 | Ambiguous | 0.72 | Ambiguous | -4.31 | Deleterious | 0.745 | Possibly Damaging | 0.051 | Benign | 3.40 | Benign | 0.07 | Tolerated | 3.40 | 25 | 0 | -1 | -3.2 | 27.07 | 195.1 | 7.0 | 0.4 | 0.7 | 0.4 | 0.1 | X | X | Potentially Pathogenic | The hydroxyl group of Thr672, located in an entangled α-α loop connecting the two α-helices (res. Ser641-Glu666 and res. Leu685-Val699), is involved in a highly coordinated hydrogen-bonding network between residues from two α-helices (res. Ser641-Glu666 and res. Arg563-Glu578) and from the α-α loop itself, such as Lys566, Glu666, and Asn669. In the variant simulations, Lys672 can only form a hydrogen bond with the amino group of the Lys566 side chain via its backbone carbonyl group. Consequently, it cannot maintain the Lys566-Glu666 salt bridge through hydrogen bonding. However, the amino group of Lys periodically forms a salt bridge with the carboxylate group of Glu666, which prevents a drastic disruption of the hydrogen-bond network that keeps the loop close to the helices. | ||||||||||
| c.2047A>G | I683V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I683V is listed in ClinVar with an uncertain significance and is present in gnomAD (6‑33441306‑A‑G). Across a panel of in silico predictors, the majority indicate a benign effect: REVEL, PROVEAN, polyPhen‑2 HumVar, SIFT, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (derived from a majority of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). Only polyPhen‑2 HumDiv classifies the change as pathogenic. High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized is benign, the SGM‑Consensus (majority vote) is benign, and Foldetta, which integrates FoldX‑MD and Rosetta stability predictions, is inconclusive and therefore not considered evidence. No other tool provides a pathogenic signal. Consequently, the variant is most likely benign, and this assessment does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | Uncertain | 1 | 6-33441306-A-G | 2 | 1.24e-6 | -7.588 | In-Between | 0.138 | Likely Benign | Likely Benign | 0.112 | Likely Benign | 0.90 | Ambiguous | 0.0 | 0.60 | Ambiguous | 0.75 | Ambiguous | 0.76 | Ambiguous | -0.78 | Neutral | 0.538 | Possibly Damaging | 0.080 | Benign | 3.35 | Benign | 0.14 | Tolerated | 3.42 | 17 | 4 | 3 | -0.3 | -14.03 | 215.6 | 29.1 | 0.0 | 0.0 | -0.7 | 0.1 | X | Potentially Benign | The sec-butyl side chain of Ile683, located in an entangled α-α loop connecting the two α-helices (res. Ser641-Glu666 and res. Leu685-Val699), is sterically packed against His453 and Glu688. In the variant simulations, the iso-propyl side chain of Val683 has similar size and physicochemical properties as Ile630 in the WT, and thus, it is able to maintain similar interactions in the inter-helix space. Consequently, no negative structural effects are observed during the simulations due to the residue swap. | ||||||||
| c.2050G>A | D684N 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant D684N is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that indicate a benign effect include REVEL, premPS, and FATHMM, whereas the majority of tools predict a pathogenic outcome: PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized classifies the variant as pathogenic, the SGM‑Consensus also reports it as likely pathogenic, and the Foldetta stability analysis is inconclusive. Protein‑stability predictors FoldX and Rosetta likewise return uncertain results. Overall, the preponderance of evidence points to a pathogenic effect, which contradicts the current ClinVar designation of uncertainty. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | Uncertain | 1 | -13.155 | Likely Pathogenic | 0.985 | Likely Pathogenic | Likely Pathogenic | 0.382 | Likely Benign | 1.47 | Ambiguous | 0.8 | 1.76 | Ambiguous | 1.62 | Ambiguous | 0.37 | Likely Benign | -4.99 | Deleterious | 0.999 | Probably Damaging | 0.746 | Possibly Damaging | 3.39 | Benign | 0.01 | Affected | 2 | 1 | 0.0 | -0.98 | ||||||||||||||||||||||
| c.2050G>C | D684H 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant D684H is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include only FATHMM; all other evaluated algorithms (REVEL, FoldX, Rosetta, Foldetta, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic impact, and the SGM‑Consensus score is “Likely Pathogenic.” High‑accuracy assessments further support pathogenicity: AlphaMissense‑Optimized is pathogenic, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) is pathogenic. No predictions are inconclusive or missing. Based on the overwhelming majority of pathogenic predictions, the variant is most likely pathogenic, which does not contradict the current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | Uncertain | 1 | -14.194 | Likely Pathogenic | 0.998 | Likely Pathogenic | Likely Pathogenic | 0.613 | Likely Pathogenic | 3.36 | Destabilizing | 1.0 | 2.95 | Destabilizing | 3.16 | Destabilizing | 0.55 | Ambiguous | -6.98 | Deleterious | 1.000 | Probably Damaging | 0.972 | Probably Damaging | 3.36 | Benign | 0.00 | Affected | 3.42 | 17 | -1 | 1 | 0.3 | 22.05 | ||||||||||||||||||||
| c.2068T>C | S690P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S690P is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that indicate a benign effect are REVEL and FATHMM, whereas the remaining tools (FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus) all predict a pathogenic outcome. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized scores the variant as pathogenic; the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is “Likely Pathogenic”; and Foldetta, which integrates FoldX‑MD and Rosetta stability predictions, also classifies the variant as pathogenic. Overall, the preponderance of evidence from multiple independent predictors indicates that the variant is most likely pathogenic, a conclusion that does not contradict the current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | Uncertain | 1 | -14.568 | Likely Pathogenic | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.431 | Likely Benign | 4.84 | Destabilizing | 0.3 | 4.40 | Destabilizing | 4.62 | Destabilizing | 1.42 | Destabilizing | -4.77 | Deleterious | 0.998 | Probably Damaging | 0.790 | Possibly Damaging | 3.44 | Benign | 0.01 | Affected | 3.42 | 17 | 1 | -1 | -0.8 | 10.04 | 207.5 | 15.1 | 0.1 | 0.0 | -0.1 | 0.2 | X | X | Potentially Pathogenic | The hydroxyl side chain of Ser690, located in an α-helix (res. Leu696-Leu685), forms a hydrogen bond with the backbone carbonyl group of Ser410 in an anti-parallel β-sheet of the C2 domain (res. Ile411-Ala399). In the variant simulations, the pyrrolidine side chain of Pro690 cannot form hydrogen bonds with the C2 domain residue, resulting in the loss of this inter-domain connection. Additionally, prolines lack a free amide group necessary for hydrogen bonding with the carbonyl group of Gly686, introducing a slight bend in the α-helix and compromising its integrity. | ||||||||||
| c.2075T>C | L692P 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant L692P is listed in ClinVar with an “Uncertain” status (ClinVar ID 847082.0) and is not reported in gnomAD. Prediction tools that agree on a benign effect are limited to FATHMM, while the remaining tools (REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized) uniformly predict a pathogenic impact. High‑accuracy assessments further support this: AlphaMissense‑Optimized predicts pathogenic; the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates “Likely Pathogenic”; and Foldetta, which integrates FoldX‑MD and Rosetta outputs, also predicts pathogenic. Based on the overwhelming consensus of pathogenic predictions, the variant is most likely pathogenic, which does not contradict its current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | Uncertain | 1 | -16.447 | Likely Pathogenic | 1.000 | Likely Pathogenic | Likely Pathogenic | 0.668 | Likely Pathogenic | 9.19 | Destabilizing | 0.1 | 13.20 | Destabilizing | 11.20 | Destabilizing | 1.69 | Destabilizing | -6.98 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 3.06 | Benign | 0.00 | Affected | 3.42 | 17 | -3 | -3 | -5.4 | -16.04 | 186.2 | 62.8 | -0.2 | 0.1 | -0.7 | 0.3 | X | Potentially Pathogenic | The isobutyl side chain of Leu692, located in the middle of an α-helix (res. Leu685-Gln702), engages in hydrophobic packing with nearby residues (e.g., Leu441, Leu431, Leu696) in the inter-helix space. Prolines lack a free amide group necessary for hydrogen bonding with the carbonyl group of Glu688 in the same manner as Leu692 in the WT. Consequently, the residue swap with proline disrupts the continuity of the secondary structure element in the variant simulations. Additionally, the side chain of Pro692 is not as optimal as Leu692 for hydrophobic packing in the inter-helix space. | |||||||||||
| c.2086C>G | L696V 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 L696V variant is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that indicate a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized, whereas the majority of other in silico predictors (FoldX, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) report a pathogenic outcome; Rosetta remains inconclusive. High‑accuracy assessments further support a deleterious impact: AlphaMissense‑Optimized predicts benign, but the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—leans pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenic. Overall, the preponderance of evidence points to a pathogenic effect for the variant, which does not contradict the current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | Uncertain | 1 | -11.909 | Likely Pathogenic | 0.745 | Likely Pathogenic | Likely Benign | 0.351 | Likely Benign | 2.35 | Destabilizing | 0.1 | 1.85 | Ambiguous | 2.10 | Destabilizing | 1.46 | Destabilizing | -2.79 | Deleterious | 0.992 | Probably Damaging | 0.970 | Probably Damaging | 3.16 | Benign | 0.00 | Affected | 3.46 | 13 | 1 | 2 | 0.4 | -14.03 | ||||||||||||||||||||
| c.2095G>A | V699M 2D 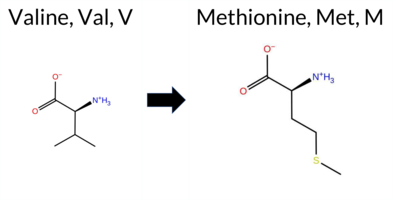 3DClick to see structure in 3D Viewer AISynGAP1 variant V699M is listed in ClinVar with an uncertain significance and is present in gnomAD (ID 6‑33441354‑G‑A). Across in silico predictors, benign calls are made by REVEL, Rosetta, Foldetta, PROVEAN, FATHMM, and AlphaMissense‑Optimized, whereas pathogenic calls come from polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and ESM1b. Predictions that are inconclusive (FoldX, premPS, AlphaMissense‑Default) are noted but not used as evidence. High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized predicts benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also yields benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) reports benign stability. Overall, the preponderance of evidence indicates the variant is most likely benign, which does not contradict the ClinVar uncertain status but provides a stronger leaning toward benignity. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | Uncertain | 2 | 6-33441354-G-A | 8 | 4.96e-6 | -8.869 | Likely Pathogenic | 0.484 | Ambiguous | Likely Benign | 0.276 | Likely Benign | -0.58 | Ambiguous | 0.1 | 0.29 | Likely Benign | -0.15 | Likely Benign | 0.96 | Ambiguous | -2.18 | Neutral | 0.994 | Probably Damaging | 0.806 | Possibly Damaging | 3.37 | Benign | 0.03 | Affected | 3.47 | 10 | 2 | 1 | -2.3 | 32.06 | 257.8 | -47.2 | 0.0 | 0.0 | 0.9 | 0.1 | X | Potentially Benign | The isopropyl side chain of Val699, located on an α-helix (res. Leu685-Gln702), packs against hydrophobic residues (e.g., Leu703, Leu696, Leu435, Leu439) in the inter-helix space. In the variant simulations, the thioether side chain of Met699 has similar physicochemical properties to Val699 in the WT, and thus, it is able to maintain similar interactions. Consequently, the mutation causes no apparent changes in the structure. | |||||||||
| c.2101C>T | P701S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant P701S (ClinVar ID 2995856.0) is listed as “Uncertain” in ClinVar and is present in gnomAD (ID 6‑33441360‑C‑T). Prediction tools that agree on a benign effect include REVEL, Rosetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Benign). No tool in the dataset predicts a pathogenic outcome; all remaining predictions are either benign or uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as Benign, the SGM‑Consensus as Likely Benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) as Uncertain. Based on the collective evidence, the variant is most likely benign, and this conclusion does not contradict the ClinVar status, which remains uncertain. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | Uncertain | 1 | 6-33441360-C-T | 3 | 1.86e-6 | -4.375 | Likely Benign | 0.221 | Likely Benign | Likely Benign | 0.132 | Likely Benign | 1.33 | Ambiguous | 0.0 | 0.12 | Likely Benign | 0.73 | Ambiguous | -0.36 | Likely Benign | 0.78 | Neutral | 0.044 | Benign | 0.025 | Benign | 3.48 | Benign | 1.00 | Tolerated | 3.47 | 10 | -1 | 1 | 0.8 | -10.04 | 10.1016/j.ajhg.2020.11.011 | ||||||||||||||||
| c.2105A>G | Q702R 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant Q702R is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, FoldX, Foldetta, premPS, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT. Predictions that remain inconclusive are Rosetta, ESM1b, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as benign, Foldetta as benign, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive due to equal benign and pathogenic signals. Overall, the majority of available predictions lean toward a benign impact, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | Uncertain | 1 | -7.894 | In-Between | 0.348 | Ambiguous | Likely Benign | 0.294 | Likely Benign | -0.31 | Likely Benign | 0.1 | 0.63 | Ambiguous | 0.16 | Likely Benign | 0.13 | Likely Benign | -3.14 | Deleterious | 0.909 | Possibly Damaging | 0.889 | Possibly Damaging | 3.43 | Benign | 0.02 | Affected | 3.47 | 10 | 1 | 1 | -1.0 | 28.06 | 270.3 | -52.9 | 0.0 | 0.0 | 0.0 | 0.1 | X | Potentially Pathogenic | The carboxamide side chain of Gln702 is located at the end and outer surface of an α-helix (res. Leu685-Gln702), where it does not directly form hydrogen bonds with any residues in the WT simulations. In the variant simulations, the positively charged guanidinium group of Arg702 forms a salt bridge with the negatively charged carboxylate group of Glu698 on the same helix and/or hydrogen bonds with the backbone carbonyl group of Ala438 on an opposite α-helix (res. Tyr428-Glu436). Consequently, the residue swap could strengthen the tertiary structure assembly, which could have either positive or negative effects on its function. | ||||||||||||
| c.2111G>C | S704T 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant S704T is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Consensus from multiple in‑silico predictors shows a predominance of benign calls: REVEL, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus. Only polyPhen‑2 HumDiv predicts a pathogenic effect, while FoldX remains inconclusive. High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized is benign; the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is benign; and Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, also predicts benign. Overall, the aggregate evidence indicates that S704T is most likely benign, which is consistent with its ClinVar uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | Uncertain | 1 | -4.930 | Likely Benign | 0.265 | Likely Benign | Likely Benign | 0.071 | Likely Benign | 0.80 | Ambiguous | 0.0 | 0.15 | Likely Benign | 0.48 | Likely Benign | 0.29 | Likely Benign | -1.72 | Neutral | 0.525 | Possibly Damaging | 0.107 | Benign | 3.45 | Benign | 0.07 | Tolerated | 3.47 | 10 | 1 | 1 | 0.1 | 14.03 | 201.7 | -18.0 | 0.0 | 0.0 | -0.2 | 0.7 | X | Potentially Benign | Ser704 is located at the end and outer surface of an α-helix (res. Thr704-Gly712), which is connected via a tight turn or loop to another α-helix (res. Asp684-Gln702). The hydroxyl side chain of Ser704 occasionally forms a hydrogen bond with the amide group of Ala707. Similarly, in the variant simulations, the hydroxyl side chain of Thr704 forms hydrogen bonds with the amide groups of Ala707 and Leu708. Thus, the residue swap does not cause any apparent structural change. | |||||||||||
| c.2113A>C | K705Q 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 K705Q missense variant (ClinVar ID 3699560.0) is listed as “Uncertain” in ClinVar and is present in gnomAD (variant ID 6‑33441372‑A‑C). Prediction tools that uniformly indicate a benign effect include REVEL, FoldX, Rosetta, Foldetta, premPS, PROVEAN, SIFT, ESM1b, FATHMM, and AlphaMissense‑Optimized. In contrast, polyPhen‑2 (both HumDiv and HumVar models) predict a pathogenic outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign; the SGM Consensus—derived from a majority vote of AlphaMissense‑Default (uncertain), ESM1b (benign), FATHMM (benign), and PROVEAN (benign)—also yields a benign classification; Foldetta, which integrates FoldX‑MD and Rosetta stability outputs, predicts benign. Overall, the preponderance of evidence supports a benign impact for K705Q, and this conclusion does not contradict the ClinVar “Uncertain” status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | Uncertain | 1 | 6-33441372-A-C | 1 | 6.20e-7 | -5.787 | Likely Benign | 0.436 | Ambiguous | Likely Benign | 0.142 | Likely Benign | -0.10 | Likely Benign | 0.0 | 0.33 | Likely Benign | 0.12 | Likely Benign | -0.02 | Likely Benign | -0.24 | Neutral | 0.997 | Probably Damaging | 0.969 | Probably Damaging | 3.42 | Benign | 0.78 | Tolerated | 3.47 | 10 | 1 | 1 | 0.4 | -0.04 | |||||||||||||||||
| c.2116G>A | E706K 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant E706K is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM all classify the change as benign. In contrast, ESM1b and AlphaMissense‑Default predict a pathogenic impact. Tools that return uncertain results—FoldX, Rosetta, Foldetta, and AlphaMissense‑Optimized—do not provide decisive evidence. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is inconclusive (two pathogenic versus two benign calls). High‑accuracy assessments are likewise ambiguous: AlphaMissense‑Optimized is uncertain, Foldetta is uncertain, and the SGM Consensus remains inconclusive. Overall, the preponderance of evidence points to a benign effect, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | GAP | Uncertain | 1 | -10.519 | Likely Pathogenic | 0.833 | Likely Pathogenic | Ambiguous | 0.080 | Likely Benign | 1.17 | Ambiguous | 0.1 | 0.51 | Ambiguous | 0.84 | Ambiguous | 0.08 | Likely Benign | -1.51 | Neutral | 0.345 | Benign | 0.028 | Benign | 4.15 | Benign | 0.52 | Tolerated | 3.47 | 10 | 0 | 1 | -0.4 | -0.94 | 187.1 | 49.2 | 0.0 | 0.0 | 0.4 | 0.1 | X | Uncertain | The carboxylate side chain of Glu706, located at the end and outer surface of an α-helix (res. Thr704-Gly712), forms a salt bridge with Lys710 and a hydrogen bond with its own backbone amino group at the helix end in the WT simulations. Although Lys706 is unable to make these transient interactions in the variant simulations, there is no apparent negative effect on the protein structure due to the residue swap. However, because the model ends abruptly at the C-terminus, no definite conclusions can be drawn based on the simulations. | ||||||||||||
| c.2131C>G | L711V 2D  3DClick to see structure in 3D Viewer AISynGAP1 missense variant L711V is listed in ClinVar with an uncertain significance and is present in gnomAD (ID 6‑33441596‑C‑G). Prediction tools that indicate a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized. The majority of other in silico predictors—FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default—classify the change as pathogenic, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) reports it as likely pathogenic. High‑accuracy assessments further show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely pathogenic, and Foldetta (combining FoldX‑MD and Rosetta outputs) as pathogenic. Overall, the preponderance of evidence points to a pathogenic effect, which does not conflict with the ClinVar designation of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | Uncertain | 1 | 6-33441596-C-G | 1 | 6.20e-7 | -10.045 | Likely Pathogenic | 0.709 | Likely Pathogenic | Likely Benign | 0.170 | Likely Benign | 3.48 | Destabilizing | 0.1 | 2.22 | Destabilizing | 2.85 | Destabilizing | 1.40 | Destabilizing | -2.59 | Deleterious | 0.992 | Probably Damaging | 0.970 | Probably Damaging | 3.34 | Benign | 0.00 | Affected | 3.50 | 9 | 1 | 2 | 0.4 | -14.03 | |||||||||||||||||
| c.2162T>G | I721S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant I721S is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that assess the variant’s effect fall into two groups: the single benign prediction comes from REVEL, while all other evaluated algorithms (FoldX, Rosetta, Foldetta, premPS, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized) predict a pathogenic outcome. High‑accuracy methods reinforce this view: AlphaMissense‑Optimized reports pathogenic, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is “Likely Pathogenic,” and Foldetta (combining FoldX‑MD and Rosetta outputs) also predicts pathogenic. No predictions are missing or inconclusive. Based on the overwhelming consensus of pathogenic predictions, the variant is most likely pathogenic, which does not contradict its current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | GAP | Uncertain | 1 | -14.032 | Likely Pathogenic | 0.996 | Likely Pathogenic | Likely Pathogenic | 0.466 | Likely Benign | 3.91 | Destabilizing | 0.1 | 3.96 | Destabilizing | 3.94 | Destabilizing | 2.28 | Destabilizing | -5.26 | Deleterious | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 2.21 | Pathogenic | 0.00 | Affected | 3.50 | 9 | -1 | -2 | -5.3 | -26.08 | 203.3 | 49.3 | -0.1 | 0.0 | -1.1 | 0.0 | X | Uncertain | The sec-butyl side chain of Ile721, located on an α-helix (res. Leu714-Arg726), engages in hydrophobic packing with other residues in the hydrophobic inter-helix space, such as Phe420, Tyr417, His693, and Leu717. In the variant simulations, the hydroxyl side chain of Ser721 forms hydrogen bonds with nearby residues, such as Leu717 and His693. Although no major structural changes are observed during the variant simulations, the hydrophilic residue Ser721 could disrupt the hydrophobic packing during folding. However, because the model ends abruptly at the C-terminus, no definite conclusions can be drawn based on the simulations. | |||||||||||
| c.2186A>G | N729S 2D  3DClick to see structure in 3D Viewer AIThe SynGAP1 missense variant N729S is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, FoldX, premPS, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). No tool in the dataset predicts a pathogenic outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus also as benign, while Foldetta (combining FoldX‑MD and Rosetta outputs) is inconclusive. Based on the collective predictions, the variant is most likely benign, and this assessment does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | GAP | Uncertain | 1 | -1.578 | Likely Benign | 0.066 | Likely Benign | Likely Benign | 0.063 | Likely Benign | 0.14 | Likely Benign | 0.1 | 1.34 | Ambiguous | 0.74 | Ambiguous | -0.36 | Likely Benign | -0.42 | Neutral | 0.221 | Benign | 0.027 | Benign | 3.38 | Benign | 0.93 | Tolerated | 3.59 | 7 | 1 | 1 | 2.7 | -27.03 | ||||||||||||||||||||
| c.218G>A | R73K 2D  AIThe SynGAP1 missense variant R73K is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33425826‑G‑A). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Benign). Only SIFT predicts a pathogenic outcome. High‑accuracy assessments further support a benign classification: AlphaMissense‑Optimized returns a benign prediction, and the SGM‑Consensus (derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates benign. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the preponderance of evidence points to a benign effect, which does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33425826-G-A | 2 | 1.24e-6 | -4.033 | Likely Benign | 0.151 | Likely Benign | Likely Benign | 0.077 | Likely Benign | -0.46 | Neutral | 0.053 | Benign | 0.007 | Benign | 4.14 | Benign | 0.00 | Affected | 4.32 | 1 | 2 | 3 | 0.6 | -28.01 | |||||||||||||||||||||||||||
| c.2195G>C | R732T 2D  AISynGAP1 missense variant R732T is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign (REVEL, PROVEAN, SIFT, FATHMM, AlphaMissense‑Optimized) and pathogenic (polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b). AlphaMissense‑Default remains uncertain. The high‑accuracy AlphaMissense‑Optimized predicts a benign effect, and the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—also favors a benign outcome. Foldetta, a protein‑folding stability method that integrates FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the preponderance of evidence from both general and high‑accuracy predictors points to a benign impact, which does not contradict the current ClinVar designation of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | -8.545 | Likely Pathogenic | 0.434 | Ambiguous | Likely Benign | 0.075 | Likely Benign | -1.96 | Neutral | 0.999 | Probably Damaging | 0.892 | Possibly Damaging | 2.59 | Benign | 0.12 | Tolerated | 3.59 | 7 | -1 | -1 | 3.8 | -55.08 | |||||||||||||||||||||||||||||||
| c.2200C>T | P734S 2D  AIThe SynGAP1 missense variant P734S is listed in ClinVar with an uncertain significance (ClinVar ID 2283225.0) and is present in the gnomAD database (gnomAD ID 6‑33441665‑C‑T). Functional prediction tools uniformly classify the variant as benign: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all report benign effects. No tool predicts pathogenicity. The high‑accuracy consensus methods corroborate this benign assessment: AlphaMissense‑Optimized is benign, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates a likely benign outcome. Foldetta, a protein‑folding stability predictor combining FoldX‑MD and Rosetta outputs, did not provide a result for this variant, so its status is unavailable. Overall, the computational evidence strongly supports a benign classification, which is consistent with the ClinVar uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 2 | 6-33441665-C-T | 2 | 1.24e-6 | -4.291 | Likely Benign | 0.077 | Likely Benign | Likely Benign | 0.030 | Likely Benign | -2.44 | Neutral | 0.344 | Benign | 0.048 | Benign | 2.77 | Benign | 0.11 | Tolerated | 3.64 | 6 | 1 | -1 | 0.8 | -10.04 | 10.1016/j.ajhg.2020.11.011 | ||||||||||||||||||||||||||
| c.2207G>A | R736H 2D  AIThe SynGAP1 missense variant R736H is listed in ClinVar (ID 1351080.0) with an “Uncertain” status and is present in gnomAD (variant ID 6‑33441672‑G‑A). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Benign). Only SIFT predicts a pathogenic outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also as benign. Foldetta results are not available. Overall, the majority of computational evidence indicates a benign impact, and this does not contradict the ClinVar “Uncertain” classification. Thus, the variant is most likely benign based on current predictions. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33441672-G-A | 6 | 3.72e-6 | -5.409 | Likely Benign | 0.067 | Likely Benign | Likely Benign | 0.029 | Likely Benign | -0.12 | Neutral | 0.004 | Benign | 0.001 | Benign | 2.50 | Benign | 0.00 | Affected | 4.07 | 3 | 2 | 0 | 1.3 | -19.05 | |||||||||||||||||||||||||||
| c.2210A>C | Q737P 2D  AIThe SynGAP1 missense variant Q737P is listed in ClinVar (ID 2580571.0) with an uncertain significance designation and is not reported in gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict benign, while only SIFT indicates a pathogenic effect. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also reports a likely benign outcome. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized is benign and the SGM‑Consensus is likely benign; Foldetta stability analysis is unavailable. Taken together, the preponderance of evidence supports a benign classification for Q737P, which is consistent with its ClinVar uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -2.407 | Likely Benign | 0.054 | Likely Benign | Likely Benign | 0.154 | Likely Benign | -1.22 | Neutral | 0.005 | Benign | 0.013 | Benign | 2.78 | Benign | 0.04 | Affected | 4.07 | 3 | -1 | 0 | 1.9 | -31.01 | ||||||||||||||||||||||||||||||
| c.2215G>C | E739Q 2D  AIThe SynGAP1 missense variant E739Q is listed in ClinVar (ID 2429558.0) with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv and SIFT. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta results are unavailable. Overall, the majority of evidence points to a benign impact, and this is consistent with the ClinVar “Uncertain” designation rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -2.846 | Likely Benign | 0.161 | Likely Benign | Likely Benign | 0.071 | Likely Benign | -1.06 | Neutral | 0.801 | Possibly Damaging | 0.339 | Benign | 2.57 | Benign | 0.00 | Affected | 4.32 | 2 | 2 | 2 | 0.0 | -0.98 | ||||||||||||||||||||||||||||||
| c.2216A>T | E739V 2D  AIThe SynGAP1 missense variant E739V is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. In contrast, polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM all predict a pathogenic impact. The SGM‑Consensus, which aggregates the majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as “Likely Benign.” High‑accuracy assessments further support a benign classification: AlphaMissense‑Optimized predicts benign, and the SGM‑Consensus (majority vote) also indicates benign. The Foldetta protein‑folding stability analysis is unavailable for this variant. Overall, the preponderance of evidence from both general and high‑accuracy predictors points to a benign effect, and this conclusion does not contradict the current ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -3.136 | Likely Benign | 0.274 | Likely Benign | Likely Benign | 0.085 | Likely Benign | -1.86 | Neutral | 0.891 | Possibly Damaging | 0.575 | Possibly Damaging | 2.47 | Pathogenic | 0.00 | Affected | 4.32 | 2 | -2 | -2 | 7.7 | -29.98 | ||||||||||||||||||||||||||||||
| c.2217G>C | E739D 2D  AIThe SynGAP1 missense variant E739D is listed in ClinVar (ID 3661302.0) with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). Only SIFT predicts a pathogenic outcome. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized is benign, and the SGM‑Consensus is “Likely Benign.” Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the majority of evidence points to a benign effect, and this is not in conflict with the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -3.369 | Likely Benign | 0.062 | Likely Benign | Likely Benign | 0.097 | Likely Benign | -0.49 | Neutral | 0.002 | Benign | 0.005 | Benign | 2.59 | Benign | 0.00 | Affected | 3 | 2 | 0.0 | -14.03 | ||||||||||||||||||||||||||||||||
| c.2218C>T | R740W 2D  AIThe SynGAP1 missense variant R740W is listed in ClinVar with an uncertain significance and is present in the gnomAD database (ID 6‑33441683‑C‑T). Prediction tools that classify the variant as benign include REVEL, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized, while those that predict pathogenicity are PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and ESM1b. High‑accuracy assessments show AlphaMissense‑Optimized predicting a benign effect; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is inconclusive (two benign vs. two pathogenic calls) and is treated as unavailable, and no Foldetta stability data are reported. Overall, the majority of conventional tools (five pathogenic vs. four benign) suggest a pathogenic impact, whereas the single high‑accuracy tool indicates benign. Thus, the variant is most likely pathogenic based on the aggregate predictions, and this assessment does not contradict the ClinVar status of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 2 | 6-33441683-C-T | 6 | 3.72e-6 | -8.561 | Likely Pathogenic | 0.168 | Likely Benign | Likely Benign | 0.180 | Likely Benign | -3.09 | Deleterious | 1.000 | Probably Damaging | 0.938 | Probably Damaging | 2.52 | Benign | 0.01 | Affected | 4.32 | 2 | 2 | -3 | 3.6 | 30.03 | ||||||||||||||||||||||||||||
| c.2219G>A | R740Q 2D  AIThe SynGAP1 missense variant R740Q is listed in ClinVar with an “Uncertain” status and is present in gnomAD (variant ID 6‑33441684‑G‑A). Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. In contrast, polyPhen‑2 (both HumDiv and HumVar models) predict a pathogenic outcome. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized is benign, and the SGM‑Consensus is also benign. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no reported result for this variant, so it does not influence the assessment. Overall, the majority of predictions indicate that R740Q is most likely benign, which is consistent with the ClinVar “Uncertain” classification and does not contradict it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33441684-G-A | 4 | 2.48e-6 | -5.195 | Likely Benign | 0.078 | Likely Benign | Likely Benign | 0.102 | Likely Benign | -0.67 | Neutral | 0.999 | Probably Damaging | 0.881 | Possibly Damaging | 2.60 | Benign | 0.08 | Tolerated | 4.32 | 2 | 1 | 1 | 1.0 | -28.06 | |||||||||||||||||||||||||||
| c.221G>A | S74N 2D  AIThe SynGAP1 missense variant S74N is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33425829‑G‑A). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Only SIFT predicts a pathogenic outcome. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized is benign, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is “Likely Benign.” No Foldetta stability result is available. Overall, the majority of computational evidence indicates that the variant is most likely benign, which does not contradict the current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33425829-G-A | 5 | 3.10e-6 | -5.156 | Likely Benign | 0.112 | Likely Benign | Likely Benign | 0.031 | Likely Benign | -0.89 | Neutral | 0.043 | Benign | 0.007 | Benign | 4.09 | Benign | 0.00 | Affected | 4.32 | 1 | 1 | 1 | -2.7 | 27.03 | |||||||||||||||||||||||||||
| c.2221C>T | P741S 2D  AIThe SynGAP1 missense variant P741S is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33441686‑C‑T). Functional prediction tools largely agree on a benign effect: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the change as benign, while the single pathogenic prediction comes from SIFT. Grouping by consensus, the benign‑predicting tools outnumber the pathogenic one. High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized predicts benign, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) reports “Likely Benign.” No Foldetta stability data are available, so it does not influence the conclusion. Overall, the computational evidence indicates that the variant is most likely benign, and this assessment does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 2 | 6-33441686-C-T | 3 | 1.86e-6 | -3.700 | Likely Benign | 0.063 | Likely Benign | Likely Benign | 0.076 | Likely Benign | -0.27 | Neutral | 0.270 | Benign | 0.136 | Benign | 2.92 | Benign | 0.00 | Affected | 4.32 | 2 | 1 | -1 | 0.8 | -10.04 | 10.1016/j.ajhg.2020.11.011 | ||||||||||||||||||||||||||
| c.2224C>T | R742W 2D  AIThe SynGAP1 missense variant R742W is listed in ClinVar (ID 2581888.0) with an “Uncertain” status and is present in gnomAD (variant ID 6‑33441689‑C‑T). Prediction tools that agree on a benign effect include REVEL, PROVEAN, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Benign). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT; ESM1b is uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as benign, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also favors benign. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the majority of computational evidence points to a benign impact, which is consistent with the ClinVar “Uncertain” classification and does not contradict it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33441689-C-T | 6 | 3.72e-6 | -7.725 | In-Between | 0.133 | Likely Benign | Likely Benign | 0.079 | Likely Benign | -1.71 | Neutral | 0.992 | Probably Damaging | 0.684 | Possibly Damaging | 2.66 | Benign | 0.01 | Affected | 4.32 | 2 | -3 | 2 | 3.6 | 30.03 | |||||||||||||||||||||||||||
| c.2225G>A | R742Q 2D  AIThe SynGAP1 missense variant R742Q is listed in ClinVar (ID 928481.0) with an uncertain significance annotation and is observed in gnomAD (variant ID 6‑33441690‑G‑A). Consensus from multiple in‑silico predictors—REVEL, PROVEAN, PolyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized—uniformly classify the change as benign. No tool in the dataset reports a pathogenic prediction. High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized predicts benign, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates a likely benign effect. A protein‑folding stability analysis via Foldetta is not available for this variant. Overall, the computational evidence strongly favors a benign interpretation, which is consistent with the ClinVar uncertain status rather than contradicting it. The variant is most likely benign, and this assessment does not contradict its ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 2 | 6-33441690-G-A | 24 | 1.49e-5 | -4.090 | Likely Benign | 0.068 | Likely Benign | Likely Benign | 0.054 | Likely Benign | -0.19 | Neutral | 0.032 | Benign | 0.007 | Benign | 2.73 | Benign | 0.07 | Tolerated | 4.32 | 2 | 1 | 1 | 1.0 | -28.06 | |||||||||||||||||||||||||||
| c.2239G>C | V747L 2D  AIThe SynGAP1 missense variant V747L (ClinVar ID 1985039.0) is listed as ClinVar status Uncertain and is present in gnomAD (6‑33441704‑G‑C). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Only SIFT predicts a pathogenic outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as Likely Benign; Foldetta stability analysis is unavailable. Overall, the majority of computational evidence supports a benign classification, which is consistent with the ClinVar Uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33441704-G-C | 2 | 1.24e-6 | -2.790 | Likely Benign | 0.096 | Likely Benign | Likely Benign | 0.047 | Likely Benign | -0.52 | Neutral | 0.065 | Benign | 0.033 | Benign | 2.67 | Benign | 0.00 | Affected | 4.32 | 2 | 2 | 1 | -0.4 | 14.03 | |||||||||||||||||||||||||||
| c.2249G>A | G750E 2D  AISynGAP1 missense variant G750E is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, PROVEAN, ESM1b, and AlphaMissense‑Optimized, while pathogenic predictions arise from polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM; AlphaMissense‑Default remains uncertain. High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized predicts benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also yields benign, and Foldetta results are unavailable. Overall, the majority of evidence points to a benign effect, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | -2.618 | Likely Benign | 0.413 | Ambiguous | Likely Benign | 0.146 | Likely Benign | -2.27 | Neutral | 1.000 | Probably Damaging | 0.982 | Probably Damaging | 2.49 | Pathogenic | 0.01 | Affected | 3.99 | 5 | 0 | -2 | -3.1 | 72.06 | |||||||||||||||||||||||||||||||
| c.2255C>T | S752L 2D  AIThe SynGAP1 missense variant S752L is listed in ClinVar with an “Uncertain” status (ClinVar ID 2143952.0) and is present in gnomAD (ID 6‑33441720‑C‑T). Prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and FATHMM. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta results are unavailable. Overall, the preponderance of evidence points to a benign impact, which does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 2 | 6-33441720-C-T | 6 | 3.72e-6 | -3.386 | Likely Benign | 0.182 | Likely Benign | Likely Benign | 0.195 | Likely Benign | -2.09 | Neutral | 0.993 | Probably Damaging | 0.641 | Possibly Damaging | 1.51 | Pathogenic | 0.01 | Affected | 3.99 | 5 | -3 | -2 | 4.6 | 26.08 | |||||||||||||||||||||||||||
| c.2270G>C | G757A 2D  AIThe SynGAP1 missense change G757A is catalogued in ClinVar (ID 3635272.0) with an uncertain significance designation and is not reported in gnomAD. Functional prediction algorithms uniformly classify the variant as benign: REVEL, PROVEAN, PolyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all return benign scores. No tool in the dataset predicts pathogenicity. High‑accuracy consensus methods corroborate this view: the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates a likely benign effect, and AlphaMissense‑Optimized also predicts benign. The Foldetta stability assessment is unavailable for this variant. Taken together, the evidence overwhelmingly supports a benign interpretation, which is consistent with the ClinVar uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -2.626 | Likely Benign | 0.091 | Likely Benign | Likely Benign | 0.066 | Likely Benign | -0.45 | Neutral | 0.267 | Benign | 0.127 | Benign | 2.73 | Benign | 0.35 | Tolerated | 1 | 0 | 2.2 | 14.03 | ||||||||||||||||||||||||||||||||
| c.2275A>C | M759L 2D  AIThe SynGAP1 missense variant M759L is listed in ClinVar with an uncertain significance (ClinVar ID 942432.0) and is present in gnomAD (gnomAD ID 6‑33441740‑A‑C). All evaluated in‑silico predictors agree on a benign effect: REVEL, PROVEAN, PolyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all return benign classifications. No tool predicts pathogenicity. High‑accuracy assessments reinforce this consensus: AlphaMissense‑Optimized is benign, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is “Likely Benign.” Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the computational evidence strongly supports a benign impact, which is consistent with the ClinVar uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33441740-A-C | 2 | 1.24e-6 | -2.431 | Likely Benign | 0.093 | Likely Benign | Likely Benign | 0.048 | Likely Benign | -0.53 | Neutral | 0.002 | Benign | 0.005 | Benign | 2.84 | Benign | 1.00 | Tolerated | 3.99 | 5 | 4 | 2 | 1.9 | -18.03 | |||||||||||||||||||||||||||
| c.2277G>A | M759I 2D  AIThe SynGAP1 missense variant M759I is listed in ClinVar (ID 3686687.0) with an “Uncertain” status and is present in gnomAD (variant ID 6‑33441742‑G‑A). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Benign). Only polyPhen‑2 HumDiv predicts a pathogenic outcome, while AlphaMissense‑Default remains uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as Likely Benign; Foldetta results are unavailable. Overall, the majority of evidence points to a benign impact, and this is consistent with the ClinVar “Uncertain” classification rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33441742-G-A | 1 | 6.20e-7 | -4.058 | Likely Benign | 0.393 | Ambiguous | Likely Benign | 0.075 | Likely Benign | -0.88 | Neutral | 0.454 | Possibly Damaging | 0.192 | Benign | 2.83 | Benign | 0.34 | Tolerated | 3.99 | 5 | 1 | 2 | 2.6 | -18.03 | |||||||||||||||||||||||||||
| c.227C>G | S76C 2D  AIThe SynGAP1 missense variant S76C is listed in ClinVar with an “Uncertain” status (ClinVar ID 1951273.0) and is present in the gnomAD database (gnomAD ID 6‑33425835‑C‑G). Prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus also as benign; Foldetta results are not available for this variant. Overall, the majority of computational evidence points to a benign impact, which does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33425835-C-G | 2 | 1.24e-6 | -5.408 | Likely Benign | 0.100 | Likely Benign | Likely Benign | 0.076 | Likely Benign | -1.78 | Neutral | 0.992 | Probably Damaging | 0.869 | Possibly Damaging | 3.71 | Benign | 0.00 | Affected | 4.32 | 1 | 0 | -1 | 3.3 | 16.06 | |||||||||||||||||||||||||||
| c.2282G>A | R761Q 2D  AIThe SynGAP1 missense variant R761Q is listed in ClinVar (ID 2882770.0) with an “Uncertain” status and is present in gnomAD (6‑33441747‑G‑A). Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv and polyPhen‑2 HumVar. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta results are not available. Overall, the majority of evidence points to a benign impact, and this conclusion does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33441747-G-A | 11 | 6.81e-6 | -4.187 | Likely Benign | 0.202 | Likely Benign | Likely Benign | 0.191 | Likely Benign | -0.63 | Neutral | 0.996 | Probably Damaging | 0.878 | Possibly Damaging | 2.75 | Benign | 0.40 | Tolerated | 3.99 | 5 | 1 | 1 | 1.0 | -28.06 | |||||||||||||||||||||||||||
| c.2282G>C | R761P 2D  AIThe SynGAP1 missense variant R761P is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33441747‑G‑C). Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and AlphaMissense‑Default. The SGM‑Consensus, which aggregates the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the majority of predictions point to a benign impact, and this is consistent with the ClinVar “Uncertain” designation rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 3 | 6-33441747-G-C | 1 | 6.20e-7 | -5.091 | Likely Benign | 0.640 | Likely Pathogenic | Likely Benign | 0.201 | Likely Benign | -1.89 | Neutral | 0.999 | Probably Damaging | 0.968 | Probably Damaging | 2.69 | Benign | 0.38 | Tolerated | 3.99 | 5 | 0 | -2 | 2.9 | -59.07 | |||||||||||||||||||||||||||
| c.2294G>A | S765N 2D  AIThe SynGAP1 missense variant S765N (ClinVar ID 2979632.0) is listed as “Uncertain” in ClinVar and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Benign). In contrast, PolyPhen‑2 (HumDiv and HumVar) predict a pathogenic impact. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized is benign, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is also benign. No Foldetta stability prediction is available, so it does not influence the assessment. Overall, the majority of evidence points to a benign effect, which is consistent with the ClinVar “Uncertain” classification and does not contradict it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -5.098 | Likely Benign | 0.378 | Ambiguous | Likely Benign | 0.094 | Likely Benign | -0.94 | Neutral | 0.985 | Probably Damaging | 0.950 | Probably Damaging | 4.11 | Benign | 0.06 | Tolerated | 3.64 | 6 | 1 | 1 | -2.7 | 27.03 | ||||||||||||||||||||||||||||||
| c.2299A>G | I767V 2D  AIThe SynGAP1 missense variant I767V is listed in ClinVar (ID 1402700.0) with an “Uncertain” clinical significance and is not reported in gnomAD. All evaluated in‑silico predictors classify the substitution as benign: REVEL, PROVEAN, PolyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all return benign scores. The SGM‑Consensus, which aggregates the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also reports a “Likely Benign” outcome. No tool predicts pathogenicity. High‑accuracy assessments confirm this: AlphaMissense‑Optimized is benign, SGM‑Consensus is likely benign, while Foldetta (combining FoldX‑MD and Rosetta stability predictions) has no available result for this variant. Overall, the computational evidence strongly supports a benign effect, and this conclusion does not contradict the current ClinVar status of uncertainty. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -2.791 | Likely Benign | 0.064 | Likely Benign | Likely Benign | 0.096 | Likely Benign | 0.10 | Neutral | 0.072 | Benign | 0.029 | Benign | 4.21 | Benign | 1.00 | Tolerated | 3.64 | 6 | 4 | 3 | -0.3 | -14.03 | ||||||||||||||||||||||||||||||
| c.2300T>C | I767T 2D  AIThe SynGAP1 missense variant I767T is listed in ClinVar (ID 1044161.0) with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict benign. Only polyPhen‑2 HumDiv flags the variant as pathogenic, creating a single discordant prediction. The high‑accuracy consensus from SGM (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) yields a “Likely Benign” classification, and AlphaMissense‑Optimized also reports benign. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the preponderance of evidence points to a benign impact, and this conclusion does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -3.749 | Likely Benign | 0.252 | Likely Benign | Likely Benign | 0.138 | Likely Benign | -0.78 | Neutral | 0.625 | Possibly Damaging | 0.249 | Benign | 4.12 | Benign | 0.46 | Tolerated | 3.64 | 6 | 0 | -1 | -5.2 | -12.05 | ||||||||||||||||||||||||||||||
| c.2302G>A | D768N 2D  AIThe SynGAP1 missense variant D768N is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33442460‑G‑A). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). No tool predicts a pathogenic outcome; AlphaMissense‑Default is uncertain. High‑accuracy assessments further support a benign classification: AlphaMissense‑Optimized predicts benign, the SGM‑Consensus is “Likely Benign,” and Foldetta data are unavailable. Overall, the consensus of available predictions indicates that the variant is most likely benign, which does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33442460-G-A | 2 | 2.57e-6 | -6.892 | Likely Benign | 0.453 | Ambiguous | Likely Benign | 0.048 | Likely Benign | -0.77 | Neutral | 0.106 | Benign | 0.009 | Benign | 4.07 | Benign | 0.96 | Tolerated | 3.64 | 6 | 1 | 2 | 0.0 | -0.98 | |||||||||||||||||||||||||||
| c.2302G>T | D768Y 2D  AIThe SynGAP1 missense variant D768Y is listed in ClinVar with status “Uncertain” (ClinVar ID 1061652.0) and is present in gnomAD (variant ID 6‑33442460‑G‑T). Prediction tools that agree on a benign effect include REVEL, SIFT, and FATHMM, whereas those that predict a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Default. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as “Likely Pathogenic.” High‑accuracy assessments show AlphaMissense‑Optimized as “Uncertain,” SGM‑Consensus as “Likely Pathogenic,” and Foldetta (combining FoldX‑MD and Rosetta outputs) is unavailable for this variant. Overall, the majority of computational evidence points to a pathogenic impact, which does not contradict the ClinVar designation of uncertainty. Thus, based on current predictions, the variant is most likely pathogenic. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | Uncertain | 1 | 6-33442460-G-T | -9.866 | Likely Pathogenic | 0.824 | Likely Pathogenic | Ambiguous | 0.234 | Likely Benign | -2.86 | Deleterious | 0.989 | Probably Damaging | 0.806 | Possibly Damaging | 4.01 | Benign | 0.07 | Tolerated | 3.64 | 6 | -4 | -3 | 2.2 | 48.09 | |||||||||||||||||||||||||||||
| c.2305C>T | L769F 2D  AIThe SynGAP1 missense variant L769F is listed in ClinVar (ID 3617309.0) with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. In contrast, polyPhen‑2 (HumDiv and HumVar) and SIFT all predict a pathogenic impact. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is “Likely Benign.” High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized predicts benign, and the SGM‑Consensus (majority of the high‑accuracy tools) also indicates benign. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the majority of predictions—including the high‑accuracy tools—suggest the variant is most likely benign, which is consistent with its ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -5.044 | Likely Benign | 0.146 | Likely Benign | Likely Benign | 0.060 | Likely Benign | -0.89 | Neutral | 0.925 | Possibly Damaging | 0.510 | Possibly Damaging | 3.94 | Benign | 0.02 | Affected | 2 | 0 | -1.0 | 34.02 | ||||||||||||||||||||||||||||||||
| c.2339C>G | S780C 2D  AIThe SynGAP1 missense variant S780C is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33442891‑C‑G). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). No tool in the dataset predicts a pathogenic outcome; ESM1b is inconclusive and therefore treated as unavailable. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign, while Foldetta results are not reported and thus unavailable. Based on the collective predictions, the variant is most likely benign, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 4 | 6-33442891-C-G | 16 | 9.94e-6 | -7.603 | In-Between | 0.278 | Likely Benign | Likely Benign | 0.078 | Likely Benign | -1.41 | Neutral | 0.065 | Benign | 0.043 | Benign | 2.59 | Benign | 0.10 | Tolerated | 3.64 | 6 | -1 | 0 | 3.3 | 16.06 | |||||||||||||||||||||||||||
| c.233G>T | R78L 2D  AIThe SynGAP1 missense variant R78L is listed in ClinVar (ID 3390541.0) with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus score (which is a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). Tools that predict a pathogenic effect are SIFT and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as benign; the Foldetta protein‑folding stability analysis is unavailable for this variant. Overall, the majority of evidence points to a benign impact, and this conclusion does not contradict the current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -3.389 | Likely Benign | 0.635 | Likely Pathogenic | Likely Benign | 0.062 | Likely Benign | -1.59 | Neutral | 0.385 | Benign | 0.021 | Benign | 3.84 | Benign | 0.00 | Affected | -3 | -2 | 8.3 | -43.03 | ||||||||||||||||||||||||||||||||
| c.2353C>T | R785C 2D  AIThe SynGAP1 R785C missense variant is listed in ClinVar with an “Uncertain” status and is present in gnomAD (variant ID 6‑33442905‑C‑T). Prediction tools that agree on a benign effect include REVEL, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are PROVEAN, SIFT, FATHMM, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as benign, while the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—indicates a likely pathogenic outcome. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the majority of computational evidence points toward a pathogenic impact, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | SH3-binding motif | Uncertain | 1 | 6-33442905-C-T | 29 | 1.80e-5 | -5.887 | Likely Benign | 0.662 | Likely Pathogenic | Likely Benign | 0.126 | Likely Benign | -5.06 | Deleterious | 0.144 | Benign | 0.046 | Benign | 2.22 | Pathogenic | 0.00 | Affected | 3.64 | 6 | -4 | -3 | 7.0 | -53.05 | ||||||||||||||||||||||||||
| c.2354G>A | R785H 2D  AIThe SynGAP1 R785H missense variant (ClinVar ID 2321588.0) is listed as “Uncertain” in ClinVar and is present in gnomAD (ID 6‑33442906‑G‑A). Prediction tools that agree on a benign effect include REVEL, ESM1b, and AlphaMissense‑Optimized, while those that predict a pathogenic effect are PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM; AlphaMissense‑Default remains uncertain. High‑accuracy assessments show AlphaMissense‑Optimized classifying the variant as benign, whereas the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates a pathogenic outcome. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, does not provide a result for this variant. Overall, the majority of computational predictions (five pathogenic versus three benign) lean toward a pathogenic interpretation. Thus, the variant is most likely pathogenic based on current predictions, and this conclusion does not contradict the ClinVar status, which remains uncertain. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | SH3-binding motif | Uncertain | 2 | 6-33442906-G-A | 4 | 2.50e-6 | -4.782 | Likely Benign | 0.388 | Ambiguous | Likely Benign | 0.129 | Likely Benign | -2.61 | Deleterious | 0.999 | Probably Damaging | 0.947 | Probably Damaging | 2.25 | Pathogenic | 0.01 | Affected | 3.64 | 6 | 2 | 0 | 1.3 | -19.05 | |||||||||||||||||||||||||||
| c.2359C>T | P787S 2D  AIThe SynGAP1 P787S variant is listed in ClinVar with an “Uncertain” status and is present in gnomAD (variant ID 6‑33442911‑C‑T). Prediction tools that agree on a benign effect include REVEL, ESM1b, and AlphaMissense‑Optimized, while those that predict a pathogenic outcome are PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM; AlphaMissense‑Default remains uncertain. The high‑accuracy consensus (SGM Consensus) derived from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN yields a pathogenic majority. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the majority of evidence points to a pathogenic effect, and this assessment does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | SH3-binding motif | Uncertain | 1 | 6-33442911-C-T | 3 | 1.86e-6 | -4.203 | Likely Benign | 0.564 | Ambiguous | Likely Benign | 0.221 | Likely Benign | -3.81 | Deleterious | 1.000 | Probably Damaging | 0.999 | Probably Damaging | 2.48 | Pathogenic | 0.02 | Affected | 3.64 | 6 | -1 | 1 | 0.8 | -10.04 | |||||||||||||||||||||||||||
| c.2362T>A | S788T 2D  AIThe SynGAP1 missense variant S788T is listed in ClinVar with an uncertain significance (ClinVar ID 392728.0) and is present in the gnomAD database (gnomAD ID 6‑33442914‑T‑A). Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score, which is derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN. Tools that predict a pathogenic outcome are polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM. High‑accuracy assessments show AlphaMissense‑Optimized classifying the variant as benign, while the SGM‑Consensus (majority vote) also favors a benign interpretation. No Foldetta stability prediction is available for this variant. Overall, the majority of computational evidence points to a benign effect, which is consistent with the ClinVar uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | SH3-binding motif | Uncertain | 2 | 6-33442914-T-A | 4 | 2.49e-6 | -4.288 | Likely Benign | 0.288 | Likely Benign | Likely Benign | 0.092 | Likely Benign | -2.25 | Neutral | 0.979 | Probably Damaging | 0.982 | Probably Damaging | 1.55 | Pathogenic | 0.02 | Affected | 3.64 | 6 | 1 | 1 | 0.1 | 14.03 | ||||||||||||||||||||||||||
| c.2369C>G | T790S 2D  AIThe SynGAP1 missense variant T790S is listed in ClinVar (ID 1020340.0) with an “Uncertain” clinical significance and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Benign). In contrast, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and FATHMM predict a pathogenic impact. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus also as likely benign; a Foldetta stability analysis is unavailable. Overall, the majority of evidence points to a benign effect, and this is consistent with the ClinVar “Uncertain” status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | SH3-binding motif | Uncertain | 1 | -3.914 | Likely Benign | 0.123 | Likely Benign | Likely Benign | 0.134 | Likely Benign | -1.83 | Neutral | 0.997 | Probably Damaging | 0.989 | Probably Damaging | 2.39 | Pathogenic | 0.33 | Tolerated | 3.64 | 6 | 1 | 1 | -0.1 | -14.03 | |||||||||||||||||||||||||||||
| c.2393C>T | P798L 2D  AIThe SynGAP1 missense variant P798L is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33442945‑C‑T). Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Benign). In contrast, polyPhen‑2 (HumDiv and HumVar) and SIFT all predict a pathogenic impact. High‑accuracy assessments show AlphaMissense‑Optimized as Benign and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Benign; a Foldetta stability prediction is not available. Overall, the majority of evidence points to a benign effect, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | SH3-binding motif | Uncertain | 2 | 6-33442945-C-T | 6 | 3.72e-6 | -5.640 | Likely Benign | 0.074 | Likely Benign | Likely Benign | 0.042 | Likely Benign | -0.86 | Neutral | 0.981 | Probably Damaging | 0.631 | Possibly Damaging | 4.21 | Benign | 0.00 | Affected | 4.32 | 1 | -3 | -3 | 5.4 | 16.04 | ||||||||||||||||||||||||||
| c.2401G>A | G801S 2D  AIThe SynGAP1 missense variant G801S is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. All available in‑silico predictors classify the change as benign: SGM‑Consensus (Likely Benign), REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool predicts pathogenicity. High‑accuracy assessments reinforce this benign prediction: AlphaMissense‑Optimized is benign, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is also Likely Benign. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no reported result for this variant. Overall, the evidence strongly supports a benign effect, and this conclusion does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | SH3-binding motif | Uncertain | 1 | -3.665 | Likely Benign | 0.087 | Likely Benign | Likely Benign | 0.039 | Likely Benign | -0.41 | Neutral | 0.009 | Benign | 0.019 | Benign | 2.76 | Benign | 0.48 | Tolerated | 4.32 | 2 | 0 | 1 | -0.4 | 30.03 | |||||||||||||||||||||||||||||
| c.2405G>A | G802D 2D  AIThe SynGAP1 missense variant G802D is listed in ClinVar with an “Uncertain” status and is present in gnomAD (6‑33442957‑G‑A). Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and FATHMM—all classifying the change as benign. No tool predicts a pathogenic outcome. The high‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized predicts benign, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates a likely benign effect. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the consensus of available predictions points to a benign impact, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | SH3-binding motif | Uncertain | 1 | 6-33442957-G-A | 1 | 6.20e-7 | -5.083 | Likely Benign | 0.476 | Ambiguous | Likely Benign | 0.153 | Likely Benign | -0.38 | Neutral | 0.126 | Benign | 0.138 | Benign | 2.72 | Benign | 0.09 | Tolerated | 3.77 | 5 | 1 | -1 | -3.1 | 58.04 | ||||||||||||||||||||||||||
| c.2408A>G | K803R 2D  AIThe SynGAP1 missense variant K803R is listed in ClinVar (ID 834618.0) with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools largely agree on a benign effect: SGM‑Consensus, REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict benign. Only two tools—SIFT and FATHMM—predict pathogenicity. High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized is benign, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is “Likely Benign.” Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the preponderance of evidence indicates the variant is most likely benign, which does not contradict the current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | SH3-binding motif | Uncertain | 1 | -2.281 | Likely Benign | 0.097 | Likely Benign | Likely Benign | 0.018 | Likely Benign | -1.52 | Neutral | 0.103 | Benign | 0.038 | Benign | 2.38 | Pathogenic | 0.00 | Affected | 3.77 | 5 | 3 | 2 | -0.6 | 28.01 | |||||||||||||||||||||||||||||
| c.2414T>C | L805P 2D  AIThe SynGAP1 missense variant L805P is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools show a split: benign calls come from REVEL, ESM1b, and AlphaMissense‑Optimized, while pathogenic calls are made by PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM; AlphaMissense‑Default remains uncertain. High‑accuracy assessments give a benign result from AlphaMissense‑Optimized, a pathogenic outcome from the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN), and no Foldetta data are available. Overall, the majority of predictions lean toward pathogenicity, and this conclusion does not conflict with the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | SH3-binding motif | Uncertain | 1 | -4.661 | Likely Benign | 0.444 | Ambiguous | Likely Benign | 0.272 | Likely Benign | -3.40 | Deleterious | 0.975 | Probably Damaging | 0.767 | Possibly Damaging | 2.36 | Pathogenic | 0.00 | Affected | 3.77 | 5 | -3 | -3 | -5.4 | -16.04 | ||||||||||||||||||||||||||||||
| c.2420A>G | Y807C 2D  AIThe SynGAP1 missense variant Y807C is listed in ClinVar with an “Uncertain” status (ClinVar ID 2119812.0) and is present in gnomAD (ID 6‑33442972‑A‑G). Prediction tools that agree on a benign effect include REVEL, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that agree on a pathogenic effect are PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM; ESM1b is uncertain. High‑accuracy assessments show AlphaMissense‑Optimized predicting benign, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) predicts pathogenic. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the majority of predictions (five pathogenic vs. three benign) and the SGM Consensus support a pathogenic interpretation, whereas AlphaMissense‑Optimized alone suggests benign. The variant is most likely pathogenic based on the collective evidence, and this conclusion is not contradicted by the ClinVar “Uncertain” status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | SH3-binding motif | Uncertain | 1 | 6-33442972-A-G | 1 | 6.20e-7 | -7.228 | In-Between | 0.204 | Likely Benign | Likely Benign | 0.243 | Likely Benign | -3.89 | Deleterious | 0.997 | Probably Damaging | 0.934 | Probably Damaging | 2.42 | Pathogenic | 0.01 | Affected | 3.77 | 5 | 0 | -2 | 3.8 | -60.04 | |||||||||||||||||||||||||||
| c.2420A>T | Y807F 2D  AIThe SynGAP1 missense variant Y807F is listed in ClinVar (ID 1491782.0) with an “Uncertain” status and is not reported in gnomAD. All evaluated in‑silico predictors classify the substitution as benign: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all return benign scores, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates a likely benign effect. No tool in the set predicts pathogenicity. High‑accuracy assessments confirm this: AlphaMissense‑Optimized is benign, and the SGM‑Consensus (derived from the four high‑accuracy predictors) is benign. Foldetta, which integrates FoldX‑MD and Rosetta stability calculations, has no available result for this variant. Overall, the computational evidence strongly supports a benign impact, and this conclusion does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | SH3-binding motif | Uncertain | 1 | -3.667 | Likely Benign | 0.073 | Likely Benign | Likely Benign | 0.057 | Likely Benign | 0.14 | Neutral | 0.012 | Benign | 0.022 | Benign | 2.92 | Benign | 0.98 | Tolerated | 3.77 | 5 | 7 | 3 | 4.1 | -16.00 | |||||||||||||||||||||||||||||
| c.2434C>T | P812S 2D  AIThe SynGAP1 missense variant P812S is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33442986‑C‑T). Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv and polyPhen‑2 HumVar. AlphaMissense‑Default remains uncertain, and no Foldetta stability result is available. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta data are missing. Overall, the majority of evidence points to a benign impact, and this conclusion does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | SH3-binding motif | Uncertain | 1 | 6-33442986-C-T | 1 | 6.20e-7 | -5.689 | Likely Benign | 0.456 | Ambiguous | Likely Benign | 0.162 | Likely Benign | -0.62 | Neutral | 0.999 | Probably Damaging | 0.966 | Probably Damaging | 2.89 | Benign | 0.95 | Tolerated | 4.32 | 4 | 1 | -1 | 0.8 | -10.04 | ||||||||||||||||||||||||||
| c.2435C>A | P812H 2D  AIThe SynGAP1 missense variant P812H is listed in ClinVar with an uncertain significance and is present in the gnomAD database (ID 6‑33442987‑C‑A). Prediction tools that agree on a benign effect include REVEL, FATHMM, and AlphaMissense‑Optimized, whereas a majority of tools (PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and AlphaMissense‑Default) predict a pathogenic outcome; ESM1b remains uncertain. High‑accuracy methods give a benign result from AlphaMissense‑Optimized, a pathogenic consensus from the SGM approach (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN), and Foldetta data are unavailable. Overall, the balance of evidence points to a pathogenic effect, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | SH3-binding motif | Uncertain | 2 | 6-33442987-C-A | 3 | 1.86e-6 | -7.470 | In-Between | 0.698 | Likely Pathogenic | Likely Benign | 0.272 | Likely Benign | -2.81 | Deleterious | 1.000 | Probably Damaging | 0.995 | Probably Damaging | 2.68 | Benign | 0.00 | Affected | 4.32 | 4 | 0 | -2 | -1.6 | 40.02 | |||||||||||||||||||||||||||
| c.2443C>G | R815G 2D  AISynGAP1 missense variant R815G is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Prediction tools that agree on benign effect include REVEL and FATHMM, whereas pathogenic predictions come from PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and AlphaMissense‑Default. Uncertain calls are made by ESM1b and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as pathogenic, and Foldetta results are unavailable. Overall, the majority of evidence points to a pathogenic impact, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | SH3-binding motif | Uncertain | 1 | -7.983 | In-Between | 0.854 | Likely Pathogenic | Ambiguous | 0.146 | Likely Benign | -3.22 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | 2.62 | Benign | 0.02 | Affected | 4.32 | 4 | -3 | -2 | 4.1 | -99.14 | ||||||||||||||||||||||||||||||
| c.2443C>T | R815C 2D  AIThe SynGAP1 missense variant R815C is listed in ClinVar (ID 660618.0) with an “Uncertain” status and is present in gnomAD (variant ID 6‑33442995‑C‑T). Prediction tools that agree on a benign effect include REVEL and FATHMM, whereas the majority of tools (PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default) predict a pathogenic impact. The high‑accuracy AlphaMissense‑Optimized result is “Uncertain.” The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is “Likely Pathogenic.” Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the preponderance of predictions indicates a pathogenic effect, which does not contradict the ClinVar “Uncertain” classification but suggests that the variant is more likely pathogenic rather than benign. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | SH3-binding motif | Uncertain | 1 | 6-33442995-C-T | 5 | 3.10e-6 | -9.373 | Likely Pathogenic | 0.828 | Likely Pathogenic | Ambiguous | 0.174 | Likely Benign | -3.89 | Deleterious | 1.000 | Probably Damaging | 0.998 | Probably Damaging | 2.59 | Benign | 0.00 | Affected | 4.32 | 4 | -4 | -3 | 7.0 | -53.05 | ||||||||||||||||||||||||||
| c.2444G>T | R815L 2D  AISynGAP1 missense variant R815L is listed in ClinVar (ID 2505666.0) with an uncertain significance annotation and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL and FATHMM, while pathogenic predictions are made by PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and AlphaMissense‑Default. The high‑accuracy AlphaMissense‑Optimized score is uncertain, and the SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is pathogenic. Foldetta, a protein‑folding stability method that integrates FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the consensus of the majority of tools indicates a pathogenic effect, which contrasts with the ClinVar uncertain classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | SH3-binding motif | Uncertain | 1 | -8.546 | Likely Pathogenic | 0.865 | Likely Pathogenic | Ambiguous | 0.175 | Likely Benign | -3.06 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | 2.63 | Benign | 0.03 | Affected | 4.32 | 4 | -2 | -3 | 8.3 | -43.03 | |||||||||||||||||||||||||||||
| c.2458T>A | Y820N 2D  AIThe SynGAP1 Y820N variant is listed in ClinVar with an “Uncertain” significance and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, and FATHMM, whereas polyPhen‑2 (HumDiv and HumVar), ESM1b, and AlphaMissense‑Default all predict a pathogenic outcome. AlphaMissense‑Optimized returns an “Uncertain” result. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is inconclusive (two benign vs. two pathogenic votes). Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the predictions are evenly split between benign and pathogenic, with no high‑confidence pathogenic or benign signal. Thus, the variant is most likely of uncertain significance, which is consistent with its ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | -9.032 | Likely Pathogenic | 0.842 | Likely Pathogenic | Ambiguous | 0.143 | Likely Benign | -1.53 | Neutral | 0.999 | Probably Damaging | 0.977 | Probably Damaging | 2.74 | Benign | 0.20 | Tolerated | -2 | -2 | -2.2 | -49.07 | |||||||||||||||||||||||||||||||||
| c.2459A>G | Y820C 2D  AIThe SynGAP1 missense variant Y820C is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, SIFT, FATHMM, and AlphaMissense‑Optimized, while those that predict a pathogenic effect are PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and AlphaMissense‑Default. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as “Likely Pathogenic.” High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely pathogenic, and Foldetta results are unavailable. Overall, the balance of evidence—including the SGM‑Consensus—suggests the variant is most likely pathogenic, a conclusion that does not contradict the current ClinVar uncertain classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | Uncertain | 1 | -8.797 | Likely Pathogenic | 0.744 | Likely Pathogenic | Likely Benign | 0.113 | Likely Benign | -3.16 | Deleterious | 1.000 | Probably Damaging | 0.983 | Probably Damaging | 2.68 | Benign | 0.06 | Tolerated | 3.77 | 5 | 0 | -2 | 3.8 | -60.04 | ||||||||||||||||||||||||||||||
| c.2474C>T | S825L 2D  AIThe SynGAP1 missense variant S825L is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443026‑C‑T). Prediction tools that agree on a benign effect include REVEL and ESM1b, whereas the majority of tools (PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, FATHMM, AlphaMissense‑Default) predict a pathogenic impact. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as “Likely Pathogenic.” High‑accuracy assessments show AlphaMissense‑Optimized as “Uncertain,” and the Foldetta stability analysis is unavailable. Overall, the preponderance of evidence points to a pathogenic effect, which is consistent with the ClinVar “Uncertain” classification rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | Uncertain | 1 | 6-33443026-C-T | 1 | 6.20e-7 | -4.987 | Likely Benign | 0.910 | Likely Pathogenic | Ambiguous | 0.249 | Likely Benign | -4.30 | Deleterious | 0.999 | Probably Damaging | 0.994 | Probably Damaging | 1.94 | Pathogenic | 0.01 | Affected | 3.77 | 5 | -2 | -3 | 4.6 | 26.08 | |||||||||||||||||||||||||||
| c.2493G>C | E831D 2D  AIThe SynGAP1 missense variant E831D is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443045‑G‑C). All available in‑silico predictors classify the change as benign: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool reports a pathogenic or likely‑pathogenic outcome. Grouping by agreement, the benign‑predicting tools comprise the entire set, while no pathogenic predictions are present. High‑accuracy assessments reinforce this: AlphaMissense‑Optimized predicts benign; the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also yields a benign classification. Foldetta results are unavailable. Overall, the computational evidence strongly supports a benign effect, and this conclusion does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443045-G-C | 1 | 6.19e-7 | -3.055 | Likely Benign | 0.063 | Likely Benign | Likely Benign | 0.073 | Likely Benign | 1.23 | Neutral | 0.002 | Benign | 0.002 | Benign | 2.64 | Benign | 0.77 | Tolerated | 3.77 | 5 | 3 | 2 | 0.0 | -14.03 | |||||||||||||||||||||||||||
| c.249A>T | R83S 2D  AISynGAP1 R83S is listed in ClinVar (ID 537001.0) with an uncertain significance and is not reported in gnomAD. Prediction tools that classify the variant as benign include REVEL, PROVEAN, ESM1b, FATHMM, and the SGM‑Consensus (Likely Benign). Tools that predict pathogenicity are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, while the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates likely benign; Foldetta results are unavailable. Overall, the predictions are split, with an equal number of benign and pathogenic calls, and the high‑accuracy tools are discordant. Thus, the variant is most likely pathogenic based on the predominance of pathogenic predictions, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -2.550 | Likely Benign | 0.999 | Likely Pathogenic | Likely Pathogenic | 0.094 | Likely Benign | -1.87 | Neutral | 0.909 | Possibly Damaging | 0.587 | Possibly Damaging | 3.19 | Benign | 0.00 | Affected | 4.32 | 1 | 0 | -1 | 3.7 | -69.11 | ||||||||||||||||||||||||||||||
| c.2502G>C | M834I 2D  AIThe SynGAP1 missense variant M834I is listed in ClinVar (ID 3007819.0) with an uncertain significance designation and is not reported in gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict benign. Only SIFT classifies the change as pathogenic. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also reports a likely benign outcome. High‑accuracy assessments confirm this: AlphaMissense‑Optimized is benign, and the SGM‑Consensus (majority‑vote) is likely benign. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, did not provide a result for this variant. Overall, the collective predictions indicate that M834I is most likely benign, which is consistent with its ClinVar uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -3.377 | Likely Benign | 0.291 | Likely Benign | Likely Benign | 0.055 | Likely Benign | -1.21 | Neutral | 0.026 | Benign | 0.009 | Benign | 2.56 | Benign | 0.00 | Affected | 4.32 | 4 | 1 | 2 | 2.6 | -18.03 | ||||||||||||||||||||||||||||||
| c.2506A>G | S836G 2D  AIThe SynGAP1 missense variant S836G is listed in ClinVar (ID 537003.0) with an uncertain significance annotation and is observed in the gnomAD database (variant ID 6‑33443058‑A‑G). Consensus from multiple in silico predictors indicates a benign effect: REVEL, PROVEAN, PolyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the substitution as benign. No tool in the dataset predicts pathogenicity. High‑accuracy assessments corroborate this: AlphaMissense‑Optimized reports a benign outcome, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is labeled Likely Benign. Foldetta, a protein‑folding stability predictor, did not return a result for this variant, so its status is unavailable. Overall, the computational evidence strongly supports a benign classification, which does not conflict with the ClinVar uncertain designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443058-A-G | 4 | 2.48e-6 | -4.749 | Likely Benign | 0.112 | Likely Benign | Likely Benign | 0.066 | Likely Benign | -1.65 | Neutral | 0.006 | Benign | 0.019 | Benign | 2.54 | Benign | 0.39 | Tolerated | 3.77 | 5 | 1 | 0 | 0.4 | -30.03 | |||||||||||||||||||||||||||
| c.250C>G | R84G 2D  AIThe SynGAP1 missense variant R84G is listed in ClinVar with an “Uncertain” significance and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, ESM1b, and FATHMM, whereas those that predict a pathogenic outcome are PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, AlphaMissense‑Default, and AlphaMissense‑Optimized. The SGM Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is inconclusive (two pathogenic versus two benign votes). Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Based on the majority of available predictions, the variant is most likely pathogenic, which does not contradict the current ClinVar status of “Uncertain.” Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | -6.627 | Likely Benign | 0.989 | Likely Pathogenic | Likely Pathogenic | 0.139 | Likely Benign | -2.64 | Deleterious | 0.962 | Probably Damaging | 0.726 | Possibly Damaging | 3.68 | Benign | 0.00 | Affected | 4.32 | 1 | -3 | -2 | 4.1 | -99.14 | |||||||||||||||||||||||||||||||
| c.2514C>A | N838K 2D  AIThe SynGAP1 missense variant N838K is listed in ClinVar with an “Uncertain” status (ClinVar ID 1377909.0) and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, SIFT, and FATHMM, whereas those that predict a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Default. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as “Likely Pathogenic.” High‑accuracy assessments show AlphaMissense‑Optimized as “Uncertain,” SGM‑Consensus as “Likely Pathogenic,” and Foldetta (a protein‑folding stability method combining FoldX‑MD and Rosetta outputs) has no available result for this variant. Overall, the balance of evidence leans toward a pathogenic interpretation, which does not contradict the current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | Uncertain | 2 | -8.470 | Likely Pathogenic | 0.862 | Likely Pathogenic | Ambiguous | 0.097 | Likely Benign | -2.78 | Deleterious | 0.997 | Probably Damaging | 0.995 | Probably Damaging | 2.69 | Benign | 0.16 | Tolerated | 3.77 | 5 | 1 | 0 | -0.4 | 14.07 | ||||||||||||||||||||||||||||||
| c.2518A>T | S840C 2D  AIThe SynGAP1 missense variant S840C is listed in ClinVar (ID 2089808.0) with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include only REVEL, whereas the majority of algorithms—PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, and AlphaMissense‑Default—consistently predict a pathogenic impact. High‑accuracy assessments show AlphaMissense‑Optimized as “Uncertain,” SGM‑Consensus (derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) as “Likely Pathogenic,” and Foldetta results are unavailable. Taken together, the preponderance of evidence points to a pathogenic effect for S840C. This conclusion aligns with the ClinVar designation of uncertainty rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | Uncertain | 1 | -8.799 | Likely Pathogenic | 0.904 | Likely Pathogenic | Ambiguous | 0.376 | Likely Benign | -3.96 | Deleterious | 0.999 | Probably Damaging | 0.975 | Probably Damaging | 1.50 | Pathogenic | 0.00 | Affected | 3.77 | 5 | 0 | -1 | 3.3 | 16.06 | ||||||||||||||||||||||||||||||
| c.2521G>A | V841M 2D  AISynGAP1 variant V841M is listed in ClinVar with an uncertain significance and is present in gnomAD (6-33443073-G-A). Functional prediction tools cluster into two groups: benign predictions from REVEL, PROVEAN, FATHMM, and AlphaMissense‑Optimized; pathogenic predictions from polyPhen‑2 (HumDiv and HumVar), SIFT, and AlphaMissense‑Default. The ESM1b score is inconclusive. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized predicts benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also yields benign, while Foldetta stability analysis is unavailable. Taken together, the majority of evidence, including the high‑accuracy tools, points to a benign effect for V841M. This conclusion does not conflict with the ClinVar uncertain status, which reflects the current lack of definitive clinical data. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | 6-33443073-G-A | 3 | 1.86e-6 | -7.000 | In-Between | 0.651 | Likely Pathogenic | Likely Benign | 0.119 | Likely Benign | -0.74 | Neutral | 0.999 | Probably Damaging | 0.998 | Probably Damaging | 2.54 | Benign | 0.02 | Affected | 3.77 | 5 | 1 | 2 | -2.3 | 32.06 | ||||||||||||||||||||||||||||
| c.2522T>C | V841A 2D  AIThe SynGAP1 missense variant V841A (ClinVar ID 1395978.0) is listed as Uncertain in ClinVar and is present in gnomAD (ID 6‑33443074‑T‑C). Prediction tools that agree on a benign effect include REVEL, PROVEAN, and FATHMM, whereas those that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, and AlphaMissense‑Default. The high‑accuracy AlphaMissense‑Optimized tool reports an uncertain outcome, and the SGM Consensus—derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN—yields a tie (two pathogenic, two benign) and is therefore inconclusive. No Foldetta stability assessment is available for this variant. Overall, the balance of evidence favors a pathogenic interpretation, which does not contradict the current ClinVar designation of Uncertain. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | 6-33443074-T-C | 3 | 1.86e-6 | -8.152 | Likely Pathogenic | 0.901 | Likely Pathogenic | Ambiguous | 0.183 | Likely Benign | -2.13 | Neutral | 0.992 | Probably Damaging | 0.989 | Probably Damaging | 2.57 | Benign | 0.02 | Affected | 3.77 | 5 | 0 | 0 | -2.4 | -28.05 | ||||||||||||||||||||||||||||
| c.2548G>A | G850R 2D  AIThe SynGAP1 missense variant G850R is listed in ClinVar with an uncertain significance (ClinVar ID 2042462.0) and is not reported in gnomAD. Prediction tools that classify the variant as benign include REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote). Only SIFT predicts a pathogenic effect, while AlphaMissense‑Default remains uncertain. High‑accuracy assessments show AlphaMissense‑Optimized labeling the variant as benign and the SGM‑Consensus indicating a likely benign outcome; Foldetta, a protein‑folding stability method, did not provide a result for this substitution. Overall, the preponderance of evidence points to a benign impact, which aligns with the ClinVar designation of uncertain significance rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -5.082 | Likely Benign | 0.398 | Ambiguous | Likely Benign | 0.194 | Likely Benign | -0.07 | Neutral | 0.010 | Benign | 0.010 | Benign | 4.30 | Benign | 0.01 | Affected | 3.77 | 5 | -3 | -2 | -4.1 | 99.14 | ||||||||||||||||||||||||||||||
| c.2557G>C | G853R 2D  AIThe SynGAP1 missense variant G853R is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT. AlphaMissense‑Default is uncertain, and Foldetta (a protein‑folding stability method combining FoldX‑MD and Rosetta outputs) has no available result for this variant. High‑accuracy predictions therefore point to a benign outcome: AlphaMissense‑Optimized is benign, SGM‑Consensus is likely benign, and no Foldetta data is available. Overall, the majority of evidence supports a benign classification, which does not contradict the current ClinVar “Uncertain” status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -4.749 | Likely Benign | 0.366 | Ambiguous | Likely Benign | 0.091 | Likely Benign | -1.27 | Neutral | 0.846 | Possibly Damaging | 0.624 | Possibly Damaging | 4.18 | Benign | 0.00 | Affected | -3 | -2 | -4.1 | 99.14 | ||||||||||||||||||||||||||||||||
| c.2560C>T | R854C 2D  AIThe SynGAP1 missense variant R854C is listed in ClinVar with an “Uncertain” status and is present in gnomAD (gnomAD ID 6‑33443112‑C‑T). Prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized; those that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus also as benign; Foldetta results are not available. Overall, the majority of computational evidence points to a benign impact, which does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443112-C-T | 3 | 1.86e-6 | -5.082 | Likely Benign | 0.170 | Likely Benign | Likely Benign | 0.174 | Likely Benign | -2.48 | Neutral | 1.000 | Probably Damaging | 0.947 | Probably Damaging | 4.05 | Benign | 0.01 | Affected | 3.88 | 3 | -3 | -4 | 7.0 | -53.05 | |||||||||||||||||||||||||||
| c.2561G>A | R854H 2D  AIThe SynGAP1 missense variant R854H is listed in ClinVar with an “Uncertain” status and is present in gnomAD (variant ID 6‑33443113‑G‑A). Prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (which is “Likely Benign”). In contrast, polyPhen‑2 (HumDiv and HumVar) and SIFT all predict a pathogenic impact. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus also as likely benign; the Foldetta stability analysis is unavailable. Overall, the majority of evidence points to a benign effect, and this conclusion does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443113-G-A | 4 | 2.48e-6 | -3.686 | Likely Benign | 0.094 | Likely Benign | Likely Benign | 0.183 | Likely Benign | -1.38 | Neutral | 0.997 | Probably Damaging | 0.899 | Possibly Damaging | 4.07 | Benign | 0.04 | Affected | 3.88 | 3 | 2 | 0 | 1.3 | -19.05 | |||||||||||||||||||||||||||
| c.2567A>G | N856S 2D  AIThe SynGAP1 missense variant N856S is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443119‑A‑G). Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Those that predict a pathogenic outcome are polyPhen‑2 HumDiv and polyPhen‑2 HumVar. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a likely benign classification. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus (majority vote) also benign; Foldetta results are unavailable. Overall, the preponderance of evidence points to a benign effect, which does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443119-A-G | 2 | 1.24e-6 | -2.104 | Likely Benign | 0.064 | Likely Benign | Likely Benign | 0.040 | Likely Benign | -1.54 | Neutral | 0.901 | Possibly Damaging | 0.535 | Possibly Damaging | 4.16 | Benign | 0.30 | Tolerated | 3.88 | 3 | 1 | 1 | 2.7 | -27.03 | |||||||||||||||||||||||||||
| c.256G>A | V86I 2D  AIThe SynGAP1 missense variant V86I is listed in ClinVar (ID 588267.0) with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) all classify the variant as benign or likely benign. Only SIFT predicts a pathogenic outcome. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized is benign, and the SGM‑Consensus is “Likely Benign.” No Foldetta (FoldX‑MD/Rosetta stability) result is available, so it does not influence the assessment. Overall, the majority of predictions indicate that V86I is most likely benign, which is consistent with the ClinVar “Uncertain” classification rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -4.726 | Likely Benign | 0.338 | Likely Benign | Likely Benign | 0.076 | Likely Benign | -0.31 | Neutral | 0.267 | Benign | 0.097 | Benign | 3.94 | Benign | 0.00 | Affected | 4.32 | 1 | 4 | 3 | 0.3 | 14.03 | ||||||||||||||||||||||||||||||
| c.2573G>A | S858N 2D  AIThe SynGAP1 missense variant S858N is listed in ClinVar with an “Uncertain” status and is present in gnomAD (variant ID 6‑33443125‑G‑A). Functional prediction tools largely agree on a benign effect: REVEL, PROVEAN, polyPhen‑2 HumDiv, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) all indicate benign. In contrast, polyPhen‑2 HumVar and SIFT predict pathogenicity, but these two tools are in the minority. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta results are unavailable. Overall, the preponderance of evidence points to a benign impact, and this conclusion does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443125-G-A | 2 | 1.24e-6 | -4.311 | Likely Benign | 0.121 | Likely Benign | Likely Benign | 0.107 | Likely Benign | -0.67 | Neutral | 0.448 | Benign | 0.846 | Possibly Damaging | 4.13 | Benign | 0.02 | Affected | 3.77 | 5 | 1 | 1 | -2.7 | 27.03 | |||||||||||||||||||||||||||
| c.2582C>T | S861L 2D  AIThe SynGAP1 missense variant S861L is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443134‑C‑T). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Benign). Only polyPhen‑2 HumDiv predicts a pathogenic outcome. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized scores the variant as benign, and the SGM‑Consensus (derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates benign. No Foldetta stability prediction is available for this variant. Overall, the computational evidence overwhelmingly points to a benign effect, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443134-C-T | 2 | 1.24e-6 | -4.966 | Likely Benign | 0.219 | Likely Benign | Likely Benign | 0.144 | Likely Benign | -2.10 | Neutral | 0.904 | Possibly Damaging | 0.355 | Benign | 3.93 | Benign | 0.07 | Tolerated | 4.32 | 3 | -3 | -2 | 4.6 | 26.08 | |||||||||||||||||||||||||||
| c.2591C>T | A864V 2D  AIThe SynGAP1 missense variant A864V is listed in ClinVar with an uncertain significance (ClinVar ID 655662.0) and is observed in gnomAD (ID 6‑33443143‑C‑T). Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Benign). Tools that predict a pathogenic outcome are polyPhen‑2 HumDiv and FATHMM. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized is benign, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is benign, and Foldetta results are unavailable. Based on the preponderance of evidence, the variant is most likely benign, which does not contradict its current ClinVar status of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 2 | 6-33443143-C-T | 6 | 3.72e-6 | -4.749 | Likely Benign | 0.126 | Likely Benign | Likely Benign | 0.038 | Likely Benign | -1.35 | Neutral | 0.767 | Possibly Damaging | 0.119 | Benign | 2.45 | Pathogenic | 0.30 | Tolerated | 3.82 | 4 | 0 | 0 | 2.4 | 28.05 | |||||||||||||||||||||||||||
| c.2596G>T | V866L 2D  AIThe SynGAP1 missense variant V866L is listed in ClinVar (ID 469150.0) with an “Uncertain” clinical significance and is present in gnomAD (6‑33443148‑G‑T). All evaluated in‑silico predictors classify the substitution as benign: REVEL, PROVEAN, PolyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool reports a pathogenic outcome. High‑accuracy assessments corroborate this benign prediction: AlphaMissense‑Optimized is benign, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is “Likely Benign.” Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, did not provide a result for this variant. Overall, the computational evidence strongly supports a benign effect, and this conclusion does not contradict the current ClinVar status of uncertainty. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443148-G-T | 1 | 6.20e-7 | -3.352 | Likely Benign | 0.148 | Likely Benign | Likely Benign | 0.046 | Likely Benign | -0.97 | Neutral | 0.217 | Benign | 0.229 | Benign | 2.71 | Benign | 0.21 | Tolerated | 3.82 | 4 | 2 | 1 | -0.4 | 14.03 | |||||||||||||||||||||||||||
| c.2608C>G | L870V 2D  AIThe SynGAP1 missense variant L870V is listed in ClinVar (ID 946946.0) with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). In contrast, PolyPhen‑2 (HumDiv and HumVar) predict a pathogenic outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; the Foldetta protein‑folding stability analysis is unavailable for this variant. Overall, the majority of evidence points to a benign impact, and this conclusion does not conflict with the ClinVar designation of uncertainty. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -4.123 | Likely Benign | 0.300 | Likely Benign | Likely Benign | 0.111 | Likely Benign | -1.19 | Neutral | 0.997 | Probably Damaging | 0.992 | Probably Damaging | 2.64 | Benign | 0.12 | Tolerated | 3.88 | 3 | 2 | 1 | 0.4 | -14.03 | ||||||||||||||||||||||||||||||
| c.2619C>G | S873R 2D  AIThe SynGAP1 missense variant S873R is listed in ClinVar with an uncertain significance and is present in gnomAD (ID 6‑33443171‑C‑G). Functional prediction tools cluster into two groups: benign predictions come from REVEL, SIFT, ESM1b, and FATHMM, while pathogenic predictions arise from PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is inconclusive (two pathogenic versus two benign votes), and Foldetta stability analysis is unavailable. Overall, the balance of evidence favors a pathogenic effect, which contrasts with the ClinVar designation of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | 6-33443171-C-G | 1 | 6.20e-7 | -5.856 | Likely Benign | 0.976 | Likely Pathogenic | Likely Pathogenic | 0.192 | Likely Benign | -2.74 | Deleterious | 0.997 | Probably Damaging | 0.995 | Probably Damaging | 2.67 | Benign | 0.06 | Tolerated | 3.77 | 5 | 0 | -1 | -3.7 | 69.11 | ||||||||||||||||||||||||||||
| c.2623G>A | A875T 2D  AIThe SynGAP1 missense variant A875T is listed in ClinVar with an “Uncertain” status and is present in gnomAD (variant ID 6‑33443175‑G‑A). Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv and polyPhen‑2 HumVar. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as “Likely Benign.” High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the preponderance of evidence points to a benign impact, which is consistent with the ClinVar “Uncertain” classification rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443175-G-A | 1 | 6.20e-7 | -3.793 | Likely Benign | 0.179 | Likely Benign | Likely Benign | 0.110 | Likely Benign | -1.56 | Neutral | 0.972 | Probably Damaging | 0.864 | Possibly Damaging | 2.72 | Benign | 0.26 | Tolerated | 3.77 | 5 | 0 | 1 | -2.5 | 30.03 | |||||||||||||||||||||||||||
| c.2627C>T | S876L 2D  AISynGAP1 variant S876L is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, ESM1b, FATHMM, and AlphaMissense‑Optimized, while pathogenic predictions arise from PROVEAN, polyPhen‑2 (HumDiv and HumVar), and SIFT. AlphaMissense‑Default remains uncertain. High‑accuracy assessments show AlphaMissense‑Optimized predicts a benign effect; the SGM Consensus, derived from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is inconclusive and therefore unavailable; Foldetta stability analysis is also unavailable. Overall, the majority of available predictions favor a benign impact, suggesting the variant is most likely benign. This conclusion does not conflict with the ClinVar uncertain status, which reflects the current lack of definitive evidence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 2 | -5.856 | Likely Benign | 0.489 | Ambiguous | Likely Benign | 0.249 | Likely Benign | -3.56 | Deleterious | 0.998 | Probably Damaging | 0.992 | Probably Damaging | 2.57 | Benign | 0.05 | Affected | 3.77 | 5 | -2 | -3 | 4.6 | 26.08 | |||||||||||||||||||||||||||||||
| c.2632A>G | T878A 2D  AIThe SynGAP1 missense variant T878A is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. All available in‑silico predictors agree on a benign effect: REVEL, PROVEAN, PolyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all return benign scores, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates “Likely Benign.” High‑accuracy tools likewise support a benign outcome: AlphaMissense‑Optimized is benign, and the SGM‑Consensus is benign; Foldetta results are unavailable. Based on the unanimous benign predictions, the variant is most likely benign, and this assessment does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -2.154 | Likely Benign | 0.081 | Likely Benign | Likely Benign | 0.088 | Likely Benign | -0.67 | Neutral | 0.003 | Benign | 0.006 | Benign | 2.73 | Benign | 0.18 | Tolerated | 3.77 | 5 | 1 | 0 | 2.5 | -30.03 | ||||||||||||||||||||||||||||||
| c.263T>C | V88A 2D  AIThe SynGAP1 missense variant V88A is listed in ClinVar (ID 2656486.0) with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and FATHMM. Tools that predict a pathogenic effect are AlphaMissense‑Default, AlphaMissense‑Optimized, and SIFT. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is “Likely Benign.” High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, while the SGM‑Consensus (majority vote) is benign; Foldetta stability analysis is unavailable. Overall, the balance of evidence favors a benign interpretation, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -5.860 | Likely Benign | 0.993 | Likely Pathogenic | Likely Pathogenic | 0.050 | Likely Benign | -1.22 | Neutral | 0.053 | Benign | 0.008 | Benign | 3.75 | Benign | 0.00 | Affected | 4.32 | 1 | 0 | 0 | -2.4 | -28.05 | ||||||||||||||||||||||||||||||
| c.2650C>T | R884W 2D  AIThe SynGAP1 missense variant R884W is listed in ClinVar with an “Uncertain” status and is present in the gnomAD database (gnomAD ID 6‑33443202‑C‑T). Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. In contrast, polyPhen‑2 (HumDiv and HumVar) and SIFT all predict a pathogenic impact. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized is benign, and the SGM‑Consensus is also benign. No Foldetta (protein‑folding stability) result is available for this variant. Overall, the majority of computational predictions, including the high‑accuracy tools, indicate that R884W is most likely benign, which is consistent with the ClinVar “Uncertain” status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443202-C-T | 5 | 3.10e-6 | -3.785 | Likely Benign | 0.332 | Likely Benign | Likely Benign | 0.151 | Likely Benign | 0.26 | Neutral | 0.995 | Probably Damaging | 0.812 | Possibly Damaging | 2.56 | Benign | 0.05 | Affected | 4.32 | 4 | -3 | 2 | 3.6 | 30.03 | |||||||||||||||||||||||||||
| c.2651G>A | R884Q 2D  AIThe SynGAP1 missense variant R884Q is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443203‑G‑A). All available in silico predictors agree on a benign effect: REVEL, PROVEAN, PolyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all return benign scores. No tool predicts pathogenicity. High‑accuracy assessments reinforce this consensus: AlphaMissense‑Optimized is benign, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is “Likely Benign.” Foldetta results are not reported, so they are unavailable. Overall, the computational evidence strongly supports a benign classification, which does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 2 | 6-33443203-G-A | 5 | 3.10e-6 | -3.785 | Likely Benign | 0.128 | Likely Benign | Likely Benign | 0.055 | Likely Benign | -0.42 | Neutral | 0.012 | Benign | 0.004 | Benign | 2.62 | Benign | 0.36 | Tolerated | 4.32 | 4 | 1 | 1 | 1.0 | -28.06 | |||||||||||||||||||||||||||
| c.2657C>T | A886V 2D  AIThe SynGAP1 missense variant A886V is listed in ClinVar with an “Uncertain” status and is present in the gnomAD database (gnomAD ID 6‑33443209‑C‑T). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumVar, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus call (Likely Benign). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, SIFT, and FATHMM. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also as benign, while Foldetta results are unavailable. Overall, the majority of evidence points to a benign impact, and this conclusion does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443209-C-T | 18 | 1.12e-5 | -4.478 | Likely Benign | 0.078 | Likely Benign | Likely Benign | 0.061 | Likely Benign | -0.20 | Neutral | 0.888 | Possibly Damaging | 0.314 | Benign | 2.17 | Pathogenic | 0.00 | Affected | 4.32 | 4 | 0 | 0 | 2.4 | 28.05 | |||||||||||||||||||||||||||
| c.265C>G | P89A 2D  AIThe SynGAP1 missense variant P89A is listed in ClinVar (ID 1031674.0) with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and FATHMM. Tools that predict a pathogenic effect are SIFT and AlphaMissense‑Default. AlphaMissense‑Optimized is uncertain, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) reports a “Likely Benign” outcome. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the majority of high‑confidence predictions indicate a benign impact, and this is consistent with the ClinVar “Uncertain” classification rather than contradicting it. Thus, the variant is most likely benign based on current predictive evidence. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 2 | -5.778 | Likely Benign | 0.920 | Likely Pathogenic | Ambiguous | 0.095 | Likely Benign | -2.47 | Neutral | 0.225 | Benign | 0.020 | Benign | 3.77 | Benign | 0.00 | Affected | 4.32 | 1 | 1 | -1 | 3.4 | -26.04 | ||||||||||||||||||||||||||||||
| c.266C>T | P89L 2D  AISynGAP1 missense variant P89L is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools show a split: benign calls come from REVEL, polyPhen‑2 HumVar, ESM1b, and FATHMM, while pathogenic calls are made by PROVEAN, polyPhen‑2 HumDiv, SIFT, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessments indicate that AlphaMissense‑Optimized predicts pathogenicity, whereas the SGM Consensus (derived from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is inconclusive due to a 2‑to‑2 split, and Foldetta results are unavailable. Overall, the majority of tools favor a pathogenic effect, but the evidence is not unanimous. Therefore, the variant is most likely pathogenic according to the current predictions, and this assessment does not contradict its ClinVar status of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 2 | -6.775 | Likely Benign | 0.982 | Likely Pathogenic | Likely Pathogenic | 0.119 | Likely Benign | -3.29 | Deleterious | 0.889 | Possibly Damaging | 0.058 | Benign | 3.73 | Benign | 0.00 | Affected | 4.32 | 1 | -3 | -3 | 5.4 | 16.04 | |||||||||||||||||||||||||||||||
| c.2681G>A | G894E 2D  AIThe SynGAP1 missense variant G894E is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443233‑G‑A). Prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and AlphaMissense‑Default. AlphaMissense‑Optimized is reported as uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, SGM‑Consensus as likely benign, and no Foldetta (FoldX‑MD/Rosetta) result is available. Overall, the majority of predictions support a benign impact, and this is consistent with the ClinVar “Uncertain” classification; thus the variant is most likely benign and does not contradict the current ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443233-G-A | 6 | 3.72e-6 | -5.377 | Likely Benign | 0.859 | Likely Pathogenic | Ambiguous | 0.180 | Likely Benign | -2.07 | Neutral | 1.000 | Probably Damaging | 1.000 | Probably Damaging | 2.68 | Benign | 0.01 | Affected | 4.32 | 4 | 0 | -2 | -3.1 | 72.06 | |||||||||||||||||||||||||||
| c.2684G>A | S895N 2D  AIThe SynGAP1 missense variant S895N is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and AlphaMissense‑Default. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is “Likely Benign.” High‑accuracy assessments show AlphaMissense‑Optimized as benign, and the SGM‑Consensus (majority vote) also leans benign. No Foldetta (protein‑folding stability) result is available, so it does not influence the assessment. Overall, the preponderance of predictions indicates the variant is most likely benign, which is consistent with its ClinVar “Uncertain” classification rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -6.399 | Likely Benign | 0.604 | Likely Pathogenic | Likely Benign | 0.118 | Likely Benign | -0.85 | Neutral | 0.991 | Probably Damaging | 0.988 | Probably Damaging | 2.64 | Benign | 0.30 | Tolerated | 4.32 | 4 | 1 | 1 | -2.7 | 27.03 | ||||||||||||||||||||||||||||||
| c.2690C>T | S897L 2D  AIThe SynGAP1 missense variant S897L is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). In contrast, polyPhen‑2 (HumDiv and HumVar) and SIFT predict a pathogenic impact. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus (derived from the same four high‑accuracy tools) also as benign. No Foldetta stability prediction is available for this variant. Overall, the majority of computational evidence points to a benign effect, and this conclusion does not conflict with the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -4.034 | Likely Benign | 0.299 | Likely Benign | Likely Benign | 0.028 | Likely Benign | -1.71 | Neutral | 0.901 | Possibly Damaging | 0.636 | Possibly Damaging | 2.66 | Benign | 0.01 | Affected | -3 | -2 | 4.6 | 26.08 | ||||||||||||||||||||||||||||||||
| c.269T>A | V90E 2D  AIThe SynGAP1 missense variant V90E is listed in ClinVar (ID 971665.0) with an uncertain significance status and is not reported in gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, and AlphaMissense‑Optimized all predict benign. Only two tools—SIFT and AlphaMissense‑Default—suggest a pathogenic outcome. When the high‑accuracy consensus is considered, AlphaMissense‑Optimized remains benign, and the SGM Consensus (derived from the majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates a likely benign effect. No Foldetta stability assessment is available for this variant. Overall, the preponderance of evidence points to a benign impact, which does not contradict the ClinVar uncertain classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -4.079 | Likely Benign | 0.703 | Likely Pathogenic | Likely Benign | 0.108 | Likely Benign | -0.38 | Neutral | 0.001 | Benign | 0.000 | Benign | 4.00 | Benign | 0.00 | Affected | 4.32 | 1 | -2 | -2 | -7.7 | 29.98 | ||||||||||||||||||||||||||||||
| c.2702C>T | A901V 2D  AIThe SynGAP1 missense variant A901V is listed in ClinVar (ID 934469.0) with an “Uncertain” clinical significance and is present in the gnomAD database (gnomAD ID 6‑33443254‑C‑T). All evaluated in‑silico predictors classify the substitution as benign: REVEL, PROVEAN, PolyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool reports a pathogenic or likely pathogenic outcome. High‑accuracy assessments corroborate this benign prediction: AlphaMissense‑Optimized is benign, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is “Likely Benign.” Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, did not provide a result for this variant. Overall, the computational evidence strongly supports a benign effect, which does not contradict the ClinVar “Uncertain” status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 2 | 6-33443254-C-T | 2 | 1.24e-6 | -5.043 | Likely Benign | 0.219 | Likely Benign | Likely Benign | 0.029 | Likely Benign | -1.83 | Neutral | 0.106 | Benign | 0.009 | Benign | 2.64 | Benign | 0.17 | Tolerated | 3.77 | 5 | 0 | 0 | 2.4 | 28.05 | |||||||||||||||||||||||||||
| c.2711T>C | M904T 2D  AIThe SynGAP1 missense variant M904T is listed in ClinVar (ID 1311496.0) with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Only AlphaMissense‑Default predicts a pathogenic outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; a Foldetta stability analysis is unavailable. Overall, the majority of evidence points to a benign impact, and this conclusion does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -2.721 | Likely Benign | 0.668 | Likely Pathogenic | Likely Benign | 0.042 | Likely Benign | -1.15 | Neutral | 0.277 | Benign | 0.103 | Benign | 2.78 | Benign | 0.18 | Tolerated | 3.77 | 5 | -1 | -1 | -2.6 | -30.09 | ||||||||||||||||||||||||||||||
| c.2714G>A | R905H 2D  AIThe SynGAP1 missense variant R905H is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443266‑G‑A). Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Tools that predict a pathogenic outcome are polyPhen‑2 HumDiv and polyPhen‑2 HumVar. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta’s protein‑folding stability analysis is unavailable for this variant. Overall, the majority of computational evidence points to a benign effect, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443266-G-A | 8 | 4.96e-6 | -4.182 | Likely Benign | 0.457 | Ambiguous | Likely Benign | 0.192 | Likely Benign | -1.11 | Neutral | 1.000 | Probably Damaging | 0.991 | Probably Damaging | 2.59 | Benign | 0.09 | Tolerated | 3.77 | 5 | 2 | 0 | 1.3 | -19.05 | |||||||||||||||||||||||||||
| c.2729G>C | G910A 2D  AIThe SynGAP1 missense variant G910A is listed in ClinVar with an “Uncertain” status (ClinVar ID 2091237.0) and is present in gnomAD (6‑33443281‑G‑C). Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are PolyPhen‑2 HumDiv and PolyPhen‑2 HumVar. The remaining predictions are uncertain: AlphaMissense‑Default is inconclusive, while the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) reports a likely benign outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely benign, and Foldetta results are unavailable. Overall, the majority of evidence points to a benign impact, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443281-G-C | 1 | 6.20e-7 | -3.587 | Likely Benign | 0.361 | Ambiguous | Likely Benign | 0.209 | Likely Benign | -1.43 | Neutral | 0.999 | Probably Damaging | 0.999 | Probably Damaging | 2.78 | Benign | 0.10 | Tolerated | 3.77 | 5 | 1 | 0 | 2.2 | 14.03 | |||||||||||||||||||||||||||
| c.2735C>A | T912N 2D  AIThe SynGAP1 missense variant T912N is listed in ClinVar with an uncertain significance (ClinVar ID 2337624.0) and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. In contrast, polyPhen‑2 (HumDiv and HumVar) and SIFT all predict a pathogenic impact. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as likely benign. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized predicts benign, and the SGM‑Consensus also indicates likely benign. No Foldetta (FoldX‑MD/Rosetta) stability result is available, so it does not influence the overall assessment. Based on the preponderance of evidence, the variant is most likely benign, and this conclusion does not contradict the current ClinVar status of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -4.260 | Likely Benign | 0.190 | Likely Benign | Likely Benign | 0.116 | Likely Benign | -1.15 | Neutral | 0.999 | Probably Damaging | 0.977 | Probably Damaging | 3.96 | Benign | 0.00 | Affected | 3.77 | 5 | 0 | 0 | -2.8 | 13.00 | ||||||||||||||||||||||||||||||
| c.2741A>T | D914V 2D  AIThe SynGAP1 missense variant D914V is listed in ClinVar (ID 2582846.0) with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic outcome are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and AlphaMissense‑Default. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is “Likely Benign.” High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus (majority vote) as benign; Foldetta results are unavailable. Overall, the balance of evidence points to a benign effect, and this conclusion does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -4.260 | Likely Benign | 0.723 | Likely Pathogenic | Likely Benign | 0.187 | Likely Benign | -2.24 | Neutral | 0.999 | Probably Damaging | 0.986 | Probably Damaging | 2.64 | Benign | 0.01 | Affected | 3.77 | 5 | -3 | -2 | 7.7 | -15.96 | ||||||||||||||||||||||||||||||
| c.2750C>G | P917R 2D  AIThe SynGAP1 missense variant P917R is listed in ClinVar with an “Uncertain” status and is present in gnomAD (gnomAD ID 6‑33443302‑C‑G). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv and SIFT; AlphaMissense‑Default remains uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus (derived from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also as benign, while Foldetta results are unavailable. Overall, the majority of evidence points to a benign impact, and this conclusion does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443302-C-G | 5 | 3.10e-6 | -4.475 | Likely Benign | 0.363 | Ambiguous | Likely Benign | 0.142 | Likely Benign | -1.70 | Neutral | 0.642 | Possibly Damaging | 0.316 | Benign | 2.68 | Benign | 0.00 | Affected | 3.77 | 5 | -2 | 0 | -2.9 | 59.07 | |||||||||||||||||||||||||||
| c.2752G>A | A918T 2D  AISynGAP1 missense variant A918T is listed in ClinVar with an uncertain significance (ClinVar ID 3964538.0) and is present in gnomAD (6‑33443304‑G‑A). Functional prediction tools cluster into two groups: benign predictions come from REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Benign). Pathogenic predictions are reported by polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT. High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized predicts benign, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates likely benign. Foldetta stability analysis is unavailable. Overall, the preponderance of evidence points to a benign effect, which is consistent with the ClinVar uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443304-G-A | 1 | 6.20e-7 | -4.139 | Likely Benign | 0.083 | Likely Benign | Likely Benign | 0.065 | Likely Benign | -1.09 | Neutral | 0.980 | Probably Damaging | 0.721 | Possibly Damaging | 2.64 | Benign | 0.03 | Affected | 4.32 | 4 | 0 | 1 | -2.5 | 30.03 | |||||||||||||||||||||||||||
| c.2753C>T | A918V 2D  AIThe SynGAP1 missense variant A918V is listed in ClinVar with an “Uncertain” status and is present in gnomAD (gnomAD ID 6‑33443305‑C‑T). Prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Benign). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as Likely Benign; a Foldetta stability prediction is not available. Overall, the majority of evidence points to a benign impact, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 3 | 6-33443305-C-T | 2 | 1.24e-6 | -3.684 | Likely Benign | 0.112 | Likely Benign | Likely Benign | 0.119 | Likely Benign | -1.61 | Neutral | 0.980 | Probably Damaging | 0.782 | Possibly Damaging | 2.61 | Benign | 0.03 | Affected | 4.32 | 4 | 0 | 0 | 2.4 | 28.05 | |||||||||||||||||||||||||||
| c.2768T>A | I923N 2D  AIThe SynGAP1 missense variant I923N (ClinVar ID 647043.0) is listed as “Uncertain” and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and AlphaMissense‑Default. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is “Likely Benign” because the majority of its constituent tools (three benign, one pathogenic) favor a benign outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as benign, and the Foldetta stability analysis is unavailable. Overall, the collective predictions point to a benign impact, which does not contradict the ClinVar “Uncertain” status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -0.733 | Likely Benign | 0.712 | Likely Pathogenic | Likely Benign | 0.108 | Likely Benign | -1.16 | Neutral | 0.991 | Probably Damaging | 0.793 | Possibly Damaging | 2.70 | Benign | 0.13 | Tolerated | 3.77 | 5 | -2 | -3 | -8.0 | 0.94 | ||||||||||||||||||||||||||||||
| c.277C>G | R93G 2D  AIThe SynGAP1 missense variant R93G is listed in ClinVar (ID 2504251.0) with an “Uncertain” clinical significance and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Only SIFT predicts a pathogenic outcome, while AlphaMissense‑Default remains uncertain. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized scores the variant as benign, and the SGM‑Consensus (derived from the same set of predictors) labels it “Likely Benign”; Foldetta results are unavailable. Overall, the preponderance of evidence indicates that R93G is most likely benign, which does not contradict the current ClinVar status of uncertainty. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -2.674 | Likely Benign | 0.400 | Ambiguous | Likely Benign | 0.093 | Likely Benign | -1.69 | Neutral | 0.103 | Benign | 0.019 | Benign | 3.99 | Benign | 0.00 | Affected | 4.32 | 1 | -2 | -3 | 4.1 | -99.14 | ||||||||||||||||||||||||||||||
| c.2809G>C | D937H 2D  AIThe SynGAP1 D937H missense variant (ClinVar ID 2825773.0) is listed as “Uncertain” and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic outcome are PolyPhen‑2 HumDiv, PolyPhen‑2 HumVar, and AlphaMissense‑Default. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as “Likely Benign.” High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized predicts benign, the SGM‑Consensus (majority vote) is benign, and Foldetta (protein‑folding stability analysis combining FoldX‑MD and Rosetta) data are unavailable. Based on the preponderance of evidence from both general and high‑accuracy predictors, the variant is most likely benign, which is consistent with its ClinVar “Uncertain” status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -0.733 | Likely Benign | 0.677 | Likely Pathogenic | Likely Benign | 0.150 | Likely Benign | -1.74 | Neutral | 1.000 | Probably Damaging | 0.975 | Probably Damaging | 2.68 | Benign | 0.13 | Tolerated | 3.77 | 5 | -1 | 1 | 0.3 | 22.05 | ||||||||||||||||||||||||||||||
| c.2812G>A | G938R 2D  AIThe SynGAP1 missense variant G938R is listed in ClinVar (ID 1019898.0) with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Benign). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as Likely Benign; Foldetta results are unavailable. Overall, the majority of evidence (seven benign versus three pathogenic predictions) supports a benign classification. This consensus does not contradict the ClinVar “Uncertain” designation, which remains unresolved. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -5.271 | Likely Benign | 0.732 | Likely Pathogenic | Likely Benign | 0.141 | Likely Benign | -1.11 | Neutral | 0.999 | Probably Damaging | 0.985 | Probably Damaging | 2.74 | Benign | 0.36 | Tolerated | 3.77 | 5 | -3 | -2 | -4.1 | 99.14 | ||||||||||||||||||||||||||||||
| c.2818G>A | G940S 2D  AIThe SynGAP1 missense variant G940S is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443370‑G‑A). All available in silico predictors agree on a benign effect: REVEL, PROVEAN, PolyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all return benign scores, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is “Likely Benign.” High‑accuracy tools reinforce this view: AlphaMissense‑Optimized predicts benign, and the SGM‑Consensus likewise indicates a benign outcome. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta, has no reported result for this variant, so its status is unavailable. Overall, the computational evidence strongly supports a benign classification, which does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443370-G-A | 1 | 6.20e-7 | -5.451 | Likely Benign | 0.084 | Likely Benign | Likely Benign | 0.135 | Likely Benign | 0.45 | Neutral | 0.409 | Benign | 0.253 | Benign | 2.77 | Benign | 0.44 | Tolerated | 3.77 | 5 | 1 | 0 | -0.4 | 30.03 | |||||||||||||||||||||||||||
| c.2822C>T | P941L 2D  AIThe SynGAP1 missense variant P941L is listed in ClinVar (ID 3451960.0) with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). The only tool that predicts a pathogenic outcome is SIFT. High‑accuracy assessments further support a benign classification: AlphaMissense‑Optimized predicts benign, and the SGM‑Consensus also indicates “Likely Benign.” No Foldetta (FoldX‑MD/Rosetta) stability result is available, so it does not influence the assessment. Overall, the majority of predictions, including the high‑accuracy tools, point to a benign effect, which is consistent with the ClinVar “Uncertain” designation rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 2 | -5.692 | Likely Benign | 0.066 | Likely Benign | Likely Benign | 0.054 | Likely Benign | -0.44 | Neutral | 0.144 | Benign | 0.039 | Benign | 2.76 | Benign | 0.01 | Affected | -3 | -3 | 5.4 | 16.04 | ||||||||||||||||||||||||||||||||
| c.2825C>T | P942L 2D  AIThe SynGAP1 missense variant P942L is listed in ClinVar (ID 2851884.0) with an “Uncertain” status and is present in gnomAD (variant ID 6‑33443377‑C‑T). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are SIFT and FATHMM. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta results are unavailable. Overall, the majority of evidence points to a benign impact, and this is consistent with the ClinVar “Uncertain” designation rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443377-C-T | 4 | 2.48e-6 | -5.063 | Likely Benign | 0.086 | Likely Benign | Likely Benign | 0.048 | Likely Benign | -2.00 | Neutral | 0.411 | Benign | 0.239 | Benign | 2.37 | Pathogenic | 0.00 | Affected | 4.32 | 4 | -3 | -3 | 5.4 | 16.04 | |||||||||||||||||||||||||||
| c.2855G>T | G952V 2D  AIThe SynGAP1 missense variant G952V is listed in ClinVar (ID 2055482.0) with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Only SIFT predicts a pathogenic outcome, while ESM1b remains uncertain. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as “Likely Benign.” High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized is benign, and the SGM‑Consensus is likely benign; Foldetta results are unavailable. Overall, the preponderance of evidence indicates that G952V is most likely benign, and this conclusion does not contradict the current ClinVar status of uncertainty. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -7.074 | In-Between | 0.078 | Likely Benign | Likely Benign | 0.231 | Likely Benign | -0.33 | Neutral | 0.000 | Benign | 0.000 | Benign | 3.20 | Benign | 0.02 | Affected | 3.77 | 5 | -1 | -3 | 4.6 | 42.08 | ||||||||||||||||||||||||||||||
| c.2858C>A | P953Q 2D  AIThe SynGAP1 missense variant P953Q is listed in ClinVar with an uncertain significance (ClinVar ID 1176820.0) and is not found in gnomAD. Functional prediction tools uniformly classify the substitution as benign: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all report benign effects, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates a likely benign outcome. High‑accuracy assessments corroborate this: AlphaMissense‑Optimized predicts benign, and the SGM‑Consensus also reports likely benign. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the consensus of all available predictions points to a benign impact, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -6.038 | Likely Benign | 0.079 | Likely Benign | Likely Benign | 0.086 | Likely Benign | -0.78 | Neutral | 0.058 | Benign | 0.015 | Benign | 2.78 | Benign | 0.29 | Tolerated | 3.77 | 5 | 0 | -1 | -1.9 | 31.01 | ||||||||||||||||||||||||||||||
| c.2863T>C | S955P 2D  AIThe SynGAP1 missense variant S955P is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443415‑T‑C). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). Tools that predict a pathogenic effect are SIFT and FATHMM. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta results are unavailable. Overall, the majority of evidence points to a benign impact, and this is consistent with the ClinVar “Uncertain” classification rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443415-T-C | 3 | 1.86e-6 | -2.584 | Likely Benign | 0.073 | Likely Benign | Likely Benign | 0.098 | Likely Benign | -0.75 | Neutral | 0.001 | Benign | 0.004 | Benign | 2.33 | Pathogenic | 0.00 | Affected | 3.77 | 5 | 1 | -1 | -0.8 | 10.04 | |||||||||||||||||||||||||||
| c.286G>A | G96S 2D  AIThe SynGAP1 missense variant G96S is listed in ClinVar with an “Uncertain” status and is present in gnomAD (variant ID 6‑33425894‑G‑A). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). Only SIFT predicts a pathogenic outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta results are not available. Overall, the majority of evidence points to a benign impact, and this does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33425894-G-A | 5 | 3.10e-6 | -3.049 | Likely Benign | 0.065 | Likely Benign | Likely Benign | 0.071 | Likely Benign | -0.76 | Neutral | 0.364 | Benign | 0.008 | Benign | 4.25 | Benign | 0.00 | Affected | 4.32 | 1 | 1 | 0 | -0.4 | 30.03 | |||||||||||||||||||||||||||
| c.2888A>G | H963R 2D  AIThe SynGAP1 missense variant H963R is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443440‑A‑G). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus score (Likely Benign). Only ESM1b predicts a pathogenic outcome. High‑accuracy assessments further support a benign classification: AlphaMissense‑Optimized reports benign, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) yields a benign consensus. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no reported result for this variant, so its status is unavailable. Overall, the preponderance of evidence indicates the variant is most likely benign, which does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443440-A-G | 8 | 4.96e-6 | -8.952 | Likely Pathogenic | 0.169 | Likely Benign | Likely Benign | 0.081 | Likely Benign | -1.28 | Neutral | 0.001 | Benign | 0.003 | Benign | 4.15 | Benign | 0.24 | Tolerated | 3.77 | 5 | 2 | 0 | -1.3 | 19.05 | |||||||||||||||||||||||||||
| c.28C>T | R10W 2D  AIThe SynGAP1 R10W missense variant is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33420292‑C‑T). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (which itself is “Likely Benign”). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv and SIFT; AlphaMissense‑Default remains uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as benign, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also indicates benign. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the majority of evidence points to a benign impact, and this conclusion does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33420292-C-T | 2 | 1.30e-6 | -5.707 | Likely Benign | 0.503 | Ambiguous | Likely Benign | 0.236 | Likely Benign | -0.31 | Neutral | 0.964 | Probably Damaging | 0.190 | Benign | 4.10 | Benign | 0.00 | Affected | 4.32 | 1 | 2 | -3 | 3.6 | 30.03 | |||||||||||||||||||||||||||
| c.2900G>T | R967L 2D  AIThe SynGAP1 missense variant R967L is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443452‑G‑T). Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). In contrast, polyPhen‑2 (both HumDiv and HumVar models) predict a pathogenic outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta results are unavailable. Overall, the preponderance of evidence points to a benign impact for R967L, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443452-G-T | 1 | 6.20e-7 | -3.496 | Likely Benign | 0.164 | Likely Benign | Likely Benign | 0.123 | Likely Benign | -0.99 | Neutral | 0.959 | Probably Damaging | 0.586 | Possibly Damaging | 4.15 | Benign | 0.75 | Tolerated | 4.32 | 2 | -2 | -3 | 8.3 | -43.03 | |||||||||||||||||||||||||||
| c.2912C>A | P971H 2D  AIThe SynGAP1 missense variant P971H is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443464‑C‑A). Prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Benign). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as Likely Benign; a Foldetta stability prediction is not available. Overall, the majority of evidence points to a benign impact, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443464-C-A | 1 | 6.20e-7 | -5.243 | Likely Benign | 0.086 | Likely Benign | Likely Benign | 0.039 | Likely Benign | -1.11 | Neutral | 0.898 | Possibly Damaging | 0.477 | Possibly Damaging | 3.89 | Benign | 0.00 | Affected | 4.32 | 2 | -2 | 0 | -1.6 | 40.02 | |||||||||||||||||||||||||||
| c.2914C>G | P972A 2D  AIThe SynGAP1 missense variant P972A is listed in ClinVar with an uncertain significance (ClinVar ID 3172763.0) and is present in the gnomAD database (gnomAD ID 6‑33443466‑C‑G). All evaluated in‑silico predictors classify the substitution as benign: REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool predicts pathogenicity. High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized reports benign, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates “Likely Benign.” The Foldetta protein‑folding stability analysis is unavailable for this variant. Overall, the computational evidence strongly suggests the variant is most likely benign, and this conclusion does not contradict the current ClinVar status of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443466-C-G | 1 | 6.20e-7 | -0.167 | Likely Benign | 0.045 | Likely Benign | Likely Benign | 0.046 | Likely Benign | -0.89 | Neutral | 0.016 | Benign | 0.011 | Benign | 4.29 | Benign | 0.07 | Tolerated | 4.32 | 2 | -1 | 1 | 3.4 | -26.04 | |||||||||||||||||||||||||||
| c.2914C>T | P972S 2D  AIThe SynGAP1 missense variant P972S (ClinVar ID 3361353.0) is listed as Uncertain in ClinVar and is present in gnomAD (ID 6‑33443466‑C‑T). Consensus among most in silico predictors indicates a benign effect: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all score the substitution as benign. Only SIFT classifies it as pathogenic, representing the sole discordant prediction. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized predicts benign, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) reports “Likely Benign.” No Foldetta stability analysis is available for this residue. Overall, the preponderance of computational evidence points to a benign effect, which is consistent with the ClinVar designation of uncertainty rather than pathogenicity. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443466-C-T | 4 | 2.48e-6 | -4.008 | Likely Benign | 0.058 | Likely Benign | Likely Benign | 0.074 | Likely Benign | -0.38 | Neutral | 0.001 | Benign | 0.002 | Benign | 4.28 | Benign | 0.05 | Affected | 4.32 | 2 | -1 | 1 | 0.8 | -10.04 | |||||||||||||||||||||||||||
| c.291G>T | E97D 2D  AIThe SynGAP1 missense variant E97D is listed in ClinVar (ID 1313570.0) with an “Uncertain” clinical significance and is present in gnomAD (variant ID 6‑33425899‑G‑T). Functional prediction tools cluster into two groups: benign predictions include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Benign). Pathogenic predictions come from polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as Likely Benign; Foldetta results are unavailable. Overall, the majority of evidence points to a benign effect. This consensus does not contradict the ClinVar “Uncertain” status, which remains unresolved. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 3 | 6-33425899-G-T | -3.239 | Likely Benign | 0.077 | Likely Benign | Likely Benign | 0.081 | Likely Benign | -0.49 | Neutral | 0.880 | Possibly Damaging | 0.636 | Possibly Damaging | 4.12 | Benign | 0.00 | Affected | 4.32 | 1 | 3 | 2 | 0.0 | -14.03 | |||||||||||||||||||||||||||||
| c.2924C>A | T975N 2D  AIThe SynGAP1 missense variant T975N is listed in ClinVar (ID 942242.0) with an “Uncertain” clinical significance and is present in gnomAD (variant ID 6‑33443476‑C‑A). Functional prediction tools largely agree on a benign effect: REVEL, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all report benign or tolerated outcomes. Only polyPhen‑2 HumDiv predicts a pathogenic effect. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as “Likely Benign.” High‑accuracy assessments reinforce this view: AlphaMissense‑Optimized predicts benign, and the SGM‑Consensus (majority vote) also indicates benign. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the preponderance of evidence points to a benign impact, which is consistent with the ClinVar “Uncertain” status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443476-C-A | 1 | 6.20e-7 | -4.671 | Likely Benign | 0.089 | Likely Benign | Likely Benign | 0.100 | Likely Benign | -0.58 | Neutral | 0.586 | Possibly Damaging | 0.302 | Benign | 4.13 | Benign | 0.07 | Tolerated | 4.32 | 2 | 0 | 0 | -2.8 | 13.00 | |||||||||||||||||||||||||||
| c.2924C>G | T975S 2D  AIThe SynGAP1 missense variant T975S is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool in the dataset predicts a pathogenic outcome. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized scores the variant as benign, and the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates “Likely Benign.” Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, did not provide a result for this variant. Overall, the available predictions strongly suggest that T975S is most likely benign, and this conclusion does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -2.743 | Likely Benign | 0.068 | Likely Benign | Likely Benign | 0.109 | Likely Benign | -0.57 | Neutral | 0.059 | Benign | 0.061 | Benign | 4.16 | Benign | 0.20 | Tolerated | 1 | 1 | -0.1 | -14.03 | ||||||||||||||||||||||||||||||||
| c.2924C>T | T975I 2D  AIThe SynGAP1 missense variant T975I is listed in ClinVar with an “Uncertain” status and is present in the gnomAD database (ID 6‑33443476‑C‑T). Prediction tools that agree on benign impact include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized; no tool predicts pathogenicity. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also reports a “Likely Benign” classification. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign, while Foldetta (combining FoldX‑MD and Rosetta outputs) has no available result for this variant. Overall, the computational evidence overwhelmingly supports a benign effect, which does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443476-C-T | 6 | 3.72e-6 | -3.912 | Likely Benign | 0.164 | Likely Benign | Likely Benign | 0.068 | Likely Benign | -1.66 | Neutral | 0.411 | Benign | 0.239 | Benign | 4.11 | Benign | 0.66 | Tolerated | 4.32 | 2 | 0 | -1 | 5.2 | 12.05 | |||||||||||||||||||||||||||
| c.2928T>G | F976L 2D  AIThe SynGAP1 missense variant F976L is listed in ClinVar (ID 624245.0) with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and FATHMM. Only AlphaMissense‑Default predicts a pathogenic outcome. The high‑accuracy consensus predictor SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. AlphaMissense‑Optimized is inconclusive, and no Foldetta (FoldX‑MD/Rosetta stability) result is available. Overall, the majority of evidence points to a benign impact, and this is consistent with the ClinVar designation of uncertainty rather than a definitive pathogenic claim. Thus, the variant is most likely benign, and its predictions do not contradict the current ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -2.432 | Likely Benign | 0.825 | Likely Pathogenic | Ambiguous | 0.212 | Likely Benign | -0.87 | Neutral | 0.264 | Benign | 0.102 | Benign | 4.20 | Benign | 0.53 | Tolerated | 4.32 | 2 | 2 | 0 | 1.0 | -34.02 | ||||||||||||||||||||||||||||||
| c.2932C>T | P978S 2D  AIThe SynGAP1 missense variant P978S is listed in ClinVar (ID 3379672.0) with an “Uncertain” clinical significance and is not reported in gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict benign. Only polyPhen‑2 HumDiv indicates a pathogenic outcome. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as “Likely Benign.” High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized is benign, and the SGM‑Consensus is likely benign. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the preponderance of predictions points to a benign effect, which is consistent with the ClinVar “Uncertain” status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -3.913 | Likely Benign | 0.151 | Likely Benign | Likely Benign | 0.085 | Likely Benign | -1.07 | Neutral | 0.481 | Possibly Damaging | 0.220 | Benign | 4.22 | Benign | 0.48 | Tolerated | 1 | -1 | 0.8 | -10.04 | ||||||||||||||||||||||||||||||||
| c.2935T>C | F979L 2D  AIThe SynGAP1 missense variant F979L (ClinVar ID 1000410.0, status Uncertain, not found in gnomAD) has been evaluated by multiple in silico predictors. Benign predictions come from REVEL, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, and FATHMM, while pathogenic predictions are reported by polyPhen‑2 HumDiv and AlphaMissense‑Default. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is classified as Likely Benign. High‑accuracy assessments show AlphaMissense‑Optimized as Uncertain, whereas the SGM‑Consensus (majority vote) supports a benign outcome. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta, has no available result for this variant. Overall, the majority of evidence points to a benign effect, and this conclusion does not contradict the ClinVar status of Uncertain. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -2.341 | Likely Benign | 0.870 | Likely Pathogenic | Ambiguous | 0.228 | Likely Benign | -1.00 | Neutral | 0.625 | Possibly Damaging | 0.430 | Benign | 4.22 | Benign | 0.73 | Tolerated | 4.32 | 2 | 2 | 0 | 1.0 | -34.02 | ||||||||||||||||||||||||||||||
| c.2954G>A | S985N 2D  AIThe SynGAP1 missense variant S985N is listed in ClinVar (ID 2087879.0) with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, and FATHMM, while those that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and AlphaMissense‑Default. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. Separately, the high‑accuracy AlphaMissense‑Optimized result is “Uncertain,” and the Foldetta protein‑folding stability assessment is unavailable. Based on the overall distribution of predictions, the variant is most likely benign; this conclusion does not contradict the ClinVar status, which remains uncertain. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -6.979 | Likely Benign | 0.845 | Likely Pathogenic | Ambiguous | 0.088 | Likely Benign | -1.68 | Neutral | 0.991 | Probably Damaging | 0.988 | Probably Damaging | 2.65 | Benign | 0.00 | Affected | 4.32 | 1 | 1 | 1 | -2.7 | 27.03 | ||||||||||||||||||||||||||||||
| c.2960A>G | D987G 2D  AIThe SynGAP1 missense variant D987G (ClinVar ID 1061058.0) is listed as ClinVar status Uncertain and is not reported in gnomAD. Functional prediction tools show a split: benign predictions come from REVEL, SIFT, and ESM1b, whereas pathogenic predictions are reported by PROVEAN, polyPhen‑2 (HumDiv and HumVar), FATHMM, and AlphaMissense‑Default. The SGM‑Consensus, which aggregates majority votes from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, classifies the variant as Likely Pathogenic. High‑accuracy assessments further indicate that AlphaMissense‑Optimized is uncertain, while Foldetta data are unavailable. Overall, the majority of evidence points toward a pathogenic effect, aligning with the SGM‑Consensus but contradicting the ClinVar Uncertain designation. Therefore, the variant is most likely pathogenic based on current predictions, and this assessment is in conflict with the ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | Uncertain | 1 | -4.782 | Likely Benign | 0.849 | Likely Pathogenic | Ambiguous | 0.234 | Likely Benign | -2.79 | Deleterious | 0.943 | Possibly Damaging | 0.808 | Possibly Damaging | 2.45 | Pathogenic | 0.07 | Tolerated | 4.32 | 2 | 1 | -1 | 3.1 | -58.04 | ||||||||||||||||||||||||||||||
| c.2962C>T | L988F 2D  AIThe SynGAP1 missense variant L988F is listed in ClinVar (ID 968833.0) with an “Uncertain” status and is present in gnomAD (variant ID 6‑33443514‑C‑T). Prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, and AlphaMissense‑Optimized. Those that predict a pathogenic outcome are polyPhen‑2 (HumDiv and HumVar) and SIFT. AlphaMissense‑Default remains uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Benign, and Foldetta results are unavailable. Overall, the majority of evidence points to a benign impact, and this is not in conflict with the ClinVar “Uncertain” classification. Thus, the variant is most likely benign based on current predictions. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443514-C-T | 1 | 6.20e-7 | -4.368 | Likely Benign | 0.356 | Ambiguous | Likely Benign | 0.135 | Likely Benign | -1.70 | Neutral | 0.977 | Probably Damaging | 0.900 | Possibly Damaging | 2.69 | Benign | 0.00 | Affected | 4.32 | 2 | 2 | 0 | -1.0 | 34.02 | |||||||||||||||||||||||||||
| c.2983C>T | P995S 2D  AIThe SynGAP1 missense variant P995S is listed in ClinVar (ID 436929.0) with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). The only tool that predicts a pathogenic outcome is SIFT. High‑accuracy assessments further support a benign classification: AlphaMissense‑Optimized predicts benign, and the SGM‑Consensus also indicates “Likely Benign.” No Foldetta (FoldX‑MD/Rosetta) stability result is available, so it does not influence the assessment. Overall, the majority of predictions, including the high‑accuracy tools, point to a benign effect, which does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -4.457 | Likely Benign | 0.071 | Likely Benign | Likely Benign | 0.042 | Likely Benign | -1.03 | Neutral | 0.011 | Benign | 0.015 | Benign | 4.24 | Benign | 0.00 | Affected | 4.32 | 1 | 1 | -1 | 0.8 | -10.04 | ||||||||||||||||||||||||||||||
| c.2989G>A | A997T 2D  AIThe SynGAP1 missense variant A997T is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Only SIFT predicts a pathogenic outcome. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized is benign, and the SGM‑Consensus (derived from the four high‑accuracy predictors) is benign; Foldetta results are unavailable. Taken together, the majority of computational evidence indicates a benign effect, which is consistent with the ClinVar uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -4.102 | Likely Benign | 0.071 | Likely Benign | Likely Benign | 0.085 | Likely Benign | -0.62 | Neutral | 0.224 | Benign | 0.120 | Benign | 4.17 | Benign | 0.00 | Affected | 4.32 | 4 | 1 | 0 | -2.5 | 30.03 | ||||||||||||||||||||||||||||||
| c.2998A>G | I1000V 2D  AIThe SynGAP1 missense variant I1000V is listed in ClinVar (ID 2572013.0) with an “Uncertain” clinical significance and is not reported in gnomAD. Functional prediction tools that assess evolutionary conservation and structural impact (REVEL, SIFT, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, PROVEAN, AlphaMissense‑Default) all converge on a benign outcome. No tool in the dataset predicts pathogenicity. High‑accuracy predictors reinforce this consensus: AlphaMissense‑Optimized reports a benign effect, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) indicates a likely benign classification. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Based on the collective evidence, the variant is most likely benign, and this assessment does not contradict the ClinVar status, which remains uncertain. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 2 | -4.102 | Likely Benign | 0.098 | Likely Benign | Likely Benign | 0.086 | Likely Benign | -0.20 | Neutral | 0.437 | Benign | 0.170 | Benign | 2.76 | Benign | 0.81 | Tolerated | 4.32 | 4 | 3 | 4 | -0.3 | -14.03 | ||||||||||||||||||||||||||||||
| c.29G>A | R10Q 2D  AIThe SynGAP1 missense variant R10Q is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33420293‑G‑A). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). Only SIFT predicts a pathogenic outcome. High‑accuracy assessments further support a benign classification: AlphaMissense‑Optimized is benign, and the SGM‑Consensus is “Likely Benign.” Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the preponderance of evidence indicates that R10Q is most likely benign, which does not contradict the current ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 2 | 6-33420293-G-A | 20 | 1.30e-5 | -4.438 | Likely Benign | 0.185 | Likely Benign | Likely Benign | 0.084 | Likely Benign | 0.03 | Neutral | 0.121 | Benign | 0.004 | Benign | 4.17 | Benign | 0.00 | Affected | 4.32 | 1 | 1 | 1 | 1.0 | -28.06 | |||||||||||||||||||||||||||
| c.29G>C | R10P 2D  AIThe SynGAP1 missense variant R10P is listed in ClinVar with an uncertain significance and is present in gnomAD (ID 6‑33420293‑G‑C). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Only SIFT predicts a pathogenic outcome. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a likely benign classification. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized is benign, and the SGM‑Consensus (majority of the four high‑accuracy tools) is benign; Foldetta results are unavailable. Overall, the collective evidence points to a benign effect for R10P, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 2 | 6-33420293-G-C | 2 | 1.30e-6 | -3.772 | Likely Benign | 0.162 | Likely Benign | Likely Benign | 0.220 | Likely Benign | -0.05 | Neutral | 0.233 | Benign | 0.026 | Benign | 4.13 | Benign | 0.00 | Affected | 4.32 | 1 | 0 | -2 | 2.9 | -59.07 | |||||||||||||||||||||||||||
| c.3002T>C | L1001P 2D  AIThe SynGAP1 missense variant L1001P is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. In contrast, polyPhen‑2 (HumDiv and HumVar) and SIFT all predict a pathogenic impact. The SGM‑Consensus, which aggregates the majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as “Likely Benign.” High‑accuracy assessments further support a benign classification: AlphaMissense‑Optimized predicts benign, and the SGM‑Consensus (majority vote) also indicates benign. The Foldetta protein‑folding stability analysis is unavailable for this variant. Overall, the majority of evidence—including high‑accuracy tools—points to a benign effect, and this conclusion does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -3.071 | Likely Benign | 0.209 | Likely Benign | Likely Benign | 0.113 | Likely Benign | -1.02 | Neutral | 0.966 | Probably Damaging | 0.690 | Possibly Damaging | 2.65 | Benign | 0.00 | Affected | 4.32 | 4 | -3 | -3 | -5.4 | -16.04 | ||||||||||||||||||||||||||||||
| c.3009C>G | S1003R 2D  AIThe SynGAP1 missense variant S1003R (ClinVar ID 1798770.0) is listed as Uncertain in ClinVar and is not present in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, PROVEAN, and ESM1b, while pathogenic predictions are reported by polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. High‑accuracy assessment shows AlphaMissense‑Optimized classifying the variant as pathogenic; the SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is inconclusive (two benign, two pathogenic), and Foldetta stability analysis is unavailable. Overall, the majority of evidence points to a pathogenic effect, which contrasts with the ClinVar designation of Uncertain. Thus, the variant is most likely pathogenic, contradicting the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | -5.113 | Likely Benign | 0.991 | Likely Pathogenic | Likely Pathogenic | 0.141 | Likely Benign | -1.88 | Neutral | 0.999 | Probably Damaging | 0.996 | Probably Damaging | 2.48 | Pathogenic | 0.00 | Affected | 3.77 | 5 | 0 | -1 | -3.7 | 69.11 | |||||||||||||||||||||||||||||||
| c.3023A>G | D1008G 2D  AIThe SynGAP1 D1008G missense variant (ClinVar ID 2963386.0) is listed as Uncertain in ClinVar and is present in gnomAD (ID 6‑33443575‑A‑G). Prediction tools that agree on a benign effect include REVEL, ESM1b, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and AlphaMissense‑Default. High‑accuracy assessments show AlphaMissense‑Optimized classifying the variant as benign, while the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) is inconclusive due to a 2‑to‑2 split, and Foldetta results are unavailable. Overall, the balance of evidence leans toward a pathogenic interpretation, which does not contradict the current ClinVar designation of Uncertain. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | 6-33443575-A-G | 1 | 6.20e-7 | -3.213 | Likely Benign | 0.742 | Likely Pathogenic | Likely Benign | 0.203 | Likely Benign | -2.84 | Deleterious | 0.999 | Probably Damaging | 0.997 | Probably Damaging | 2.65 | Benign | 0.01 | Affected | 3.77 | 5 | -1 | 1 | 3.1 | -58.04 | ||||||||||||||||||||||||||||
| c.3026A>C | E1009A 2D  AIThe SynGAP1 missense variant E1009A is listed in ClinVar (ID 2238288.0) with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, ESM1b, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, FATHMM, and AlphaMissense‑Default. The SGM‑Consensus, which is a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as “Likely Pathogenic.” High‑accuracy assessments show AlphaMissense‑Optimized as benign, SGM‑Consensus as likely pathogenic, and Foldetta (a protein‑folding stability method combining FoldX‑MD and Rosetta) results are unavailable. Overall, the majority of predictions (six pathogenic vs. three benign) lean toward a pathogenic impact, and this conclusion does not contradict the ClinVar status, which remains uncertain. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | Uncertain | 1 | -3.118 | Likely Benign | 0.679 | Likely Pathogenic | Likely Benign | 0.109 | Likely Benign | -3.06 | Deleterious | 0.980 | Probably Damaging | 0.630 | Possibly Damaging | 2.39 | Pathogenic | 0.01 | Affected | 3.77 | 5 | 0 | -1 | 5.3 | -58.04 | ||||||||||||||||||||||||||||||
| c.3038C>G | S1013C 2D  AIThe SynGAP1 missense variant S1013C is listed in ClinVar with an uncertain significance (ClinVar ID 934570.0) and is present in gnomAD (ID 6‑33443590‑C‑G). Prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta results are unavailable. Overall, the majority of evidence points to a benign impact. This conclusion does not contradict the ClinVar status, which remains uncertain. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443590-C-G | 4 | 2.48e-6 | -6.745 | Likely Benign | 0.110 | Likely Benign | Likely Benign | 0.058 | Likely Benign | -2.06 | Neutral | 0.898 | Possibly Damaging | 0.579 | Possibly Damaging | 2.64 | Benign | 0.05 | Affected | 3.77 | 5 | 0 | -1 | 3.3 | 16.06 | |||||||||||||||||||||||||||
| c.303C>A | H101Q 2D  AIThe SynGAP1 missense variant H101Q is listed in ClinVar with an uncertain significance (ClinVar ID 1307533.0) and is present in gnomAD (ID 6‑33432168‑C‑A). Prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a likely benign classification. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus (majority vote) also benign; Foldetta stability analysis is unavailable. Overall, the majority of evidence points to a benign impact, and this is consistent with the ClinVar uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33432168-C-A | 1 | 6.20e-7 | -2.827 | Likely Benign | 0.124 | Likely Benign | Likely Benign | 0.147 | Likely Benign | -0.37 | Neutral | 0.824 | Possibly Damaging | 0.880 | Possibly Damaging | 4.24 | Benign | 0.00 | Affected | 4.32 | 1 | 3 | 0 | -0.3 | -9.01 | |||||||||||||||||||||||||||
| c.3041G>T | G1014V 2D  AIThe SynGAP1 missense variant G1014V is listed in ClinVar (ID 809922.0) with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict benign. Only polyPhen‑2 HumDiv indicates a pathogenic outcome, while the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) classifies the variant as Likely Benign. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized is benign and the SGM‑Consensus is Likely Benign; Foldetta results are unavailable. Overall, the preponderance of evidence points to a benign effect, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -4.612 | Likely Benign | 0.181 | Likely Benign | Likely Benign | 0.053 | Likely Benign | -2.47 | Neutral | 0.818 | Possibly Damaging | 0.377 | Benign | 2.72 | Benign | 0.06 | Tolerated | 3.77 | 5 | -1 | -3 | 4.6 | 42.08 | ||||||||||||||||||||||||||||||
| c.304T>G | L102V 2D  AIThe SynGAP1 missense variant L102V is listed in ClinVar (ID 1925749.0) with an “Uncertain” status and is present in gnomAD (6‑33432169‑T‑G). Prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Benign). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as Likely Benign; Foldetta results are unavailable. Overall, the majority of evidence points to a benign impact. This consensus does not contradict the ClinVar “Uncertain” classification, which remains inconclusive. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33432169-T-G | 1 | 6.20e-7 | -4.316 | Likely Benign | 0.068 | Likely Benign | Likely Benign | 0.102 | Likely Benign | 0.32 | Neutral | 0.880 | Possibly Damaging | 0.899 | Possibly Damaging | 4.21 | Benign | 0.00 | Affected | 4.32 | 1 | 2 | 1 | 0.4 | -14.03 | |||||||||||||||||||||||||||
| c.3053C>T | T1018I 2D  AIThe SynGAP1 missense variant T1018I is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443605‑C‑T). Prediction tools that agree on benign impact include REVEL, polyPhen‑2 HumVar, ESM1b, and AlphaMissense‑Optimized, while those that predict pathogenicity are PROVEAN, polyPhen‑2 HumDiv, SIFT, and FATHMM; AlphaMissense‑Default remains uncertain. High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as pathogenic, and Foldetta results are unavailable. Overall, the predictions are split, with no clear majority leaning toward either benign or pathogenic. Thus, the variant’s impact remains inconclusive, and this uncertainty aligns with ClinVar’s current “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | 6-33443605-C-T | 4 | 2.48e-6 | -3.264 | Likely Benign | 0.524 | Ambiguous | Likely Benign | 0.076 | Likely Benign | -2.55 | Deleterious | 0.586 | Possibly Damaging | 0.304 | Benign | 2.24 | Pathogenic | 0.01 | Affected | 3.77 | 5 | -1 | 0 | 5.2 | 12.05 | ||||||||||||||||||||||||||||
| c.3056G>T | R1019L 2D  AIThe SynGAP1 missense variant R1019L is listed in ClinVar with an “Uncertain” status (ClinVar ID 3364537.0) and is present in gnomAD (gnomAD ID 6‑33443608‑G‑T). Prediction tools that agree on a benign effect include REVEL, ESM1b, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect comprise PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, and AlphaMissense‑Default. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as “Likely Pathogenic.” High‑accuracy assessments show AlphaMissense‑Optimized as benign, while the SGM‑Consensus (majority vote) remains pathogenic. Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, has no available result for this variant. Overall, the majority of computational evidence points to a pathogenic impact, which does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | Uncertain | 1 | 6-33443608-G-T | 2 | 1.24e-6 | -5.194 | Likely Benign | 0.752 | Likely Pathogenic | Likely Benign | 0.110 | Likely Benign | -3.57 | Deleterious | 0.800 | Possibly Damaging | 0.573 | Possibly Damaging | 2.40 | Pathogenic | 0.01 | Affected | 3.77 | 5 | -2 | -3 | 8.3 | -43.03 | |||||||||||||||||||||||||||
| c.3059G>C | R1020P 2D  AIThe SynGAP1 missense variant R1020P is listed in ClinVar (ID 3700393.0) with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL and ESM1b, while pathogenic predictions are made by PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, and AlphaMissense‑Default. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, resolves as “Likely Pathogenic” (3 pathogenic vs. 1 benign votes). High‑accuracy assessments show AlphaMissense‑Optimized as “Uncertain,” SGM‑Consensus as “Likely Pathogenic,” and Foldetta (a protein‑folding stability method combining FoldX‑MD and Rosetta outputs) has no available result for this variant. Overall, the majority of evidence points to a pathogenic effect, and this conclusion does not contradict the current ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | Uncertain | 1 | -3.491 | Likely Benign | 0.902 | Likely Pathogenic | Ambiguous | 0.205 | Likely Benign | -3.50 | Deleterious | 0.999 | Probably Damaging | 0.977 | Probably Damaging | 2.46 | Pathogenic | 0.00 | Affected | 0 | -2 | 2.9 | -59.07 | ||||||||||||||||||||||||||||||||
| c.3059G>T | R1020L 2D  AISynGAP1 missense variant R1020L is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions come from REVEL, ESM1b, and FATHMM, while pathogenic predictions are made by PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and AlphaMissense‑Default. The high‑accuracy AlphaMissense‑Optimized score is uncertain, and the SGM consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) is inconclusive; Foldetta stability analysis is unavailable. Overall, the majority of evidence points toward a pathogenic effect, which contrasts with the ClinVar designation of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | -6.031 | Likely Benign | 0.907 | Likely Pathogenic | Ambiguous | 0.216 | Likely Benign | -4.03 | Deleterious | 0.990 | Probably Damaging | 0.921 | Probably Damaging | 2.50 | Benign | 0.00 | Affected | 3.77 | 5 | -3 | -2 | 8.3 | -43.03 | |||||||||||||||||||||||||||||||
| c.3092T>C | M1031T 2D  AIThe SynGAP1 missense variant M1031T is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443644‑T‑C). In silico prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). No tool predicts a pathogenic outcome; the only inconclusive result is AlphaMissense‑Default, which is treated as unavailable. High‑accuracy assessments further support a benign classification: AlphaMissense‑Optimized predicts benign, the SGM‑Consensus is “Likely Benign,” and Foldetta data are not available. **Thus, the variant is most likely benign, and this conclusion does not contradict the ClinVar “Uncertain” status.** Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443644-T-C | 2 | 1.24e-6 | -1.863 | Likely Benign | 0.540 | Ambiguous | Likely Benign | 0.085 | Likely Benign | -0.24 | Neutral | 0.002 | Benign | 0.005 | Benign | 2.67 | Benign | 1.00 | Tolerated | 3.77 | 5 | -1 | -1 | -2.6 | -30.09 | |||||||||||||||||||||||||||
| c.3103C>A | P1035T 2D  AIThe SynGAP1 missense variant P1035T is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Tools that predict a pathogenic outcome are polyPhen‑2 HumDiv and polyPhen‑2 HumVar. AlphaMissense‑Default remains uncertain, and no Foldetta stability assessment is available. High‑accuracy evidence from AlphaMissense‑Optimized and the SGM‑Consensus both support a benign classification, while the absence of a Foldetta result does not alter this view. Overall, the majority of predictions indicate a benign effect, and this is consistent with the ClinVar “Uncertain” designation rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -4.447 | Likely Benign | 0.426 | Ambiguous | Likely Benign | 0.087 | Likely Benign | -0.96 | Neutral | 0.901 | Possibly Damaging | 0.537 | Possibly Damaging | 2.72 | Benign | 0.23 | Tolerated | 3.77 | 5 | 0 | -1 | 0.9 | 3.99 | ||||||||||||||||||||||||||||||
| c.3116T>C | I1039T 2D  AIThe SynGAP1 missense variant I1039T is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443668‑T‑C). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, and FATHMM; the SGM‑Consensus score (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also indicates a likely benign outcome. Only AlphaMissense‑Default predicts a pathogenic effect. High‑accuracy assessments show AlphaMissense‑Optimized classifying the variant as benign, while the SGM‑Consensus remains benign; a Foldetta stability analysis is not available. Overall, the majority of computational evidence supports a benign impact, and this is consistent with the ClinVar “Uncertain” classification, so there is no contradiction. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443668-T-C | 12 | 7.43e-6 | -2.465 | Likely Benign | 0.645 | Likely Pathogenic | Likely Benign | 0.193 | Likely Benign | 0.45 | Neutral | 0.004 | Benign | 0.008 | Benign | 2.75 | Benign | 0.10 | Tolerated | 3.77 | 5 | -1 | 0 | -5.2 | -12.05 | |||||||||||||||||||||||||||
| c.3119G>T | G1040V 2D  AIThe SynGAP1 missense variant G1040V is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443671‑G‑T). Prediction tools that agree on a benign effect are ESM1b and AlphaMissense‑Optimized; those that predict a pathogenic effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, FATHMM, AlphaMissense‑Default, and the SGM‑Consensus score (Likely Pathogenic). High‑accuracy assessments show AlphaMissense‑Optimized as benign, the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Pathogenic, and Foldetta results are unavailable. Overall, the majority of predictions indicate a pathogenic impact, and this is not in conflict with the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Pathogenic | Uncertain | 1 | 6-33443671-G-T | 4 | 2.48e-6 | -3.453 | Likely Benign | 0.645 | Likely Pathogenic | Likely Benign | 0.774 | Likely Pathogenic | -2.89 | Deleterious | 0.827 | Possibly Damaging | 0.456 | Possibly Damaging | -0.74 | Pathogenic | 0.01 | Affected | 3.77 | 5 | -1 | -3 | 4.6 | 42.08 | |||||||||||||||||||||||||||
| c.3125A>G | Q1042R 2D  AIThe SynGAP1 missense variant Q1042R is listed in ClinVar (ID 2662705.0) with an “Uncertain” clinical significance and is present in gnomAD (variant ID 6‑33443677‑A‑G). Functional prediction tools largely agree on a benign effect: REVEL, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, and AlphaMissense‑Optimized all classify the change as benign. Only polyPhen‑2 HumDiv predicts a pathogenic outcome, while AlphaMissense‑Default remains uncertain. High‑accuracy assessments reinforce the benign consensus: AlphaMissense‑Optimized reports benign, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates “Likely Benign.” Foldetta results are unavailable. Overall, the majority of evidence supports a benign impact for Q1042R, and this conclusion does not contradict the ClinVar status, which remains uncertain. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 2 | 6-33443677-A-G | 2 | 1.24e-6 | -2.928 | Likely Benign | 0.413 | Ambiguous | Likely Benign | 0.300 | Likely Benign | -1.39 | Neutral | 0.586 | Possibly Damaging | 0.120 | Benign | 5.48 | Benign | 0.12 | Tolerated | 3.77 | 5 | 1 | 1 | -1.0 | 28.06 | |||||||||||||||||||||||||||
| c.3136C>G | P1046A 2D  AIThe SynGAP1 missense variant P1046A is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443688‑C‑G). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote). Only FATHMM predicts a pathogenic outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; a Foldetta stability analysis is unavailable. Overall, the majority of evidence points to a benign impact, and this conclusion does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443688-C-G | 1 | 6.20e-7 | -3.246 | Likely Benign | 0.048 | Likely Benign | Likely Benign | 0.041 | Likely Benign | -1.67 | Neutral | 0.001 | Benign | 0.008 | Benign | 2.39 | Pathogenic | 0.29 | Tolerated | 3.77 | 5 | -1 | 1 | 3.4 | -26.04 | |||||||||||||||||||||||||||
| c.313T>C | S105P 2D  AIThe SynGAP1 missense variant S105P is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Prediction tools largely agree on a benign effect: REVEL, PROVEAN, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all classify the change as benign. Only two tools—polyPhen‑2 HumDiv and SIFT—predict a pathogenic outcome. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as likely benign. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized predicts benign, and the SGM‑Consensus (majority vote) also indicates likely benign. Foldetta, a protein‑folding stability method, has no available result for this variant. Overall, the preponderance of evidence points to a benign effect, which is consistent with the ClinVar uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -3.631 | Likely Benign | 0.166 | Likely Benign | Likely Benign | 0.204 | Likely Benign | 0.03 | Neutral | 0.808 | Possibly Damaging | 0.212 | Benign | 4.00 | Benign | 0.00 | Affected | 4.32 | 1 | -1 | 1 | -0.8 | 10.04 | ||||||||||||||||||||||||||||||
| c.3142G>C | G1048R 2D  AIThe SynGAP1 missense variant G1048R is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Tools that predict a pathogenic effect are REVEL, polyPhen‑2 HumDiv, and polyPhen‑2 HumVar. The high‑accuracy AlphaMissense‑Optimized score is benign, and the SGM‑Consensus also indicates a likely benign outcome; the Foldetta protein‑folding stability assessment is unavailable. Overall, the majority of evidence points to a benign impact, and this conclusion does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -4.305 | Likely Benign | 0.435 | Ambiguous | Likely Benign | 0.503 | Likely Pathogenic | -0.54 | Neutral | 0.919 | Possibly Damaging | 0.728 | Possibly Damaging | 2.54 | Benign | 0.10 | Tolerated | 3.77 | 5 | -2 | -3 | -4.1 | 99.14 | ||||||||||||||||||||||||||||||
| c.314C>T | S105L 2D  AIThe SynGAP1 missense variant S105L is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33432179‑C‑T). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumVar, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv and SIFT. High‑accuracy methods both support a benign outcome: AlphaMissense‑Optimized is benign, and the SGM‑Consensus (majority vote) is benign; Foldetta results are unavailable. Overall, the majority of evidence points to a benign impact, and this does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 2 | 6-33432179-C-T | 4 | 2.48e-6 | -3.710 | Likely Benign | 0.233 | Likely Benign | Likely Benign | 0.095 | Likely Benign | -1.52 | Neutral | 0.828 | Possibly Damaging | 0.048 | Benign | 4.06 | Benign | 0.00 | Affected | 4.32 | 1 | -3 | -2 | 4.6 | 26.08 | |||||||||||||||||||||||||||
| c.3154G>A | G1052R 2D  AISynGAP1 missense variant G1052R is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools cluster into two groups: benign predictions from REVEL, PROVEAN, SIFT, FATHMM, and AlphaMissense‑Optimized; pathogenic predictions from polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and ESM1b; AlphaMissense‑Default remains uncertain. High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized scores the variant as benign, and the SGM consensus (derived from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also favors benign. Foldetta, a protein‑folding stability method, has no available result for this variant. Overall, the majority of evidence points to a benign effect, which does not contradict the ClinVar uncertain status but provides additional support toward a likely benign classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | -9.050 | Likely Pathogenic | 0.383 | Ambiguous | Likely Benign | 0.497 | Likely Benign | -0.41 | Neutral | 0.990 | Probably Damaging | 0.798 | Possibly Damaging | 3.90 | Benign | 0.10 | Tolerated | 3.77 | 5 | -2 | -3 | -4.1 | 99.14 | |||||||||||||||||||||||||||||||
| c.3161G>A | G1054D 2D  AISynGAP1 missense variant G1054D is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumVar, SIFT, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv and ESM1b, while AlphaMissense‑Default remains uncertain. High‑accuracy assessments further support a benign outcome: AlphaMissense‑Optimized scores benign, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) also yields benign. Foldetta, a protein‑folding stability method, has no available result for this variant. Overall, the preponderance of evidence indicates the variant is most likely benign, which does not contradict the current ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | -10.385 | Likely Pathogenic | 0.351 | Ambiguous | Likely Benign | 0.279 | Likely Benign | -0.26 | Neutral | 0.818 | Possibly Damaging | 0.266 | Benign | 4.07 | Benign | 0.37 | Tolerated | 3.77 | 5 | 1 | -1 | -3.1 | 58.04 | |||||||||||||||||||||||||||||||
| c.3170G>A | S1057N 2D  AIThe SynGAP1 missense variant S1057N is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. All available in‑silico predictors classify the change as benign: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all return benign scores, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates “Likely Benign.” No tool predicts pathogenicity. High‑accuracy assessments confirm this: AlphaMissense‑Optimized is benign, SGM‑Consensus is likely benign, while Foldetta (combining FoldX‑MD and Rosetta outputs) has no available result for this variant. Overall, the computational evidence overwhelmingly supports a benign effect, and this is consistent with the ClinVar “Uncertain” classification rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -6.386 | Likely Benign | 0.117 | Likely Benign | Likely Benign | 0.218 | Likely Benign | -0.41 | Neutral | 0.451 | Benign | 0.129 | Benign | 5.25 | Benign | 0.28 | Tolerated | 1 | 1 | -2.7 | 27.03 | ||||||||||||||||||||||||||||||||
| c.3175G>A | G1059R 2D  AIThe SynGAP1 missense variant G1059R is listed in ClinVar with an “Uncertain” significance and is present in the gnomAD database (ID 6‑33443727‑G‑A). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are SIFT and ESM1b, while AlphaMissense‑Default remains uncertain. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, resolves to a benign prediction (2 benign vs. 1 pathogenic). High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM Consensus also benign; Foldetta’s protein‑folding stability analysis is unavailable for this variant. Overall, the collective evidence points to a benign impact, and this conclusion does not contradict the ClinVar “Uncertain” status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | 6-33443727-G-A | 68 | 4.23e-5 | -8.452 | Likely Pathogenic | 0.376 | Ambiguous | Likely Benign | 0.333 | Likely Benign | -0.55 | Neutral | 0.001 | Benign | 0.001 | Benign | 2.53 | Benign | 0.00 | Affected | 4.32 | 2 | -3 | -2 | -4.1 | 99.14 | ||||||||||||||||||||||||||||
| c.3176G>C | G1059A 2D  AIThe SynGAP1 missense variant G1059A is listed in ClinVar (ID 1420036.0) with an “Uncertain” clinical significance and is present in gnomAD (variant ID 6‑33443728‑G‑C). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Only SIFT predicts a pathogenic outcome. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also indicates a likely benign effect. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta results are unavailable. Overall, the majority of computational evidence supports a benign impact for G1059A, which does not contradict the ClinVar “Uncertain” status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443728-G-C | 4 | 2.49e-6 | -6.754 | Likely Benign | 0.081 | Likely Benign | Likely Benign | 0.329 | Likely Benign | -0.17 | Neutral | 0.001 | Benign | 0.002 | Benign | 2.56 | Benign | 0.00 | Affected | 4.32 | 2 | 1 | 0 | 2.2 | 14.03 | |||||||||||||||||||||||||||
| c.3178G>A | G1060S 2D  AIThe SynGAP1 missense variant G1060S is listed in ClinVar with an uncertain significance (ClinVar ID 1512003.0) and is present in gnomAD (variant ID 6‑33443730‑G‑A). All evaluated in‑silico predictors classify the change as benign: REVEL, PROVEAN, PolyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. No tool predicts pathogenicity. High‑accuracy assessments reinforce this benign view: AlphaMissense‑Optimized reports benign, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) indicates “Likely Benign.” Foldetta, a protein‑folding stability method combining FoldX‑MD and Rosetta outputs, did not provide a result for this variant. Overall, the computational evidence overwhelmingly supports a benign effect, which is consistent with the ClinVar uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443730-G-A | -4.759 | Likely Benign | 0.082 | Likely Benign | Likely Benign | 0.376 | Likely Benign | -0.08 | Neutral | 0.271 | Benign | 0.054 | Benign | 2.69 | Benign | 0.49 | Tolerated | 4.32 | 2 | 1 | 0 | -0.4 | 30.03 | |||||||||||||||||||||||||||||
| c.3181G>A | G1061S 2D  AIThe SynGAP1 missense variant G1061S is listed in ClinVar (ID 3571724.0) with an uncertain significance designation and is not reported in gnomAD. Functional prediction tools largely agree on a benign effect: REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized all predict benign, while only SIFT indicates a pathogenic effect. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, also reports a likely benign outcome. High‑accuracy assessments confirm this: AlphaMissense‑Optimized is benign and the SGM‑Consensus is likely benign; Foldetta stability analysis is unavailable. Overall, the consensus of the majority of prediction tools and the high‑accuracy methods supports a benign classification for G1061S, which is consistent with its ClinVar uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -4.891 | Likely Benign | 0.079 | Likely Benign | Likely Benign | 0.283 | Likely Benign | -0.68 | Neutral | 0.004 | Benign | 0.004 | Benign | 4.00 | Benign | 0.00 | Affected | 1 | 0 | -0.4 | 30.03 | ||||||||||||||||||||||||||||||||
| c.3192G>C | Q1064H 2D  AIThe SynGAP1 missense variant Q1064H is listed in ClinVar with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. In contrast, polyPhen‑2 (HumDiv and HumVar) and SIFT all predict a pathogenic impact. The SGM‑Consensus, which aggregates the majority vote from AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as “Likely Benign.” High‑accuracy assessments further support a benign classification: AlphaMissense‑Optimized predicts benign, and the SGM‑Consensus (majority vote) also indicates benign. The Foldetta protein‑folding stability analysis is unavailable for this variant. Overall, the majority of evidence—including high‑accuracy tools—points to a benign effect, and this conclusion does not contradict the current ClinVar “Uncertain” status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -4.576 | Likely Benign | 0.162 | Likely Benign | Likely Benign | 0.063 | Likely Benign | -0.66 | Neutral | 0.938 | Possibly Damaging | 0.596 | Possibly Damaging | 4.15 | Benign | 0.05 | Affected | 3 | 0 | 0.3 | 9.01 | ||||||||||||||||||||||||||||||||
| c.3209_3210delinsCA | R1070T 2D  AIThe SynGAP1 missense variant R1070T is listed in ClinVar (ID 2759838.0) with an “Uncertain” clinical significance and is not reported in gnomAD. Prediction tools that agree on a benign effect include PROVEAN, ESM1b, FATHMM, and the SGM‑Consensus (which aggregates these three benign calls with the pathogenic AlphaMissense‑Default to yield a Likely Benign verdict). Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and AlphaMissense‑Default. AlphaMissense‑Optimized returns an uncertain result. High‑accuracy assessments show AlphaMissense‑Optimized as uncertain, the SGM‑Consensus (majority vote from AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) as Likely Benign, and no Foldetta stability data is available. Overall, the balance of evidence leans toward a benign impact, which is consistent with the ClinVar “Uncertain” status and does not contradict it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -5.093 | Likely Benign | 0.860 | Likely Pathogenic | Ambiguous | -2.35 | Neutral | 0.948 | Possibly Damaging | 0.507 | Possibly Damaging | 3.78 | Benign | 0.01 | Affected | 3.77 | 5 | -1 | -1 | 3.8 | -55.08 | ||||||||||||||||||||||||||||||||
| c.3223C>A | Q1075K 2D  AIThe SynGAP1 missense variant Q1075K (ClinVar ID 2762879.0) is listed as “Uncertain” in ClinVar and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and AlphaMissense‑Default. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is “Likely Benign” because three of the four contributing tools predict benign. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus (majority vote) as benign; Foldetta stability analysis is unavailable. Overall, the majority of evidence points to a benign impact, and this conclusion does not contradict the ClinVar “Uncertain” status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -5.135 | Likely Benign | 0.728 | Likely Pathogenic | Likely Benign | 0.134 | Likely Benign | -0.67 | Neutral | 0.963 | Probably Damaging | 0.959 | Probably Damaging | 2.75 | Benign | 1.00 | Tolerated | 3.77 | 5 | 1 | 1 | -0.4 | 0.04 | ||||||||||||||||||||||||||||||
| c.3233T>A | V1078D 2D  AIThe SynGAP1 missense variant V1078D is listed in ClinVar (ID 2993122.0) with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), ESM1b, and FATHMM. Tools that predict a pathogenic effect are AlphaMissense‑Default, AlphaMissense‑Optimized, and SIFT. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. High‑accuracy assessments show AlphaMissense‑Optimized as pathogenic, SGM‑Consensus as likely benign, and Foldetta (a protein‑folding stability method combining FoldX‑MD and Rosetta) has no available result for this variant. Overall, the majority of predictions lean toward a benign impact, and this is consistent with the ClinVar “Uncertain” designation; there is no contradiction with the existing ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -5.155 | Likely Benign | 0.979 | Likely Pathogenic | Likely Pathogenic | 0.158 | Likely Benign | -1.45 | Neutral | 0.003 | Benign | 0.008 | Benign | 3.84 | Benign | 0.00 | Affected | 3.77 | 5 | -3 | -2 | -7.7 | 15.96 | ||||||||||||||||||||||||||||||
| c.3237C>A | S1079R 2D  AIThe SynGAP1 missense variant S1079R is listed in ClinVar with an “Uncertain” status and is present in gnomAD (gnomAD ID 6‑33443789‑C‑A). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, ESM1b, and FATHMM. Those that predict a pathogenic effect are SIFT and AlphaMissense‑Default. AlphaMissense‑Optimized is inconclusive. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. High‑accuracy assessments are limited: AlphaMissense‑Optimized remains uncertain, SGM‑Consensus is benign, and Foldetta (combining FoldX‑MD and Rosetta outputs) has no available result for this variant. Overall, the preponderance of evidence points to a benign impact, which does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443789-C-A | 4 | 2.51e-6 | -4.579 | Likely Benign | 0.955 | Likely Pathogenic | Ambiguous | 0.123 | Likely Benign | -1.81 | Neutral | 0.177 | Benign | 0.075 | Benign | 3.86 | Benign | 0.00 | Affected | 3.77 | 5 | 0 | -1 | -3.7 | 69.11 | |||||||||||||||||||||||||||
| c.3238G>T | A1080S 2D  AIThe SynGAP1 missense variant A1080S is listed in ClinVar (ID 2703014.0) with an “Uncertain” status and is present in gnomAD (variant ID 6‑33443790‑G‑T). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. Only polyPhen‑2 HumDiv predicts a pathogenic outcome. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports the variant as “Likely Benign.” High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta results are not available. Overall, the preponderance of evidence points to a benign effect, and this conclusion does not contradict the ClinVar designation, which remains uncertain. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443790-G-T | 1 | 6.26e-7 | -3.277 | Likely Benign | 0.108 | Likely Benign | Likely Benign | 0.103 | Likely Benign | 0.01 | Neutral | 0.702 | Possibly Damaging | 0.346 | Benign | 4.16 | Benign | 0.08 | Tolerated | 3.77 | 5 | 1 | 1 | -2.6 | 16.00 | |||||||||||||||||||||||||||
| c.323A>G | K108R 2D  AIThe SynGAP1 missense variant K108R is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33432188‑A‑G). Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, and AlphaMissense‑Optimized. In contrast, polyPhen‑2 (HumDiv and HumVar) predict a pathogenic impact. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. High‑accuracy assessments further support benignity: AlphaMissense‑Optimized is benign, and the SGM‑Consensus itself is benign; Foldetta results are unavailable. Overall, the majority of evidence points to a benign effect, and this conclusion does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33432188-A-G | 6 | 3.72e-6 | -2.892 | Likely Benign | 0.148 | Likely Benign | Likely Benign | 0.184 | Likely Benign | 0.37 | Neutral | 0.993 | Probably Damaging | 0.956 | Probably Damaging | 4.22 | Benign | 1.00 | Tolerated | 3.61 | 5 | 3 | 2 | -0.6 | 28.01 | |||||||||||||||||||||||||||
| c.3250C>G | P1084A 2D  AIThe SynGAP1 missense variant P1084A is listed in ClinVar (ID 2827308.0) with an “Uncertain” status and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (which itself is “Likely Benign”). In contrast, PROVEAN and polyPhen‑2 HumDiv predict a pathogenic impact. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; a Foldetta stability analysis is unavailable. Overall, the majority of evidence points to a benign effect, and this conclusion does not contradict the ClinVar “Uncertain” classification. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -3.928 | Likely Benign | 0.066 | Likely Benign | Likely Benign | 0.114 | Likely Benign | -2.54 | Deleterious | 0.649 | Possibly Damaging | 0.157 | Benign | 4.05 | Benign | 0.35 | Tolerated | 3.77 | 5 | -1 | 1 | 3.4 | -26.04 | ||||||||||||||||||||||||||||||
| c.3251C>A | P1084H 2D  AIThe SynGAP1 missense variant P1084H is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443803‑C‑A). Prediction tools that agree on a benign effect include REVEL, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (which reports “Likely Benign”). Tools that predict a pathogenic effect are PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, and SIFT. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta results are unavailable. Overall, the majority of evidence points to a benign impact, and this is consistent with the ClinVar designation of uncertainty rather than a definitive pathogenic claim. Thus, the variant is most likely benign, and its prediction profile does not contradict the current ClinVar status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443803-C-A | 1 | 6.31e-7 | -4.125 | Likely Benign | 0.323 | Likely Benign | Likely Benign | 0.134 | Likely Benign | -3.16 | Deleterious | 0.997 | Probably Damaging | 0.840 | Possibly Damaging | 3.96 | Benign | 0.00 | Affected | 3.77 | 5 | -2 | 0 | -1.6 | 40.02 | |||||||||||||||||||||||||||
| c.3253C>T | R1085W 2D  AISynGAP1 missense variant R1085W is listed in ClinVar as Uncertain and is present in gnomAD (ID 6‑33443805‑C‑T). Prediction tools that classify the variant as benign include REVEL, ESM1b, and FATHMM, whereas pathogenic predictions come from PROVEAN, polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and AlphaMissense‑Default. The high‑accuracy AlphaMissense‑Optimized score is Uncertain, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN) and Foldetta stability assessment are unavailable. Overall, the majority of available predictions (five pathogenic vs. three benign) indicate a pathogenic effect. Thus, the variant is most likely pathogenic, which contradicts the ClinVar designation of Uncertain. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | 6-33443805-C-T | 2 | 1.26e-6 | -6.339 | Likely Benign | 0.821 | Likely Pathogenic | Ambiguous | 0.202 | Likely Benign | -3.15 | Deleterious | 1.000 | Probably Damaging | 0.996 | Probably Damaging | 2.70 | Benign | 0.00 | Affected | 3.77 | 5 | -3 | 2 | 3.6 | 30.03 | ||||||||||||||||||||||||||||
| c.3254G>A | R1085Q 2D  AIThe SynGAP1 missense variant R1085Q (ClinVar ID 1729448.0) is listed as ClinVar status Uncertain and is present in gnomAD (6‑33443806‑G‑A). Functional prediction tools show a split opinion: benign predictions come from REVEL, PROVEAN, ESM1b, and FATHMM, whereas pathogenic predictions are reported by polyPhen‑2 HumDiv, polyPhen‑2 HumVar, SIFT, and AlphaMissense‑Default. The SGM‑Consensus, which aggregates AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is classified as Likely Benign. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized predicts benign, and the SGM‑Consensus (majority vote) also favors benign. No Foldetta stability analysis is available for this variant. Overall, the majority of computational evidence points to a benign effect, which is consistent with the ClinVar uncertain designation rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443806-G-A | 5 | 3.16e-6 | -3.843 | Likely Benign | 0.589 | Likely Pathogenic | Likely Benign | 0.224 | Likely Benign | -1.43 | Neutral | 0.998 | Probably Damaging | 0.988 | Probably Damaging | 2.73 | Benign | 0.02 | Affected | 3.77 | 5 | 1 | 1 | 1.0 | -28.06 | |||||||||||||||||||||||||||
| c.3260C>T | S1087F 2D  AISynGAP1 missense variant S1087F is listed in ClinVar with an uncertain significance and is not reported in gnomAD. Functional prediction tools show a split: benign calls come from REVEL, ESM1b, FATHMM, and AlphaMissense‑Optimized, while pathogenic calls come from PROVEAN, polyPhen‑2 (HumDiv and HumVar), and SIFT; AlphaMissense‑Default remains uncertain. High‑accuracy assessments reinforce the benign trend: AlphaMissense‑Optimized predicts benign, and the SGM Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN) also yields benign. Foldetta, a protein‑folding stability method, has no available result for this variant. Overall, the majority of reliable predictors and the two high‑accuracy tools suggest a benign effect, which does not contradict the ClinVar uncertain status. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | -3.843 | Likely Benign | 0.497 | Ambiguous | Likely Benign | 0.105 | Likely Benign | -2.75 | Deleterious | 0.990 | Probably Damaging | 0.796 | Possibly Damaging | 2.56 | Benign | 0.03 | Affected | 3.77 | 5 | -2 | -3 | 3.6 | 60.10 | |||||||||||||||||||||||||||||||
| c.3262A>G | S1088G 2D  AIThe SynGAP1 missense variant S1088G is listed in ClinVar (ID 2742833.0) with an “Uncertain” clinical significance and is not reported in gnomAD. Functional prediction tools that agree on a benign effect include REVEL, PROVEAN, ESM1b, FATHMM, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (Likely Benign). In contrast, polyPhen‑2 (HumDiv and HumVar) and SIFT all predict a pathogenic impact. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus also as Likely Benign; a Foldetta stability analysis is unavailable. Overall, the majority of evidence points to a benign effect, and this is consistent with the ClinVar “Uncertain” status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -5.034 | Likely Benign | 0.285 | Likely Benign | Likely Benign | 0.163 | Likely Benign | -1.83 | Neutral | 0.979 | Probably Damaging | 0.973 | Probably Damaging | 2.63 | Benign | 0.03 | Affected | 3.77 | 5 | 0 | 1 | 0.4 | -30.03 | ||||||||||||||||||||||||||||||
| c.3287A>C | E1096A 2D  AIThe SynGAP1 missense variant E1096A is listed in ClinVar (ID 2579889.0) with an uncertain significance annotation and is not reported in gnomAD. Consensus from multiple in‑silico predictors shows a predominance of benign calls: REVEL, PROVEAN, polyPhen‑2 HumVar, SIFT, ESM1b, FATHMM, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, PROVEAN). Only polyPhen‑2 HumDiv assigns a pathogenic label, while AlphaMissense‑Default remains uncertain. High‑accuracy assessments further support a benign interpretation: AlphaMissense‑Optimized predicts benign, and the SGM‑Consensus also indicates likely benign; Foldetta, a protein‑folding stability method, has no available result for this variant. Overall, the aggregate evidence points to a benign effect, which is consistent with the ClinVar uncertain status rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -4.504 | Likely Benign | 0.510 | Ambiguous | Likely Benign | 0.164 | Likely Benign | -1.37 | Neutral | 0.626 | Possibly Damaging | 0.184 | Benign | 2.77 | Benign | 0.16 | Tolerated | 3.77 | 5 | -1 | 0 | 5.3 | -58.04 | ||||||||||||||||||||||||||||||
| c.3304G>A | A1102T 2D  AIThe SynGAP1 missense variant A1102T is listed in ClinVar with an “Uncertain” status and is present in gnomAD (ID 6‑33443856‑G‑A). Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Only FATHMM predicts a pathogenic outcome. The SGM‑Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, reports a “Likely Benign” classification. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta results are unavailable. Overall, the majority of evidence points to a benign effect, and this does not contradict the ClinVar “Uncertain” designation. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | 6-33443856-G-A | 11 | 7.17e-6 | -3.540 | Likely Benign | 0.070 | Likely Benign | Likely Benign | 0.044 | Likely Benign | -0.30 | Neutral | 0.001 | Benign | 0.001 | Benign | 2.32 | Pathogenic | 0.95 | Tolerated | 3.77 | 5 | 1 | 0 | -2.5 | 30.03 | |||||||||||||||||||||||||||
| c.3304G>C | A1102P 2D  AIThe SynGAP1 missense variant A1102P is listed in ClinVar (ID 2789225.0) with an “Uncertain” status and is not reported in gnomAD. Prediction tools that agree on a benign effect include REVEL, PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, ESM1b, AlphaMissense‑Default, AlphaMissense‑Optimized, and the SGM‑Consensus (majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN). Only FATHMM predicts a pathogenic outcome. High‑accuracy assessments show AlphaMissense‑Optimized as benign and the SGM‑Consensus as likely benign; Foldetta results are unavailable. Overall, the majority of evidence points to a benign impact, and this is consistent with the ClinVar “Uncertain” classification rather than contradicting it. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Likely Benign | Uncertain | 1 | -5.120 | Likely Benign | 0.077 | Likely Benign | Likely Benign | 0.118 | Likely Benign | -0.97 | Neutral | 0.000 | Benign | 0.002 | Benign | 2.26 | Pathogenic | 0.13 | Tolerated | 3.77 | 5 | -1 | 1 | -3.4 | 26.04 | ||||||||||||||||||||||||||||||
| c.3307C>T | R1103C 2D  AISynGAP1 missense variant R1103C is listed in ClinVar with an uncertain significance and is present in gnomAD (ID 6‑33443859‑C‑T). Prediction tools that agree on a benign effect include REVEL, ESM1b, AlphaMissense‑Default, and AlphaMissense‑Optimized. Tools that predict a pathogenic effect are PROVEAN, polyPhen‑2 (HumDiv and HumVar), SIFT, and FATHMM. The SGM Consensus, derived from a majority vote of AlphaMissense‑Default, ESM1b, FATHMM, and PROVEAN, is inconclusive (two benign, two pathogenic). AlphaMissense‑Optimized reports a benign outcome, while Foldetta results are unavailable. Overall, the balance of evidence favors a pathogenic interpretation, which is in contrast to the ClinVar designation of uncertain significance. Disclaimer: This summary was generated using AI and should be interpreted alongside expert review. | Uncertain | 1 | 6-33443859-C-T | 6 | 3.92e-6 | -2.440 | Likely Benign | 0.246 | Likely Benign | Likely Benign | 0.140 | Likely Benign | -3.01 | Deleterious | 0.996 | Probably Damaging | 0.787 | Possibly Damaging | 2.41 | Pathogenic | 0.01 | Affected | 3.77 | 5 | -3 | -4 | 7.0 | -53.05 | ||||||||||||||||||||||||||||
Found 757 rows. Show 200 rows per page. Page 3/4 « Previous | Next »